Abstract
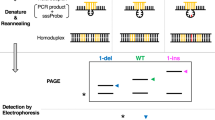
The surprising finding that amplification of genomic DNA can be directed by only one oligonucleotide primer of arbitrary sequence to produce a characteristic spectrum of short DNA products of varying complexity, was applied as a strategy to detect genetic differences between organisms. This approach, DNA amplification fingerprinting (DAF), does not depend on cloning or DNA sequence information and can generate fingerprints from DNA of viral, bacterial, fungal, plant and animal origins. Primers as short as 5 nucleotides in length can produce complex banding patterns that are resolved by polyacrylamide gel electrophoresis and silver staining. Amplification fragment length polymorphisms (AFLPs) were detected between different human individuals as well as between soybean cultivars. It is anticipated that DAF will have wide application for DNA analysis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Jeffreys, A.J., Wilson, V. and Thein, S.L. 1985. Hypervariable ‘minisatcllite’ regions in human DNA. Nature 314: 67–73.
Nakamura, Y., Leppert, M., O'Connell, P., Wolff, R., Holm, T., Culver, M., Martin, C., Fujimoto, E., Hoff, M., Kumlin, E. and White, R. 1987. Variable number of tandem repeat (VNTR) markers for human gene mapping. Science 235: 1616–1622.
Wong, Z., Wilson, V., Patel, I., Povey, S. and Jeffreys, A.J. 1987. Characterization of a panel of highly variable minisatellites cloned from human DNA. Annu. Hum. Genet. 51: 269–288.
Watkins, P.C. 1988. Restriction fragment length polymorphism (RFLP): applications in human chromosome mapping and genetic disease research. Biotechniques 6: 310–320.
Donis-Keller, H., Green, P., Helms, C., Cartinhour, S. et al. 1987. A genetic linkage map of the human genome. Cell 51: 319–337.
Landegren, U., Kaiser, R., Caskey, C.T. and Hood, L. 1988. DNA diagnostics-molecular techniques and automation. Science 242: 229–237.
Jeffreys, A.J., Wilson, V. and Thein, S.L. 1985. Individual-specific ‘fingerprints’ of human DNA. Nature 316: 76–79.
Vassart, G., Georges, M., Monsieur, R., Broca, H., Lequarre, A.S. and Cristophe, D. 1987. A sequence in M13 phage detects hypervariable minisatellites in human and animal DNA. Science 235: 683–684.
Armour, J.A.L., Wong, Z., Wilson, V., Royle, N.J. and Jeffreys, A.J. 1989. Sequences flanking the repeat arrays of human minisatellites: association with tandem and dispersed repeat elements. Nucleic Acids Res. 17: 4925–4935.
Wong, Z., Wilson, V., Jeffreys, A.J. and Thien, S.L. 1986. Cloning a selected fragment from a human DNA fingerprint: isolation of an extremely polymorphic minisatellite. Nucleic Acids Res. 14: 4607–4616.
Jeffreys, A.J., Wilson, V., Neumann, R. and Keyte, J. 1988. Amplification of human minisatellites by the polymerase chain reaction: towards DNA fingerprinting of single cells. Nucleic Acids Res. 16: 10953–10971.
Boerwinkle, E., Xiong, W., Fourest, E. and Chan, L. 1989. Rapid typing of tandemly repeated hypervariable loci by the polymerase chain reaction: application to the apolipoprotein B 3′ hypervariable region. Proc. Natl. Acad. Sci. USA 86: 212–216.
Horn, G.T., Richards, B. and Klinger, K.W. 1989. Amplification of a highly polymorphic VNTR segment by the polymerase chain reaction. Nucleic Acids Res. 17: 2140.
Tautz, D. 1989. Hypervariability of simple sequences as a general source for polymorphic DNA markers. Nucleic Acids Res. 17: 6463–6472.
Jeffreys, A.J., Neumann, R. and Wilson, V. 1990. Repeat unit sequence variation in minisatellites: a novel source of DNA polymorphism for studying variation and mutation by single molecule analysis. Cell 60: 473–485.
Williams, J.G.K., Kubelik, A.R., Livak, K.J., Rafalski, J.A. and Tingey, S.V. 1990. DNA polymorphisms amplified by arbitrary primers are useful as genetic markers. Nucleic Acids Res. 18: 6531–6535.
Welsh, J. and McClelland, M. 1990. Fingerprinting genomes using PCR with arbitrary primers. Nucl. Acids Res. 18: 7213–7218.
Mullis, K.B. and Faloona, F.A. 1987. Specific synthesis of DNA in vitro via a polymerase catalysed reaction. Meth. Enzymol. 255: 335–350.
Saiki, R.K., Gelfand, D.H., Stoffel, S., Scharf, S.J., Higuchi, R., Horn, G.T. and Erlich, H.A. 1988. Primer-directed enzymatic amplification of DNA with a thermostable DNA polymerase. Science 239: 487–491.
Ochman, H., Gerber, A.S. and Hartl, D.L. 1988. Genetic applications of an inverse polymerase chain reaction. Genetics 120: 621–623.
Doyle, J.J. and Beachy, R.N. 1985. Ribosomal gene variation in soybean (Glycine) and its relatives. Theor. Appl. Genet. 70: 369–376.
Doyle, J.J. 1988. 5S ribosomal gene variation in the soybean and its progenitor. Theor. Appl. Genet. 75: 621–624.
Apuya, N., Frazier, B., Keim, P., Roth, E.J. and Lark, K.J. 1988. Restriction length polymorphisms as genetic markers in soybean, Glycine max (L.) Merr. Theor. Appl. Genet. 75: 889–901.
Keim, P., Shoemaker, R.C. and Palmer, R.G. 1989. Restriction fragment length polymorphism diversity in soybean. Theor. Appl. Genet. 77: 786–792.
Botstein, D., White, R.L., Skolnick, M. and Davis, R.W. 1980. Construction of a genetic linkage map in man using restriction fragment length polymorphisms. Am. J. Hum. Genet. 32: 314–331.
Chesney, R.H., Scott, J.R. and Vapnek, D. 1979. Integration of the plasmid prophages P1 and P7 into the chromosome of Escherichia coli. J. Mol. Biol. 130: 161–173.
Dellaporta, S.L., Wood, J. and Hicks, J.B. 1983. A plant DNA minipreparation: Version II. Plant Mol. Biol. Reporter 1: 19–21.
Sarkar, G. and Sommer, S.S. 1990. Avoidance of false positives. Nature 343: 27.
Goldman, D. and Merril, C.R. 1982. Silver staining of DNA in polyacrylamide gels: linearity and effect of fragment size. Electrophoresis 3: 24–26.
Blum, H., Beier, H. and Gross, H.J. 1987. Improved silver staining of plant proteins, RNA and DNA in polyacrylamide gels. Electrophoresis 8: 93–99.
Bassam, B.J., Caetano-Anollés, G. and Gresshoff, P.M. 1991. Fast and sensitive silver staining of DNA in polyacrylamide gels. Anal. Biochem. In press.
Franssen, H.J., Nap, J.P., Gloudemans, T., Stiekema, W., van Dam, H., Govers, F., Louwerse, J., van Kammen, A. and Bisseling, T. 1987. Characterization of cDNA for nodulin-75 of soybean: a gene product involved in early stages of root nodule development. Proc. Natl. Acad. Sci. USA 84: 4495–4499.
Dickstein, R., Bisseling, T., Reinhold, V.N. and Ausubel, F.M. 1988. Expression of nodule-specific genes in alfalfa root nodules blocked at an early stage of development. Genes Dev. 2: 677–687.
Scheres, B., Van De Wiel, C., Zalensky, A., Horvath, B., Spaink, H., Van Eck, H., Zwartkruis, F., Wolters, A.M., Gloudemans, T., Van Kammen, A. and Bisseling, T. 1990. The ENOD12 gene product is involved in the infection process during the pest-Rhizobium interaction. Cell 60: 281–294.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Caetano-Anollés, G., Brant J., B. & Peter M., G. DNA Amplification Fingerprinting Using Very Short Arbitrary Oligonucleotide Primers. Nat Biotechnol 9, 553–557 (1991). https://doi.org/10.1038/nbt0691-553
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nbt0691-553
This article is cited by
-
Genetic diversity analysis in chrysanthemum (Dendranthema grandiflora Tzvelev) using SSR markers: corroborating mutant behaviour of newly evolved genotypes
Genetic Resources and Crop Evolution (2023)
-
Non-specificity of sequence characterised amplified region as an alternative molecular epidemiology marker for the identification of Salmonella enterica subspecies enterica serovar Typhi
BMC Research Notes (2018)