Abstract
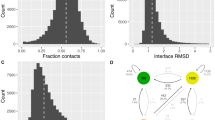
Immunoglobulin (Ig) amino acid sequences are highly conserved and often have sequence homology ranging from 70 to 95%. Antigen binding fragments (Fab), variable region fragments (Fv), and single chain Fv (scFv) of more than 50 myeloma proteins and monoclonal antibodies (mAb) have been crystallized and display a high degree of structural similarity. Based on this observation, several homology modeling approaches have been developed for the prediction of Fab and Fv structures prior to their experimental determination. We have extracted features from existing Ig sequences, 44 known Fab and Fv structures to create an automated AntiBody structure GENeration (ABGEN) algorithm for obtaining structural models of antibody fragments. ABGEN utilizes a homology based scaffolding technique, and includes the use of invariant and strictly conserved residues, structural motifs of known Fab, canonical features of hypervari-able loops, torsional constraints for residue replacements and key inter-residue interactions. The validity of the ABGEN algorithm has been tested using a five-fold cross validation with the existing Fab structures. Molecular mechanics and dynamics methods have been implemented with ABGEN models to accurately predict two Fab structures of anti-sweetener antibodies prior to crystallographic determinations.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Padlan, E.A. 1994. Anatomy of the antibody molecule. Molec. Immunol. 31: 169–217.
Kabat, E.A., Wu, T.T. and Bilofsky, H. 1977. Unusual distribution of amino acids in complementarity-determining (hypervariable) segments of heavy and light chains of immune-globulins and their possible roles in specificity of antibody-combining sites. J. Blol. Chem. 252: 6609–6616.
Kabat, E.A., Wu, T.T., Perry, H.M., Gottesman, K.S. and Foeller, C. 1991. Sequences of Proteins of Immunological Interest(Fifth Edition). US Dept. HHS NIH Publication 91-3242.
Chothia, C. and Lesk, A.M. 1987. Canonical structures for the hypervariable regions of immunoglobulins. J. Mol. Biol. 196: 901–917.
Chothia, C., Lesk, A.M., Tramontano, A., Levitt, M., Smith-Gill, S.J., Air, G., Sheriff, S., Padlan, E.A., Davies, D., Tulip, W.R., Colman, P.M., Spinelli, S., Alzari, P.M. and Poljak, R.J. 1989. Conformations of immunoglobulin hypervariable regions. Nature 342: 877–883.
de la Paz, P., Sutton, B., Darsley, M. and Rees, A. 1986. Modeling of the combining sites of three anti-lysozyme monoclonal antibodies and of the complex between one of the antibodies and its epitope. EMBO J. 5: 415–425.
Rees, A.R. and de la Paz, P. 1986. Investigating antibody specificity using computer graphics and protein engineering. Trends in Biological Science 11: 144–148.
Martin, A., Cheetham, J. and Rees, A. 1989. Modeling antibody hypervariable loops: A combined algorithm. Proc. Natl. Acad. Sci. USA 86: 9268–9277.
Bernstein, F.C., Koetzle, T.F., Williams, E.J.B., Meyer, E.F.J., Kennard, O., Shimanouchi, T. and Tasumi, M., 1977. Bank: A computer-based archival file for molecular structures. J. Molec. Biol. 112: 535–542.
Anchin, J.M. and Linthicum, D.S. 1993. Variable region sequence and characterization of monoclonal antibodies to a N,N′,N′′-trisubstituted guanidine high potency sweetener. Molec. Immunol. 30: 1463–1471.
Thornton, J.M. 1991. Modeling antibody combining sites, p. 55–71; in Catalytic Antibodies. Chichester(Ciba Found Symp 159) Wiley, NY.
Brunger, A.T., Kuriyan, J. and Karplus, M.A., 1987. R-factor refinement by molecular dynamics. Science 235: 458–460.
Hermans, J., Berendsen, H.J.C., van Gunsteren, W.F. and Postma, J.P.M. 1984. A consistent empirical potential for water-protein interactions. Biopolymers 23: 1513–1521.
Viswanathan, M., Anchin, J.A., Droupadi, P.R., Mandal, C., Linthicum, D.S. and Subramaniam, S. 1995. Structural predictions of the binding site architecture for monoclonal antibody NC6.8 using ligand binding, spectroscopy and computer-aided molecular modeling. Biophysical! 69: 741–753.
Purvis, G.D. and Culberson, J.C. 1986. On the graphical display of molecular electrostatic force-fields and gradients of the density. J. Molec. Graphics 4: 88–92.
Needleman, S.B. and Wunsch, C.D. 1970. A general method applicable to the search for similarities in the amino acid sequences of two proteins. J. Molec. Biol. 48: 443–53.
Smith, T.F. and Waterman, M.S. 1981. Identification of molecular subsequences. J. Molec. Biol. 147: 195–197.
Altschul, S.F., Gish, W., Miller, W., Myers, E.W. and Lipman, D.J. 1990. Basic local alignment search tool. J. Molec. Biol. 215: 403–410.
Bruccoleri, R., Haber, E. and Novotny, J. 1988. Structure of antibody hypervariable loops reproduced by a conformational search algorithm. Nature 335: 564–568.
Kingery, B.D., Culberson, C. and Linthicum, D.S., 1996. Automated antibody model generation for X. Biotechnol. Software J. 12: 20–28.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Mandal, C., Kingery, B., Anchin, J. et al. ABGEN: A Knowledge-Based Automated Approach for Antibody Structure Modeling. Nat Biotechnol 14, 323–328 (1996). https://doi.org/10.1038/nbt0396-323
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nbt0396-323
This article is cited by
-
Detection and characterization of a sialoglycosylated bacterial ABC-type phosphate transporter protein from patients with visceral leishmaniasis
Glycoconjugate Journal (2009)
-
Search for fucose binding domains in recently sequenced hypothetical proteins using molecular modeling techniques and structural analysis
Glycoconjugate Journal (2006)
-
Structure-based design of immunologically active therapeutic peptides
Immunologic Research (1998)