Abstract
Ash trees (genus Fraxinus, family Oleaceae) are widespread throughout the Northern Hemisphere, but are being devastated in Europe by the fungus Hymenoscyphus fraxineus, causing ash dieback, and in North America by the herbivorous beetle Agrilus planipennis1,2. Here we sequence the genome of a low-heterozygosity Fraxinus excelsior tree from Gloucestershire, UK, annotating 38,852 protein-coding genes of which 25% appear ash specific when compared with the genomes of ten other plant species. Analyses of paralogous genes suggest a whole-genome duplication shared with olive (Olea europaea, Oleaceae). We also re-sequence 37 F. excelsior trees from Europe, finding evidence for apparent long-term decline in effective population size. Using our reference sequence, we re-analyse association transcriptomic data3, yielding improved markers for reduced susceptibility to ash dieback. Surveys of these markers in British populations suggest that reduced susceptibility to ash dieback may be more widespread in Great Britain than in Denmark. We also present evidence that susceptibility of trees to H. fraxineus is associated with their iridoid glycoside levels. This rapid, integrated, multidisciplinary research response to an emerging health threat in a non-model organism opens the way for mitigation of the epidemic.
Similar content being viewed by others
Main
We sequenced a European ash (F. excelsior) tree generated from self-pollination of a woodland tree in Gloucestershire, UK. The sequenced tree (Earth Trust accession number 2451S) appeared free of ash dieback (ADB) when sampled in 2013 and 2014, but showed symptoms in February 2016. The haploid genome size was measured by flow cytometry as 877.24 ± 1.41 megabase pairs (Mbp). Total genomic DNA was sequenced to 192× coverage (see Supplementary Table 1). We assembled the genome into 89,514 nuclear scaffolds with an N50 (the length at which scaffolds include half the bases of the assembly) of 104 kilobase pairs (kbp), 26 mitochondrial scaffolds, and one plastid chromosome (Supplementary Tables 2 and 3), where the non-N assembly constitutes 80.5% of the predicted genome size. RepeatMasker estimated 35.90% of the assembly to be repetitive elements, with long terminal repeat retrotransposons predominating (Supplementary Table 4). Compared with other eudicot genomes of similar size4,5 this repeat content is low. The 17% of the assembly composed of undetermined bases probably contains additional repeats; 27% of reads that do not map to the assembly align to ash repeats (Supplementary Table 5). We generated approximately 160 million RNA sequencing (RNA-seq) read pairs from tree 2451S leaf tissue and from leaf, cambium, root and flower tissue of its parent tree (Supplementary Table 6); low expression of repetitive elements was found in all tissues (Supplementary Table 7).
We annotated the genome using an evidence-based workflow incorporating protein and RNA-seq data, predicting 38,852 protein-coding genes and 50,743 transcripts (Supplementary Table 4). This gene count is within 12% that of tomato (version of genome (v)2.3)4, potato (v3.4)6 and hot pepper (v1.5)7 but higher than monkey flower (v2.0; 26,718 genes)8. Evidence for completeness and coherence of our models is shown in Extended Data Fig. 1. Of 38,852 predicted genes, 97.67% (and 98.18% of transcripts) were supported by ash RNA-seq data, 81.80% showed high similarity to plant proteins (>50% high-scoring segment pair coverage) (Supplementary Table 8), 97.05% had matches in the non-redundant databases (excluding hits to ash), 82.74% generated hits to InterPro signatures and 78.09% were assigned Gene Ontology terms. We also identified 107 microRNA (miRNA), 792 transfer RNA (tRNA) and 51 ribosomal RNA (rRNA) genes.
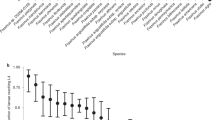
Past whole-genome duplication events are commonly inferred from the distributions of pairwise synonymous site divergence (Ks) within paralogous gene groups9. We plotted these for ash and six other plant species (Fig. 1a and Supplementary Table 9). Ash and olive shared a peak near Ks = 0.25, suggesting an Oleaceae-specific whole-genome duplication. A peak near Ks = 0.6 shared by ash, olive, monkey flower and tomato but not by bladderwort, coffee and grape does not fit a common origin hypothesis, unless bladderwort has an accelerated substitution rate and the tomato peak is not restricted to the Solanales as evidenced previously4. Synteny analysis between ash and monkey flower did not provide conclusive evidence for shared whole-genome duplication (Extended Data Fig. 2). Duplicated genes in the ash genome that were not locally duplicated (that is, within ten genes of each other in our assembly) show no significantly enriched Gene Ontology terms at a false discovery rate level of 0.05. By contrast 1,005 locally duplicated genes showed significant enrichment of terms relating to oxidoreductase, catalytic and monooxygenase activity compared with all other genes, suggesting evolution of secondary metabolism by local duplications.
a, Distribution of Ks values between paralogous gene pairs within the genomes of ash (F. excelsior), tomato (Solanum lycopersicum), coffee (Coffea canephora), bladderwort (Utricularia gibba), grape (Vitis vinifera) and monkey flower (Mimulus guttatus), and transcriptome of olive (O. europaea). b, Venn diagram of gene sharing by five asterid species. c, Venn diagram of gene sharing by six woody species. Numbers in parentheses are the total number of OrthoMCL groups found for that species; numbers in intersections show the total number of groups shared between given combinations of taxa.
We analysed gene families shared between ash and 10 other species (Supplementary Table 10). In total, 279,603 proteins (77.14% of the input sequences) clustered into 27,222 groups, of which 4,292 contained sequences from all species, 3,266 were angiosperm-specific and 462 Eudicot-specific. Patterns of gene-family sharing among asterids and among woody species are shown in Fig. 1b, c. For 38,852 ash proteins, 30,802 clustered into 14,099 groups, of which 643 were ash-specific, containing 1,554 proteins. There were also 8,050 singleton proteins unique to ash. Of the 9,604 ash-specific proteins, 6,405 matched at least one InterPro signature. The 20 largest groups in ash are listed in Extended Data Table 1: several are putatively associated with disease resistance.
To investigate genomic diversity in F. excelsior, we sequenced 37 ash trees from central, northern and western Europe (Fig. 2 and Supplementary Table 11), to an average of 8.4× genome coverage by trimmed and filtered reads. Together with reads from Danish ‘Tree35’ (http://oadb.tsl.ac.uk/), these were mapped to the reference genome. We found 12.48 million polymorphic sites with a variant of high confidence in at least one individual (quality > 300 using freebayes10): we refer to these as the ‘genome-wide SNP set’ in the ‘European Diversity Panel’. Of these, 6.85 million (54.88%) occur inside or within 5 kbp of genes (Supplementary Table 12). We found 259,946 amino-acid substitutions and 71,513 variants that affect stop or start codons, or splice sites. We selected 23 amino-acid variants, and 26 non-coding variants from the ‘genome-wide SNP set’ with a range of call qualities for validation using KASP: individual genotype calls with quality greater than 300 have a false-positive rate of 6% and those with quality greater than 1,000 have a false-positive rate of zero (Supplementary Table 13). We ran a more stringent variant calling restricted to regions of the genome with between 5× and 30× coverage in all 38 samples. These totalled 20.6 Mbp (2.3% of the genome), within which 529,812 variants were called with CLC Genomics Workbench. Of these, 394,885 were bi-allelic single nucleotide polymorphisms (SNPs) with minimum allele frequency above 0.05, which we refer to as the ‘reduced SNP set’. We also found about 31,300 singleton simple sequence repeat (SSR) loci in the ash genome, and designed primers for 664 (Supplementary Data 1). In a sample of 366 of these, 48% were polymorphic in the European Diversity Panel sequences. We PCR tested 48 of these in multiplexes with European Diversity Panel genomic DNA and found that 41 amplified successfully (Supplementary Data 1).
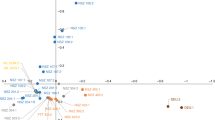
a, Map showing the distribution of plastid haplotypes (n = 37), on the basis of a median-joining plastid haplotype network for the European Diversity Panel (inset). b, Map showing diversity structure of genomic SNPs, on the basis of average Q value for each individual (inset), from three runs of STRUCTURE with different sets of 8,955 SNPs and k = 3. c, PCA of 34,607 nuclear SNPs in the European Diversity Panel, PC2 plotted against PC3, with points coloured by plastid haplotype. d, From the same PCA, PC1 plotted against PC2, with points coloured by groupings found by STRUCTURE using genomic SNPs. e, Linkage disequilibrium decay between SNPs in the European Diversity Panel. f, Effective population size (ne) history estimated using the PSMC method on the reference genome, with 100 bootstraps (shown in light blue).
We analysed population structure of the European Diversity Panel using a plastid haplotype network; STRUCTURE11 runs on genomic SNPs and principal component analysis (PCA) of the ‘reduced SNP set’ (Fig. 2a–d and Extended Data Fig. 3). Clearest differentiation was found in the plastid network, with four distinct haplotype groups each separated from each other by at least 20 substitutions. One group was more frequent in Great Britain than on the continental Europe. The second and third principal components of the PCA corresponded with the plastid data somewhat (Fig. 2c). Previous analyses of SSRs in plastids identified variants unique to the British Isles and Iberia12. Linkage disequilibrium in the European Diversity Panel decayed logarithmically, with an average r2 of 0.15 at 100 bp between SNPs, reaching an r2 of 0.05 at ~40 kbp (Fig. 2e). This is similar to long-range linkage disequilibrium estimates found in Populus tremuloides13. An apparent long-term effective population size decline of F. excelsior in Europe was shown by analyses based on heterozygosity in the reference genome (using pairwise sequentially Markovian coalescent (PSMC)14, Fig. 2f). Such patterns may also reflect a complex history of population subdivision in ash15.
We used associative transcriptomics to predict ADB damage in Great Britain. We used the full coding DNA sequence (CDS) models from our genome annotation as a mapping reference for previously generated3 RNA-seq reads from 182 Danish ash accessions (‘Danish Scored Panel’) that have been exposed to H. fraxineus, and scored for damage (Supplementary Data 2). This yielded 40,133 gene expression markers (GEMs; Supplementary Data 3) and 394,006 SNPs (Supplementary Data 4). Twenty GEMs were associated with ADB damage scores, including eight MADS-box proteins, and two cinnamoyl-CoA reductase 2 genes that may be involved in the hypersensitive response (Supplementary Data 5). Four assays representing the top five GEMs were applied to 58 Danish accessions (‘Danish Test Panel’) to validate the top markers. Results were combined into a single predicted damage score for each tree (Supplementary Data 6), which was compared with the observed damage scores (Fig. 3; r2 = 0.25, P = 6.9 × 10−5): predictions of damage less than 50% consistently detected trees with very low observed damage scores. The same assays were also applied to 130 accessions from across the British range of F. excelsior (‘British Screening Panel’; Supplementary Data 6). Strikingly, this provided lower predictions for ADB damage in the British Screening Panel: 25% were predicted to have <25% canopy damage compared with 9% of the Danish Test Panel. Trees with low predicted damage are scattered throughout Britain (Fig. 3).
Map points are scaled by hue (high predicted damage scores in brown, low in green) and plotted according to the geographical origin of the parent trees of the British Screening Panel (n = 130) and the Danish Test Panel (n = 58). Single leaf samples taken from grafts of each individual tree were used for predicting damage scores. Inset: damage predictions for the Danish Test Panel (n = 58) correlated with log(mean observed damage scores) from 2013 to 2014 (r2 = 0.25, P = 6.9 × 10−5).
We also examined expression of the top five GEM loci using reads per kilobase pair per million aligned reads (RPKM) values from our shotgun Illumina read data for the reference tree (Extended Data Fig. 4), comparing these with RPKM values from the Danish Scoring Panel. Expression patterns in the reference tree were highly correlated with those of the most susceptible Danish quartile (r2 = 0.995, P < 0.001), but not the least susceptible (P = 0.24), consistent with observations that the reference tree is now succumbing to the disease. We correlated the expression of all 20 top GEM markers in leaf, flower, cambium and root transcriptomes of the parent of the reference tree. This revealed that leaf expression levels were positively correlated with those in the cambium (r2 = 0.65, P < 0.001) and flower (r2 = 0.38, P = 0.0041), but not with the root (P = 0.3594).
We identified putative orthologues of the five GEM loci using our OrthoMCL results (Supplementary Data 5) and BLAST searches of GenBank, and conducted maximum likelihood and Bayesian analyses of relevant hits (Extended Data Fig. 5). FRAEX38873_v2_000173540.4, FRAEX38873_v2_000048340.1 and FRAEX38873_v2_000048360.1 clustered into the SVP/StMADS11 group16 of type II MADS-box genes. FRAEX38873_v2_000261470.1 and FRAEX38873_v2_000199610.1 clustered into the SOC1/TM3 group of type II MADS-box proteins16,17. Both groups have roles in flower development18,19,20,21, and appear to be involved in stress response in Brassica rapa22. Many genes involved in regulation of flowering time in Arabidopsis thaliana are involved in controlling phenology in perennial trees species23, and genes belonging to the SVP/StMADS11 clade have potential roles in growth cessation, bud set and dormancy23. In A. thaliana, AGL22/SVP may be required for age-related resistance24.
One mechanism by which transcriptional cascades, such as those involving MADS box genes, might be involved in tolerance or resistance to pathogens is via modulation of secondary metabolite concentrations. For five high-susceptibility and five low-susceptibility Danish trees, we profiled methanol-extracted leaf samples by liquid chromatography/mass spectrometry on a quadrupole time-of-flight mass spectrometer. Partial least squares discriminant analysis (PLS-DA) clearly discriminated high- and low-susceptibility trees (Fig. 4a). By using accurate mass to identify the chemical nature of discriminant features, we found greater abundance (Fig. 4b) of iridoid glycosides (for details see Extended Data Figs 6, 7, 8, 9 and Supplementary Data 9) in genotypes with high susceptibility to ADB than in low-susceptibility genotypes. A tandem mass spectrometry (MS/MS) fragmentation network identified several product ions expected from fragmentation of iridoid glycosides (Fig. 4c). Iridoid glycosides are a well-known anti-herbivore defence mechanism in the Oleaceae25,26,27. They can also enhance fungal growth in vitro28, although their aglycone hydrolysis product formed following tissue damage can also mediate fungal resistance29. Our data suggest there may be a trade-off between ADB susceptibility and herbivore susceptibility. This is of particular concern given the threat of A. planipennis to ash in both North America1 and Europe30 and may hamper efforts to breed trees with low susceptibility to both threats.
a, Multivariate analysis PLS-DA score plot of metabolic profiles of five high-susceptibility and five low-susceptibility trees (n = 3 per genotype). b, Box-plots from these profiles showing normalized (internal standard) intensity (log2 transformed) of five discriminatory features observed in negative mode; m/z and retention time (RT) are given for each feature. c, Fragmentation network of discriminatory features, highlighted in black (positive mode) and grey (negative mode). Each product ion is labelled with its size (m/z), also depicted by its circle size. Blue shading increases with the number of times each ion is present in the precursor discriminatory features. Product ions not shared among precursors are shown as unlabelled tips. The edges are in shades of red on the basis of retention time; the paler the colour the earlier the retention time. Those fragment masses shaded in green have been previously reported from fragmentation of iridoid glycosides.
Methods
No statistical methods were used to predetermine sample size. The experiments were not randomized. The investigators were not blinded to allocation during experiments and outcome assessment.
Tree material
Reference tree. In 2013 twig material was collected from tree 2451S growing at Paradise Wood, Earth Trust, Oxfordshire, UK. This tree was produced via self-pollination of a hermaphroditic F. excelsior tree growing in woodland in Gloucestershire (latitude 52.020592, longitude −1.832804), UK, in 2002 as part of the FRAXIGEN project31. The parent tree was one of 19 trees that produced seed from self-pollination, and had lower heterozygosity at four microsatellite loci than the other 18 trees (D.B., unpublished observations). DNA was extracted from bud, cambial and wood tissues using CTAB32 and Qiagen DNeasy protocols. RNA was extracted using the Qiagen RNeasy protocol from leaf tissue of tree 2451S and from leaf, cambium, root, and flower tissue of its parent tree in Gloucestershire.
European Diversity Panel. In 2014, twig material was collected from 37 trees representing 37 European provenances in a trial of F. excelsior established in 2004 at Paradise Wood, Earth Trust, Oxfordshire, UK, as part of the Realizing Ash’s Potential project. DNA was extracted from cambial tissue of the twigs using a CTAB protocol.
British Screening Panel. In 2015, freshly flushed leaf material was collected from a clonal seed orchard of F. excelsior growing at Paradise Wood, Earth Trust, Oxfordshire, UK, for RNA extraction and complementary DNA (cDNA) synthesis as in ref. 3. Single whole leaves were harvested from four ramets of each of 130 ash trees selected from phenotypically superior parents throughout Britain, which had been cloned by grafting.
2451S DNA sequencing and genome assembly
The genome size of 2451S was estimated by flow cytometry with propidium iodide staining of nuclei, using leaf tissue co-chopped with an internal standard using a razor blade. Three preparations were made: two with Petroselinum crispum ‘Curled Moss’ parsley as standard (2C genome size = 4.50 pg)33 and one with S. lycopersicum ‘Stupicke polnı rane’ (2C = 1.96 pg)34 as standard. The Partec CyStain Absolut P protocol was used (Partec, Germany). Each preparation was measured six times, with the relative fluorescence of over 5,000 particles per replicate recorded on a Partec Cyflow SL3 (Partec, Germany) flow cytometer fitted with a 100-mW green solid state laser (Cobolt Samba; Cobolt, Sweden). The resulting histograms were analysed with the Flow-Max software (version 2.4, Partec). The measurement with the tomato internal standard was used as the best estimate of genome size, because the tomato genome size is closest to that of 2451S, yielding a more accurate result.
Genomic DNA of 2451S was sequenced using the following methods: (1) HiSeq 2000 (Illumina, San Diego, California, USA) at Eurofins, Ebersberg, Germany, with 100 bp reads and shotgun libraries with fragment sizes of 200 bp, 300 bp and 500 bp, and long jumping distance libraries with 3 kbp, 8 kbp, 20 kbp and 40 kbp insert sizes, generating 188× genome coverage; (2) 454 FLX+ (Roche, Switzerland) at Eurofins with shotgun libraries and maximum read length of 1,763 bp and mean length of 642 bp giving 4.3× genome coverage; and (3) MiSeq (Illumina, San Diego, California) at the Earlham Institute, Norwich, UK, with 300 bp paired-end reads from a Nextera library with ~5 kbp insert size, giving 16× genome coverage (see Supplementary Table 1). We assembled and released five genome assembly versions over the course of 3 years, details of which can be found in Supplementary Table 3. The most recent version assembled first into 235,463 contigs with a total size of 663 Mbp and an N50 of 5.7 kbp (Supplementary Table 2), and after scaffolding and removing organellar scaffolds, the assembly comprised 89,487 scaffolds totalling 867 Mbp (17% ‘N’) with an N50 of 104 kbp (Supplementary Table 2). The plastid genome was assembled separately into one circular contig of 155,498 bp, including an inverted repeat region of approximately 25,700 bp. The mitochondrial genome initially assembled into 296 contigs totalling 232 kbp. After several rounds of contig extension using overlaps of mapped 454 reads, the final assembly consisted of 26 contigs totalling 581 kbp with an N50 of 60.6 kbp.
All Illumina reads from 2451S were trimmed using CLC Genomics Workbench (QIAGEN Aarhus, Denmark) versions 6–8 (depending on when the data were received) to a minimum quality score of 0.01 (equivalent to Phred quality score of 20), a minimum length of 50 bp, and were trimmed of any adaptor and repetitive telomere sequences. The MiSeq Nextera reads were also run through FLASH35 to merge overlapping paired reads, and NextClip36 to remove adaptor sequences, both used with default parameters. Roche 454 reads were trimmed to a minimum Phred score of 0.05, and minimum length of 50 bp. De novo assembly was performed with the CLC Genomics Workbench, using the 200 bp, 300 bp, 500 bp and 5 kbp insert size Illumina library reads to build the De Bruijn graphs. The remaining Illumina reads and the 454 reads were used as ‘guidance only reads’ to help select the most supported path through the De Bruijn graphs. A word size (k-mer, a substring of length k in DNA sequence data) of 50 and maximum bubble size of 5,000 were used to assemble the reads into contigs with a minimum length of 500 bp. Contigs were then scaffolded with the stand-alone tool SSPACE37 Basic version 2.0 using all paired Illumina reads, with the ‘-k’ parameter (number of mapped paired reads required to join contigs) set to 7. Gaps in the scaffolds were closed using the GapCloser version 1.12 program using all paired reads (except for long jumping distance libraries), with pair_num_cutoff parameter set at 7. Four hundred and fifty-four reads were mapped to the assembly and used to join overlapping scaffolds using the Jelly.py script from PBSuite38 version 14.7.14 with the following blasr parameters: -minMatch 11 -minPctIdentity 70 -bestn 1 -nCandidates 10 -maxScore -500 -noSplitSubreads. Contig57544 was removed from the assembly because it aligned fully to the PhiX bacteriophage genome, indicating it derived from the PhiX control library added to Illumina sequencing runs.
To assemble the plastid and mitochondrial genomes, high read depth 50 bp k-mers were extracted from the 200, 300 and 500 bp read libraries. Jellyfish39 version 2.1.1 was used to count the depth for each k-mer, and these values were plotted in a scatterplot to identify peaks that could correspond to the organellar genomes. Every k-mer over 600× coverage was used in a BLAST search against the NCBI non-redundant (nr) database with a filter allowing only plant sequences; k-mers were then extracted on the basis of whether their first hit contained a ‘mitochondrion’ or ‘plastid/chloroplast’ related description. Reads from the 200, 300 and 500 bp libraries were then filtered against the k-mer sets, and were kept if the first and last 50 bp matched k-mers from the extracted sets (reads were at most 90 bp long). Each set of reads (mitochondrial and plastid) were then assembled de novo using the CLC Genomics Workbench. The plastid genome assembled initially into two contigs, which were joined using an alignment to the O. europaea plastid genome (GenBank accession number NC_015401.1), with the inverted repeat region being identified also. Reads from the 454 library were mapped to the assembly to check the sequence and especially the join region. The mitochondrial genome assembled first into 296 contigs. To fill in gaps and join the contigs together, 454 reads were mapped against the assembly and contig ends were extended using the Extend Contigs tool in the CLC Genome Finishing Module. The Join Contigs tool was then used to join overlapping ends together, and 454 reads were mapped to the resulting assembly to check any joined regions. Using this method of ‘Map-Extend-Join’ iteratively (approximately ten times in total), a more contiguous assembly of 26 contigs was obtained.
RNA sequencing
The five RNA samples (see ‘Tree Material’ above) were sequenced paired-end on Illumina HiSeq 2000 with 200 bp insert sizes, and a read length of 100 bp, at the QMUL Genome Centre, London, UK. Reads were trimmed using CLC Genomics Workbench to a minimum quality score of 0.01 (equivalent to Phred score of 20) and minimum length of 50 bp, and adaptors were also removed (Supplementary Table 6).
Analysis of repetitive DNA
The repetitive element (transposable elements and tandem repeats) content of the ash genome was analysed via two approaches: (1) de novo identification of the most abundant repeat families from unassembled 454 and Illumina reads; (2) de novo and similarity-based identification of repeats from the ash genome assembly.
De novo identification of repeat families from unassembled reads
Individual 454 reads and Illumina read pairs from the 500 bp insert library (after adaptor trimming, but before any further quality control or filtering: see above) were used for de novo repeat identification. Reads were quality filtered and trimmed using the FASTX-Toolkit version 0.0.13 (http://hannonlab.cshl.edu/fastx_toolkit/index.html). Using fastx_trimmer, the first 10 bp of all reads (454 and Illumina) were removed (owing to skewed base composition). The 454 reads were clipped to a maximum of 250 bp and Illumina reads to a maximum of 90 bp; all shorter reads were removed using a custom Perl script. Reads were then quality filtered with the fastq_quality_filter tool to retain only those where 90% of bases had a Phred score of at least 20. Exact duplicates (which are probably artefacts from the emulsion PCR40) were removed from the 454 reads using the fastx_collapser tool.
The complete set of quality filtered and trimmed 454 reads (3,330,483) was used as input for the RepeatExplorer pipeline on Galaxy41, with a minimum of 138 bp overlap for clustering and a minimum of 100 bp overlap for assembly. All clusters containing at least 0.01% of the input reads were examined manually to identify clusters that required merging (that is, where there was evidence that a single repeat family had been split over multiple clusters). Clusters were merged if they met the following three criteria: (1) they shared a significant number of similarity hits (for example, in a pair of clusters, 10% of the reads in the smaller cluster had BLAST hits to reads in the larger cluster); (2) they were the same repeat type (for example, LINEs); (3) they could be merged in a logical position (for example, for repetitive elements containing conserved domains these domains would be joined in the correct order). The re-clustering pipeline was run with a minimum of 100 bp overlap for assembly; merged clusters were examined manually to verify that all domains were in the correct orientation.
Quality filtered and trimmed Illumina reads were paired using the FASTA interlacer tool (version 1.0.0) in RepeatExplorer, resulting in 111,230,011 pairs; unpaired reads were discarded. An initial run of RepeatExplorer with a sample of 100,000 read pairs was performed to obtain an estimate of the maximum number of reads that could be handled by the pipeline. A random sample of 3.5 million read pairs was then taken using the sequence sampling tool (version 1.0.0) in RepeatExplorer and used as input for the clustering pipeline, which further randomly subsampled the reads down to 3,370,186 pairs. The pipeline was run with a minimum of 50 bp overlap for clustering and a minimum of 36 bp overlap for assembly. Clusters containing at least 0.01% of the input reads were merged if kx,y passed the 0.2 cut-off (for clusters x and y, kx,y is defined as k1,2 = 2W/(n1 + n2) where W is the number of read pairs shared between clusters x and y and nx is the number of reads in cluster x which does not include the other read from its pair within the same cluster); clusters that passed this threshold but which had no similarity hits to each other were not merged. The re-clustering pipeline was run with a minimum of 36 bp overlap for assembly.
Repeat families identified by RepeatExplorer were annotated according to the results of BLAST searches to the Viridiplantae RepeatMasker library, to a database of conserved protein coding domains from transposable elements and to a custom RepeatMasker library comprising all Fraxinus sequences (excluding shotgun sequences), all mitochondrial genome sequences from asterids and all plastid genome sequences from Oleaceae available from NCBI (downloaded on 13 February 2014); these BLAST searches were performed as part of the RepeatExplorer pipeline. For repeat families that were not annotated in RepeatExplorer (that is, no significant BLAST hits), or where only very few reads (<2%) had a BLAST hit or separate reads matched different repeat types (that is, inconsistent BLAST hits), contigs were also searched against the nr/nt database in GenBank using BLASTN with an E value cut-off42 of 1 × 10−10, against the non-redundant database using BLASTX with an E value cut-off of 1 × 10−5, and submitted to Tandem Repeat Finder version 4.07b with default parameters43. Annotation of repeat families from the clustering of the 454 and Illumina data was cross-validated by BLAST searching the contigs from each analysis against each other using the BLASTN program in the BLAST+ package (version 2.2.28+) with an E value cut-off of 1 × 10−10 and the DUST filter switched off. Any repeat families annotated as plastid or mitochondrial DNA were removed before downstream analyses (see below).
Identification of repeats from the genome assembly
De novo identification of repetitive elements from the assembled ash genome sequence was conducted with RepeatModeler version 1.0.7 (http://www.repeatmasker.org/RepeatModeler.html) using RMBlast as the search engine. All unannotated (‘unknown’) repeat families from the RepeatModeler library were searched against a custom BLAST database of organellar genomes (see above) using BLASTN with an E value cutoff of 1 × 10−10 in the BLAST+ package (version 2.2.28+ (ref. 44)). Any repeat families matching plastid or mitochondrial DNA were removed.
To prevent any captured gene fragments within repetitive element families causing the masking of protein coding genes within the ash assembly, the custom repeat libraries were pre-masked using the TAIR10 CDS data set45 (TAIR10_cds_20101214_updated; downloaded from http://www.arabidopsis.org). First, transposonPSI version 2 (http://transposonpsi.sourceforge.net) was run with the ‘nuc’ option to identify any transposable-element-related genes within the TAIR10 CDS data set. Sequences with a significant hit to transposable-element-related sequences (E value cut-off of 1 × 10−5) were removed from the TAIR10 CDS file (n = 308); a further 19 sequences that included the term ‘transposon’ in their annotation, but which did not have a hit using transposonPSI, were also removed. The filtered TAIR10 CDS data set was used to hard mask the RepeatModeler library, the RepeatExplorer libraries (454 and Illumina) and the library from RepeatMasker using RepeatMasker version 4.0.5 (http://www.repeatmasker.org) with RMblast as the search engine and the following parameter settings: -s –no_is –nolow. The four pre-masked libraries were combined into a single custom repeat library; any repeat families annotated as ‘rRNA’, ‘low-complexity’ or ‘simple’ were removed before combining the libraries. The combined library was then used to identify repetitive elements in the ash genome assembly with RepeatMasker version 4.0.5, using the same parameter settings as above. RepeatMasker results were summarized using ProcessRepeats with the species set to ‘eudicotyledons’ and using the ‘nolow’ option.
In addition to the analysis with the combined custom ash repeat library, repeats within the assembly were also annotated by running RepeatMasker separately with each of the four individual repeat libraries with parameter settings as described above. The results were saved in gff format and combined into a single gff file that was then used to inform the process of annotating protein coding genes (see below, ‘Gene annotation’).
Although the ash genome assembly covers about 99% of the expected genome size based on flow cytometry, about 17% is composed of Ns. Therefore, the repeat content of the genome assembly may be an underestimate of the actual amount of repetitive DNA within the genome. To test whether the about 18% of missing sequence includes additional repetitive elements we analysed the repeat content of individual Illumina reads that do not map to the genome assembly. Quality-trimmed and length-filtered reads from the Illumina short insert libraries (Supplementary Table 1) were mapped to the assembly using the ‘Map Reads to Reference’ tool in the CLC Genomics Workbench, with both similarity match and length match parameters set to 0.90. Unmapped reads from the 200 bp, 300 bp and 500 bp insert libraries (equating to about 4.8% of all reads from these libraries; see Supplementary Table 1) were searched against the custom library of ash repeats using BLASTN (see Supplementary Table 5) with an E value cut-off of 1 × 10−10 and the DUST filter switched off in the BLAST+ package (version 2.2.29+ (ref. 44)).
To test for evidence of the expression of transposable elements, trimmed RNA sequencing reads from five different tissue types (see Supplementary Table 7) were searched against the custom library of ash repeats using BLASTN as described above for the unmapped DNA sequencing reads.
Gene annotation
Protein coding genes were predicted using an evidence-based annotation workflow incorporating protein, cDNA and RNA-seq alignments. Protein sequences from nine species (Amborella trichopoda, A. thaliana, Fraxinus pennsylvanica, M. guttatus, Populus trichocarpa, S. lycopersicum, Solanum tuberosum, V. vinifera and Pinus taeda; Supplementary Table 8) were soft masked for low complexity (segmasker-blast-2.2.30) and aligned to the softmasked (for repeats) final 2451S assembly with exonerate46 protein2genome version 2.2.0; alignments were filtered at a minimum 60% identity and 60% coverage, except for F. pennsylvanica, which were filtered at a minimum of 80% identity and 60% coverage. Publicly available F. excelsior expressed sequence tags (12,083 from GenBank) were aligned with GMAP (r20141229)47 and filtered at a minimum 95% identity and 80% coverage.
RNA-seq reads from the five sequenced RNA samples were filtered for adaptors and quality trimmed, rRNA reads were identified and removed48 (trim_galore-0.3.3 http://www.bioinformatics.babraham.ac.uk/projects/trim_galore/: -q 20–stringency 5–length 60; sortmerna-1.9: -r 0.25–paired-out). RNA-seq reads were aligned using TopHat (version 2.0.13/Bowtie 2.2.3)49 and transcript assemblies were generated using three alternative methods: Cufflinks (version 2.2.1)50, StringTie (version 1.04)51 and Trinity (genome-guided assembly)52. Assembled Trinity transcripts were mapped to the F. excelsior assembly using GMAP (r20141229) at 80% coverage and 95% identity. A comprehensive transcriptome assembly was created using Mikado (version 0.8.5, https://github.com/lucventurini/mikado, L. Venturini, manuscript in preparation) on the basis of the GMAP Trinity alignments, Cufflinks and StringTie transcript assemblies. Mikado leverages transcript assemblies generated by multiple methods to improve transcript reconstruction. Loci are first defined across all input assemblies with each assembled transcript scored on the basis of metrics relating to open reading frame and cDNA size, relative position of the open reading frame within the transcript, untranslated region length and presence of multiple open reading frames. The best scoring transcript assembly is then returned along with additional transcripts (splice variants) compatible with the representative transcript.
Protein coding genes were predicted using AUGUSTUS53 by means of a generalized hidden markov model that took both intrinsic and extrinsic information into account. An AUGUSTUS ab initio model was generated on the basis of a subset of cufflinks assembled transcripts identified by similarity support as containing full-length open reading frames. Gene models were predicted using the trained ab initio model with the nine sets of cross species protein alignments, RNA-seq junctions (defining introns), and Mikado transcripts as evidence hints. RNA-seq read density was provided as exon hints and repeat information (interspersed repeats) as nonexonpart hints. We generated two alternative AUGUSTUS models by either including or excluding the RNA-seq read depth information. A set of integrated gene models was derived from the two AUGUSTUS runs along with the transcriptome and protein alignments via EVidenceModeller:r20120625 (EVM)54. Weights of evidence were manually set following an initial testing and review process as AUGUSTUS predictions with RNA-seq read depth hint, weight 2; AUGUSTUS predictions without RNA-seq read depth hint, weight 1; protein alignment high confidence (greater than 90% coverage, 60% identity) weight 5; protein alignment low confidence (lower than 90% coverage, 60% identity) weight 1; cufflinks transcripts, weight 1; Mikado transcripts, weight 10; RNA-seq splice junctions, weight 1. We identified examples of EVM errors resulting from incomplete genes in the AUGUSTUS gene predictions or non-canonical splicing; to rectify these problems we substituted the EVM model for the overlapping AUGUSTUS model (with RNA-seq read depth hints). To add untranslated region features and alternative splice variants we ran PASA55 with Mikado transcript assemblies and available F. excelsior expressed sequence tags using the corrected EVM models as the reference annotation.
The PASA updated EVM models were further refined by removing gene models that showed no expression support (using all available RNA-seq libraries) or had no support from cross species protein alignments or no BLAST similarity support with a Viridiplantae (without F. excelsior) protein database (<50% BLAST high-scoring segment pair coverage) or where the CDS length was less than 100 bp (retaining those transcripts with ≥50% BLAST high-scoring segment pair coverage). Gene models were also excluded if they aligned with at least 30% similarity and 40% coverage to the TransposonPSI (version 08222010) library (http://transposonpsi.sourceforge.net/) and had at least 40% coverage by the RepeatModeler/RepeatMasker derived interspersed repeats. In addition, gene models that had at least 30% similarity and 60% coverage to the TransposonPSI library or had at least 60% coverage by the RepeatModeler/RepeatMasker derived interspersed repeats were also excluded. The functional annotation of protein coding genes was generated using an in-house pipeline, AnnotF-1.01, which executes and integrates the results from InterProSCAN (version 5) and Blast2GO (version 2.5.0). Completeness of transcript models was classified by Full-lengther Next56 and coherence in gene length examined by comparison with single copy gene BLAST hits in monkey flower (Extended Data Fig. 1).
Transfer RNA genes were predicted by tRNAscanSE-1.3.1 with eukaryote parameters57 and rRNAs using RNAmmer-1.2 (ref. 58). miRNA was predicted by BLASTN searches with precursor miRNAs from miRBase59 21.0 against the reference genome sequence (BLAST 2.2.30, E value 1 × 10−6) and miRCat60 using the mature miRNAs from miRBase with default plant parameters, except modifying the flanking window to 200 bp. Putative miRNA precursors from these methods were combined and were folded using RNAfold61 and mature miRNAs from miRBase were aligned to precursor hairpins using PatMaN62. These predictions were checked manually for RNA secondary structure.
Organellar genes were annotated manually using the BLAST tool within the CLC Genomics Workbench version 7.5. Mitochondrial genes were identified using CDS from M. guttatus, Nicotiana tabacum and A. thaliana (all downloaded from NCBI). Plastid genes were identified using CDS from O. europaea and N. tabacum (both downloaded from NCBI). An E value cut-off of 1 × 10−4 was used. Gene and CDS annotations were added manually to the F. excelsior organellar scaffolds using the sequence editing tools available within the CLC Genomics Workbench. In the plastid genome, we annotated 72 protein-coding, 7 putative coding (ycf), rRNA and tRNA genes. On the mitochondrial scaffolds, we annotated 37 protein-coding, rRNA and tRNA genes.
Analysis of whole-genome duplications
To examine evidence for past whole-genome duplication, CDS and protein sequences (one transcript per gene) were taken from our ash genome annotation, and downloaded from Phytozome version 10.3 for tomato (S. lycopersicum), monkey flower (M. guttatus) and grape (V. vinifera), the CoGe database for bladderwort (U. gibba) and http://coffee-genome.org for coffee (C. canephora). For olive (O. europaea) we predicted open reading frames from transcriptome data63 using Transdecoder52 with all parameters set to defaults (version 2.01, http://transdecoder.github.io). Olive63 is in the same family as ash (Oleaceae); monkey flower8 and bladderwort64 are in the same order as ash (Lamiales); tomato4 and coffee65 are in different orders (Solanales and Gentianales, respectively), but like ash in the asterids; and grape66 is a rosid. An all-against-all comparison using protein sequences was performed on each species separately using BLASTP version 2.2.29, with an E value cut-off of 1 × 10−5. BLAST alignments were further filtered to retain pairs for which the shorter sequence was at least 50% of the longer sequence, and the alignment was at least 50% of the shorter sequence. If one sequence had multiple matches meeting the length and E value thresholds, these were grouped into a paralogue group, including any other genes that were associated with the matches (for example, if gene A matches gene B and gene C, and gene C also matches gene D, then one group of A, B, C and D would be formed).
Next, all possible pairs of protein sequences within each group were aligned using muscle version 3.8.31 with default parameters67. A nucleotide alignment was generated from the protein alignment using a Python script. Synonymous substitutions were estimated using the codeml program from PAML version 4.8 (ref. 68). The Ks scores within each group were then corrected to remove redundant values; only those representing duplication events within the group were retained (in a group of n genes, there are n − 1 possible duplication events) using the method described in refs 9 and 69. These steps are implemented in a Python script available online: http://github.com/EndymionCooper/KSPlotting.
To examine patterns of conserved synteny, we constructed syntenic dotplots using the SynMap70 with default parameters (Extended Data Fig. 2). The default uses LAST71 to perform similarity searches, and DAGchainer72 to find syntenic regions. By default DAGchainer requires a minimum of 5 aligned gene pairs with no more than 20 genes between neighbouring pairs.
Pairs of genes were categorized as ‘local’ duplications if they were located on the same chromosome or scaffold and resided within ten genes of each other, and as ‘tandem’ duplications if they reside directly next to each other. Gene Ontology term enrichment was performed on ash proteins using the BLAST2GO plugin suite of tools within the CLC Genomics Workbench version 8.5. Three separate BLAST searches were run against the RefSeq protein database: first using CDS from all genes as queries, second using CDS from genes involved in whole-genome duplication (excluding locally duplicated genes), and third using CDS from locally duplicated genes (genes located within ten genes of each other). The E value cut-off for all BLAST runs was 1 × 10−5. BLAST results were annotated with Gene Ontology terms using the ‘Mapping’ and ‘Annotation’ tools within the BLAST2GO plugin, using default parameters except for Annotation Cutoff = 55 and high-scoring segment pair-hit coverage cutoff = 40. Significantly enriched Gene Ontology terms were identified using the Fisher’s exact test tool within the plugin, where the reference set was the Gene Ontology terms for all genes, and a false discovery rate of 0.05 was used.
Analysis of gene families
The OrthoMCL pipeline (version 2.0.9)73 was used to identify clusters of orthologous and paralogous genes from F. excelsior and the following: Amborella74, Arabidopsis75, barrel medic76, bladderwort64, coffee65, grape66, loblolly pine77, monkey flower8, poplar78 and tomato4 (Supplementary Table 10). Input proteomes contained a single transcript per gene and were filtered with orthomclFilterFasta to remove any sequences of fewer than ten amino acids in length and/or >20% stop codons. Similar sequences were identified via an all versus all BLASTP search for the 362,741 proteins remaining after filtering. The BLAST search was performed in the BLAST+ package44 (version 2.2.29+), using an E value cut-off of 1 × 10−5. BLAST results were filtered with orthomclPairs to retain protein pairs that match across at least 50% of the length of the shorter sequence in the pair. Clustering of sequences was performed with mcl79 (version 14.137) using a setting of 1.5 for the inflation parameter. The output from OrthoMCL was summarized using a custom Perl script to obtain counts of the number of sequences from each species belonging to each group. Venn diagrams for selected taxa were generated using InteractiVenn80.
European Diversity Panel sequencing
DNA from the 37 European Diversity Panel trees was sequenced at the Earlham Institute on Illumina HiSeq, using paired-end insert sizes between 100 and 700 bp, and a read length of 150 bp. This generated an average of 63.6 million 150 bp reads (10.9× genome coverage) per tree. Filtering and trimming steps reduced this average to 55.3 million reads. An average of 85.8% of these reads per tree mapped to our reference genome. In addition, DNA reads from Danish Tree35 library ‘3077’ were downloaded from the Open Ash Dieback website (http://oadb.tsl.ac.uk); these were 250 bp paired-end reads with an insert size between 200 and 400 bp. Tree35 is given the sample number ‘38’ in all further population analysis.
European Diversity Panel genome-wide SNP calling
The raw reads from the 37 trees in the European Diversity Panel (Supplementary Table 11) were aligned to the reference genome using Bowtie 2.2.5 (ref. 81). The alignments were converted to BAM format and duplicated reads were removed with samtools 1.2 (ref. 82). To assign each read to its corresponding tree, the flag ‘rg’ was added to each BAM file with picard tools 1.119 (http://broadinstitute.github.io/picard/). SNPs were called with freebayes 1.0.2 (ref. 10) to produce a VCF file. The SNPs with quality less than 300 were filtered with bio-samtools 2.1 (ref. 83). SnpEff 4.1g (ref. 84) was used to predict the effect of the putative SNPs (see Supplementary Table 12). Genic regions were within 5 kbp from a gene model. Amino-acid changes were labelled as missense_variant.
SNP call validation using the KASP platform
To test the reliability of SNP calls in the genome-wide SNP calling, we designed KASP assays for 53 SNPs, which ranged in their level of confidence (see Supplementary Table 13). None of the SNP calls tested by KASP were present in the reduced SNP set used for population genetic analyses. Primers were designed with a modified version of PolyMarker85 including FAM or HEX tails (FAM tail: 5′-GAAGGTGACCAAGTTCATGCT-3′; HEX tail: 5′-GAAGGTCGGAGTCAACGGATT-3′). The primer mix was prepared as recommended by the manufacturer (46 μl distilled H2O, 30 μl common primer (100 μM) and 12 μl of each tailed primer (100 μM)) (http://www.lgcgroup.com/services/genotyping). The assays were run on 37 individuals from the European Diversity Panel, in 384-well plates as 4 μl reactions (2-μl template (10–20 ng of DNA), 1.944 μl of V4 2× Kaspar mix and 0.056 μl primer mix). PCR was done with the following protocol: hotstart at 95 °C for 15 min, followed by ten touchdown cycles (95 °C for 20 s; touchdown 65 °C, −1 °C per cycle, 25 s) then followed by 30 cycles of amplification (95 °C for 10 s; 57 °C for 60 s). Fluorescence was detected on a Tecan Safire at ambient temperature. Genotypes were called using Klustercaller software (version 2.22.0.5; LGC Hoddesdon, UK). Four of the individuals did not amplify and were discarded from the analysis. The results of the calls are in Supplementary Data 7.
European Diversity Panel population genetics and history using a reduced set of SNPs
For population structure analyses and effective population size estimation, variants were only called at SNP sites in the genome where all 38 samples had between 5× and 30× coverage. We refer to this as the ‘reduced SNP set’.
First, all reads were trimmed in the CLC Genomics Workbench to a minimum quality score of 0.01 (equivalent to Phred quality score of 20), a minimum length of 50 bp, and were also trimmed of any adaptor and repetitive telomere sequences. Filtered reads were mapped to the reference assembly using the ‘Map Reads to Reference’ tool in the CLC Genomics Workbench, setting both similarity match and length match parameters to 0.95. Regions with coverage of between 5 and 30 reads in all samples were extracted using the ‘Create Mapping Graph’, ‘Identify Graph Threshold Areas’ and ‘Calculus Track’ tools. These extracted regions totalled 20.6 Mbp (2.3% of the genome).
Variant calling was performed on a read mapping pooled from all samples, using the ‘Low Frequency Variant Caller’ tool in the CLC Genomics Workbench, with the coverage-restricted regions from the previous step used as a track of target regions. This prevented variants being called where some samples did not have read coverage, and in the organellar scaffolds where the read coverage was very high. The following parameters were changed from default: Ignore positions with coverage above = 1,000, Ignore broken pairs = no, Ignore non-specific matches = Reads, Minimum Coverage = 190 (38 samples with at least 5 reads each should have a combined total coverage of >189), Minimum Count = 10, Minimum Frequency = 5%, Base Quality Filter = Yes, Neighbourhood radius = 5, Minimum Central Quality = 20, Minimum neighbourhood quality = 15, Read Direction Filter = yes, Direction Frequency = 5%. As a result, 529,812 variants were called, comprising 468,237 SNPs, 14,850 equal replacements (where >1 nucleotide is replaced by an equal number of nucleotides), 26,043 deletions, 19,085 insertions and 1,597 unequal replacements (where at least one SNP lies directly beside an indel). The average quality of all reads at these variant positions was 36.2.
To genotype each sample individually at the variant loci called in the previous steps, the ‘Identify Known Mutations from sample mappings’ tool within the CLC Biomedical Genomics workbench was used. The workflow takes a track of known variants as input (such as those called from the pooled read mapping) and reports the presence, absence, coverage, count and other statistics of each variant locus in the read mapping of another sample (in this case, the read mapping from each of the 38 trees). The ‘Identify Candidate Variants’ tool was then used to filter variants with a minimum coverage of 5, minimum count of 3 and minimum frequency of 20%. VCF files for each tree were exported from the CLC Workbench and merged into one file using the vcf-merge tool from VCFtools86. The merged VCF file was then filtered using vcftools, to remove indels, multi-allelic loci, and loci with a minimum allele frequency < 0.05, with 394,885 SNP loci remaining. This set of high-quality SNPs with comprehensive knowledge of the genotype of every sample was referred to as the ‘reduced SNP set’ and used for further population analyses.
To visualize similarities and differences among the genomes of the European Diversity Panel, PCA was performed using the SNPRelate version 1.4.2 (ref. 87) package in R version 3.1.2. The filtered VCF file was converted into gds using the snpgdsVCF2GDS command, and was filtered on a linkage disequilibrium value of 0.1 using the snpgdsLDpruning command, leaving 34,607 SNPs. PCA was performed on the pruned set of SNPs using the snpgdsPCA command with default options, and the results of the first three PCs were plotted in R.
To analyse population structure in the European Diversity Panel, scaffolds were selected that contained 10 or more SNPs in the filtered VCF file (8,955 nuclear scaffolds in total). Three different SNPs were selected at random from each of these scaffolds, and placed into three different files in STRUCTURE input format (26,865 SNPs in total, 8,955 in each set). STRUCTURE version 2.3.4 (ref. 88) was run with admixture from k = 1 to k = 20 for each of the three sets of SNPs, with both BURNIN and NUMREPS set to 100,000. All output results were run through Structure Harvester Web version 0.6.94 (ref. 89), which found k = 3 to have the largest Δk value of 32.91 (Extended Data Fig. 3). Next, the three runs of k = 3 were used as input into CLUMPP version 1.1.2 (ref. 90) to align the clusters, and samples within each cluster. Aligned results were imported back into STRUCTURE version 2.3.4 to generate Q value bar plots. Average Q values from the three runs were used to generate a map with pie charts, using Tableau version 9.3 (Tableau, Seattle, USA) with Tableau base-map country outlines. Each section of the pie represented the average Q value of the individual belonging to the coloured cluster (Fig. 2b).
To analyse relationships among plastid sequences in plastid haplotype networks, a consensus sequence of the large single copy plastid region was extracted for each of the 38 samples. The sequences were then aligned using the Create Alignment tool in the CLC Genomics Workbench, and the alignment was exported in Phylip format. The alignment was imported into PopArt version 1.7 (http://popart.otago.ac.nz), where a Median Joining network was generated. Results were visualized on a map using Tableau version 9.3 (Fig. 2a) with Tableau base-map country outlines.
We estimated the effective population size history of F. excelsior using two complementary methods: the PSMC14 model estimated the history in the non-recent past, whereas by using linkage disequilibrium, we could estimate the population size more recently. The PSMC model calculated the effective population size using a time to most recent common ancestor approach. The effective population size history was then estimated from the number of recombination events separating segments of constant time to most recent common ancestor. The program PSMC 0.6.5 (ref. 14) took only a diploid consensus sequence as input. To estimate past effective population size, PSMC analysis was used on the reference tree. DNA reads from the 2451S 200, 300 and 500 bp libraries were mapped to the 2451S reference sequence using CLC Genomics Workbench ‘Map Reads to Reference’ tool (length fraction = 0.95 and similarity fraction = 0.9). The mapping was exported in BAM format, and a consensus sequence was obtained following PSMC recommendations, by using samtools version 0.1.18 ‘mpileup’ command with options -C 50 -A -Q 20 -u, bcftools version 1.1 to convert the BCF file to VCF format, and finally using vcfutils.pl to convert the VCF file to a consensus sequence where the coverage was between 5 and 200. The PSMC program was then run with default parameters except for -p ‘4+25*2+4+6’, with 100 bootstraps. To scale the results, the psmc_plot.pl script was used with default parameters except for the following: -u 7.5e-09 -g 15 -N 0.25 (the mutation rate of F. excelsior was unknown, so the substitution rate of 7.5 × 10−9 was taken from a study on A. thaliana91). Effective population size estimates were then plotted in R version 3.1.2 (Fig. 2f).
Effective population size estimation by linkage disequilibrium in the European Diversity Panel was performed using the program SNeP version 1.1 (ref. 92), which takes genome-wide polymorphism data from several individuals in a population as input. The European Diversity Panel filtered VCF file with the reduced SNP set of 38 trees (the same as used in PCA and STRUCTURE analysis) was converted into Map and Ped files. The third column in the Map file (linkage distance in morgans) was set to zero for all SNPs, as these values were unknown and SNeP calculates this value from each SNP’s physical distance. SNeP was then run with a minimum distance between SNPs of 10,000 bp and a maximum of 400,000 bp, with Sved’s modifier for recombination rate, and with 50 bins. Estimated effective population sizes were plotted in R (Extended Data Fig. 3c), as well as linkage disequilibrium decay over distance between 100 and 300,000 bp (Fig. 2e).
Simple-sequence repeat analysis
To develop accessible population genetic markers, the repeat masked version 0.4 2451S genome was mined for simple sequence repeat (SSR) sequences (a repeat motif of 2–5 bp in length repeated a minimum of five times) using the QDD version 3.1 pipeline93. Downstream QDD version 3.1 pipes screened SSR loci (inclusive of the SSR repeat motif and 200 bp forward and reverse flanking regions) for singleton sequences in an all-against-all BLAST (-task blastn -evalue 1e-40 -lcase_masking -soft_masking true) and designed primer pairs within 200 bp flanking regions using PRIMER3 software94.The approximately 31,300 singleton SSR loci identified in the ash genome were screened using RepeatMasker Open-4.0 (http://www.repeatmasker.org) in QDD version 3.1 to eliminate loci that hit known transposable elements in the RepBase Viridiplantae repeat library (http://www.girinst.org), leaving about 28,800 SSR loci. The final primer table output by the QDD version 3.1 pipeline allows selection of the best primer pair design for each SSR loci. To select candidate markers for further development, these primer pairs were filtered according to parameters provided by QDD version 3.1. The selected SSR loci had a: maximum primer alignment score of 5; minimum 20 bp forward and reverse flanking region between SSR and primer sequences; high-quality primer design (defined by QDD pipeline as an absence of homopolymer, nanosatellite and microsatellite sequence in primer and flanking sequences); and minimum number of 7 motif repeats within the SSR sequence. This filtering gave a set of 837 SSR loci, which was screened against the combined custom ash repeat library for v0.5 of the 2451S genome assembly (see above: ‘Analysis of repetitive DNA’) via a BLASTN search with an E value of 1 × 10−10 in the BLAST+ package (version 2.2.31+). Elimination of all sequences with a hit to known repetitive elements left 681 candidate loci. These were compared with the v0.5 assembly via a BLASTN search with an E value cut-off of 1 × 10−10. This returned a set of 664 loci with a unique match to the v0.5 assembly for use as population genetic markers (see Supplementary Data 1).
In silico analysis of allelic diversity (that is, locus polymorphism) of these SSR loci was performed by screening a subset of loci (366) against a variance table composed of insertions and deletions recorded for the European Diversity Panel. Approximately half (48%) of the loci tested were variable among 37 of the re-sequenced genomes (sample 38 not included). Twenty candidate SSR loci with the greatest in silico allelic diversity were selected for wet laboratory testing on seven individuals from the European Diversity Panel. Primer pairs with a fluorescent tag on the 5′ end of the forward primer (FAM, HEX or TAM) were used. For singleplex PCR, primer aliquots were used at a concentration of 10 pmol/μl. PCR amplification of target regions was performed in singleplex reactions with a final reaction volume of 10 μl, containing 1 μl genomic DNA, 0.2 μl of each primer (10 pmol/μl), 3.6 μl of RNase free water, and 5 μl of Qiagen Type-it Multiplex PCR Master Mix, in a G-Storm GS2 Multi Block Thermal Cycler. The amplification conditions were as follows: 5 min at 95 °C; 18 cycles of 30 s at 95 °C, 90 s at 62 °C with a 0.5 °C reduction per cycle, 30 s at 72 °C; 20 cycles of 30 s at 95 °C, 1 min 30 s at 51 °C, 30 s at 72 °C; a final extension step of 30 min at 60 °C. PCR samples were diluted to 1:10 with distilled H2O and run (on an Applied Biosystems 3730xl 96 capillary sequencing instrument with Applied Biosystems GeneScan 400HD Rox dye size standard). Negative control samples were included for each primer pair PCR reaction mix. Allele calling was performed using GeneMarker version 2.6.4 (http://www.softgenetics.com).
Primer pairs that produced interpretable allele peaks from capillary sequencing of singleplex reactions were arranged into four multiplex primer mixes (containing five primer pairs each) according to PCR product size and fluorescent tag. Multiplex primer mixes were tested on DNA extractions for a further 14 of the 37 trees from the European Diversity Panel. For each multiplex, primer pair mixes were prepared at a final concentration of 10 pmol/μl and amplified via PCR in 10 μl reaction volumes (1 μl genomic DNA, 1 μl primer mix, 3 μl of RNase free water, and 5 μl of Qiagen Type-it Multiplex PCR Master Mix) under the amplification conditions described above. PCR product size range, allele counts, primer design and successful multiplex panels for the 20 wet laboratory tested candidate SSR markers developed for European ash are described in Supplementary Data 1.
Further multiplex primer mixes were tested on 7 trees from the European Diversity Panel for amplification of the longest SSR loci (14 or more repeated motifs). Primer pair mixes were prepared at a final concentration of 10 pmol/μl and amplified via PCR in 8 μl reaction volumes (1 μl genomic DNA from a 1:10 dilution with nuclease free water, 1 μl primer mix, 2 μl of RNase free water, and 4 μl of Qiagen Type-it Multiplex PCR Master Mix.). The amplification conditions were as follows: 5 min at 95 °C; 32 cycles of 30 s at 95 °C, 90 s at 62 °C with a 0.35 °C reduction per cycle, 30 s at 72 °C; a final extension step of 30 min at 60 °C. Amplification was performed in a G-Storm GS2 Multi Block Thermal Cycler. Size fraction analysis of PCR products was performed for two samples of each tested primer multiplex using a 12 sample DNA1000/7500 chip in an Agilent 2100 Bioanalyzer (http://www.genomics.agilent.com). Of the 28 primer pairs tested, 22 successfully amplified across the six primer multiplexes tested (Supplementary Data 1).
Association of transcriptomic markers with reduced susceptibility to ADB in Denmark
Sequence reads for the ‘Danish Scored Panel’ of 182 Danish ash accessions (as described in ref. 3; sequence reads are available in the European Nucleotide Archive under the study accession number PRJEB10202) were mapped to a reference composed of the complete set of CDS models (including 229 genes identified as possible transposable elements; see above: ‘Gene annotation’). This provided transcript abundance estimates for 40,133 CDS models (Supplementary Data 2). Transcript abundance was quantified and normalized as reads per kilobase pairs per million aligned reads (RPKM). After filtering out models exhibiting negligible expression (mean RPKM value of below 0.4), 33,204 CDS models were analysed as potential gene expression markers (GEMs; Supplementary Data 3). SNPs were called by the meta-analysis of alignments (as described in ref. 95) of mRNA-seq reads obtained from each of the 182 accessions. SNP positions were excluded if they did not have a read depth in excess of 20, a base call quality above Q20, missing data below 0.25, and three alleles or fewer. An additional noise threshold was used to reduce the effect of sequencing errors, whereby ambiguous bases were only allowed to be called if both bases were present at 0.15 or above. This resulted in a final set of 394,006 SNPs (Supplementary Data 4) of which 234,519 had minor allele frequencies in excess of 0.05, and all of which were within the CDS models constituting the GEM panel.
The SNP data set for the 182 accessions was entered into the program PSIKO96 to produce a Q matrix, which was composed of two population clusters. The SNP genotypes, Q matrix and ADB damage scores for these trees3 were incorporated into a compressed mixed linear model97 implemented in the GAPIT R package98, with missing data imputed to the major allele. The kinship matrix used in this analysis was also generated by GAPIT.
GEM associations were calculated by a fixed effect linear model in R with RPKM values and the Q matrix inferred by PSIKO as the explanatory variables and damage score the response variable. Coefficients of correlation (r2), regression coefficients, constants and significance values were outputted for each regression.
Twenty GEMs were associated with damage scores (Supplementary Data 3). A previous analysis of the gene expression data, based on a simple mRNA transcript reference, identified only 13 GEMs associated with ADB damage in ash3, with the strongest associations exhibiting higher P values than the present study (best P values 5.31 × 10−12 and 9.83 × 10−13, respectively). The CDS models for the top three GEMs identified in the present study had very high BLAST similarity to the transcripts for two of the GEMs identified in the previous study. FRAEX38873_v2_000173540.4 (P = 1.95 × 10−10) corresponded with Gene_23247_Predicted_mRNA_scaffold3380 from the previous study, but Gene_19216_Predicted_mRNA_scaffold2427 resolved into two distinct CDS models in the present study (FRAEX38873_v2_000261470.1, P = 9.83 × 10−13 and FRAEX38873_v2_000199610.1, P = 6.01 × 10−12). The qRT–PCR primers designed for the previous analysis3 were adequate for assaying FRAEX38873_v2_000173540.4 and FRAEX38873_v2_000261470.1 and new primers were designed for FRAEX38873_v2_000199610.1.
Two of the 20 significantly associated GEMs in the present study, FRAEX38873_v2_000048360.1 (P = 1.77 × 10−9) and FRAEX38873_v2_000048340.1 (P = 3.48 × 10−7), did not have high BLAST similarity to GEMS found in the previous study. However, these GEMs were highly similar to a cDNA transcript containing a predictive A/G SNP (termed a cSNP) identified previously, where presence of a G allele was associated with low damage scores. Both of these GEMS contained the ‘less susceptible’ G variant. A third paralogous gene in this family with the A variant was also found (FRAEX38873_v2_000184430.1), and was not identified as a GEM associated with damage score (P = 0.02). The present study therefore resolves this cSNP marker into three paralogous genes, two fixed for a ‘less susceptible’ G nucleotide, and one a ‘susceptible’ A nucleotide.
These five GEMs were applied using qRT–PCR, and, in the case of FRAEX38873_v2_000048360.1 and FRAEX38873_v2_000048340.1 RT–PCR, to a small test panel of 58 Danish accessions (henceforth ‘Danish Test Panel’) to assess their predictive capabilities in a similar way as in ref. 3. Unlike this previous study, however, ratios between the bases of the FRAEX38873_v2_000048360.1 and FRAEX38873_v2_000048340.1 were scored by eye (instead of simply scoring the presence or absence of the ‘less susceptible’ nucleotide), to estimate levels of gene expression for the ‘less susceptible’ paralogue, while maintaining the simplicity of the assay. These ratios and the qRT–PCR assays for the other three GEMs were combined into a single predicted damage score for each of the Danish Test Panel, which could then be compared with the observed damage scores for these trees. The combined prediction was correlated with the log mean damage scores for 2013–2014 (r2 = 0.25, P = 6.9 × 10−5) which gave a small improvement in predictive power from the previous analysis (r2 = 0.24, P < 8.4 × 10−5).
Screening of UK F. excelsior accessions for markers of reduced susceptibility to ADB
Four markers were selected for predictive marker assays on the basis of this analysis and previous work on the Danish Test Panel of 58 trees3. The three GEM markers most highly associated with disease damage were assayed by qRT–PCR using the following primer combinations: FRAEX38873_v2_000261470.1 (GTCGAGGAGGATGGTCAGTCAT, AATCTTGCGGAGGACCTATCG), FRAEX38873_v2_000199610.1 (GGTGAGAGGAAAGGTTCAAATGA, TGCGTTTTGAGAAGGAAACCA), FRAEX38873_v2_000173540.4 (AGGGCAAGGCTTGGAAACAT, TAGGCTTTTTTCTAGCTGCTTGTCA) and GAPDH reference (CTGGGATCGCTCTTAGCAAGA, CGATCAAATCAATCACACGAGAA).
Using RNA extracted from the British Screening Panel, qRT–PCR reactions were performed with SYBR Green fluorescence detection in a qPCR thermal cycler (ViiATM 7, Applied Biosystems, San Francisco, California) using optical grade 384-well plates, allowing all reactions to be performed simultaneously for each target gene. Each reaction was prepared using 3 μl from a 2 ng/μl dilution of cDNA derived from the RT reaction, 5 μl of SYBR Green PCR Master Mix (Applied Biosystems), 200 nM forward and reverse primers, in a total volume of 10 μl. The cycling conditions were 2 min at 50 °C, 10 min at 95 °C, followed by 40 cycles of 95 °C for 15 s and 60 °C for 1 min with the final dissociation at 95 °C for 15 s, 60 °C for 1 min and 95 °C for 15 s. Three technical replicates were used for quantification analysis. Melting curve analysis was performed to evaluate the presence of non-specific PCR products and primer dimers. The specificity and uniqueness of the primers and the amplicons were verified by amplicon sequencing (GATC Biotech LIGHTrun). The results were exported as raw data, and the LinRegPCR99 software was used for baseline correction. The resulting means of triplicate NO values, representing initial concentrations of a target and reference gene, were used to analyse gene expression. For each marker, the set of qRT–PCR quantifications were standardized and rescaled to better emulate the range of RPKM values observed in the original association panel, and then predicted damage scores generated using the regression coefficient and constant from the GEM associations.
An additional GEM marker was assayed as a cSNP by PCR using 1 μl undiluted cDNA, 11.5 μl Thermo Scientific Fermentas PCR Master Mix (2×), 200 nM forward (GGTTTCTCTTCTGCAGCGAG) and reverse (TCCATGATCATCTTGCTGAG) primers in a total volume of 25 μl. The touchdown PCR was performed in using a BIORAD Tetrad PCR machine with the following cycling conditions: 5 min at 94 °C, followed by 15 cycles of 94 °C for 30 s, 63 °C for 30 s −1 °C per cycle, 72 °C for 1 min, and 30 cycles of 94 °C for 30 s, 53 °C for 30 s, 72 °C for 1 min and a final elongation step at 72 °C for 7 min.
Sanger sequences obtained using the forward primer co-amplify GEM FRAEX38873_v2_000048360.1, which is highly associated with ADB disease damage, and another member of the gene family that is not. Owing to a polymorphism between the two (at position 203 of the CDS model mentioned above), the relative abundance of the G nucleotide found in the highly associated GEM could be scored by eye relative to the A nucleotide found in the other paralogue as a cSNP. Previously3, this marker was scored in the Danish Test Panel as the presence or absence of a G nucleotide at this position, but predictions using this method did not incorporate the dynamic range of the gene expression observed. So, for this analysis, G:A peak height ratios were approximated directly from the sequence chromatograms using Softgenetics Mutation Surveyor software for the British Screening Panel and the Danish Test Panel. These ratios were then standardized and rescaled to the RPKM values for FRAEX38873_v2_000048360.1 to predict damage scores as before.
Combined predictions were made by ranking and standardizing the individual predictions for all four markers, and then calculating the mean rank score for each individual tree (Supplementary Data 6). Combined predictions were calculated for the Danish Test Panel and compared with the observed ADB damage scores to ensure that the assay was predictive (Fig. 3).
The four assays were applied in the same way to analyse a panel of 130 accessions originating from across the UK range of F. excelsior (‘British Screening Panel’). Strikingly, when assayed by RT–PCR, expression of the ‘G’ variant paralogues was seen at much higher frequency in the British Screening Panel than in the Danish panels and the mean G:A ratio across the British Screening Panel was 0.67 compared with a mean of 0.03 observed in the Danish Test Panel. Likewise, the gene expression estimates for the British Screening Panel exhibited wider ranges and were more favourable in terms of their expected effect on damage scores. The qRT–PCR results for the GEMs negatively correlated with disease damage (FRAEX38873_v2_000261470.1 and FRAEX38873_v2_000199610.1) exhibited higher mean expression in the UK (0.1 ± 0.11 and 0.12 ± 0.14) versus the Danish Test Panel (0.09 ± 0.08, 0.12 ± 0.11), and the positively correlated FRAEX38873_v2_000173540.4 was on average expressed at a lower level in the British Screening Panel (0.48 ± 0.26) than the Danish Test Panel (0.59 ± 0.17). As expected, this translated to lower combined predictions for ADB damage in the British Screening Panel. Only 9% of the Danish Test Panel accessions were predicted to have a low damage score (defined as 25% canopy damage or less) compared with 25% of the British Screening Panel (Fig. 3).
Analysis of predictive genes
To predict the susceptibility of the reference tree 2451S to ADB, we calculated RPKM values for the five GEM marker CDS models (FRAEX38873_v2_000173540.4, FRAEX38873_v2_000048340.1, FRAEX38873_v2_000048360.1, FRAEX38873_v2_000261470.1 and FRAEX38873_v2_000199610.1) from leaf transcriptome read data. We also did this for each of the trees in the Danish Scoring Panel, and the average of these predictions was taken to provide combined predictions. The top and bottom quartiles from the distribution of predicted scores, which represent the trees with the most susceptible and least susceptible gene expression patterns at these five loci, were then correlated with the RPKM values for the genome sequenced tree 2451S (Extended Data Fig. 4).
RPKM data were also generated for four tissue types: leaf, flower, cambium and root, of the parent of sequenced tree 2451S by mapping raw reads to the CDS reference as before. RPKM data for the 20 CDS models found to be significantly associated with susceptibility to ADB in the GEM analysis were selected and compared for the four tissue types.
The five CDS models represented in the ADB susceptibility predictions were translated using the standard codon usage table and were searched against the non-redundant database in GenBank using BLASTP with default settings to identify top hits to protein sequences in A. thaliana: FRAEX38873_v2_000199610.1 and FRAEX38873_v2_000261470.1 show high similarity to AGAMOUS-LIKE 42/FOREVER YOUNG FLOWER (AGL42/FYF; AT5G62165); FRAEX38873_v2_000173540.4, FRAEX38873_v2_000048340.1 and FRAEX38873_v2_000048360.1 have top hits to SHORT VEGETATIVE PHASE/AGAMOUS-LIKE 22 (SVP/AGL22; AT2G22540). Both AGL42/FYF and SVP/AGL22 are encoded by type II MADS-box genes16. To find potential orthologues from other species, we examined the results of the OrthoMCL analysis for clusters containing AGL42/FYF and SVP/AGL22; all sequences from these clusters were extracted and added to the appropriate F. excelsior sequences to create two data sets, one of AGL42/FYF-like sequences and one of SVP/AGL22-like sequences. To ensure adequate representation of putative orthologues, we further expanded these data sets to include sequences from the OrthoMCL clusters containing A. thaliana proteins from closely related MADS lineages, as identified by previous phylogenetic analyses of type II MADS-box sequences16,17.
Preliminary phylogenetic analysis of these data sets revealed that, despite showing high sequence similarity in BLAST searches, FRAEX38873_v2_000048340.1 and FRAEX38873_v2_000048360.1 do not fall within the clade containing SVP/AGL22 and similar A. thaliana sequences. Therefore, to identify potentially more closely related sequences we performed a BLASTP search of FRAEX38873_v2_000048340.1 and FRAEX38873_v2_000048360.1 against the complete set of 362,741 protein sequences used for the OrthoMCL analysis (see Supplementary Table 10), using the BLAST+ package44 (version 2.2.31+) with an E value cut-off of 1 × 10−5 (FRAEX38873_v2_000048340.1 and FRAEX38873_v2_000048360.1 were not included in the OrthoMCL analysis because they were flagged as putative transposable-element-related genes during annotation). This identified several highly similar sequences from other species with better ranking BLAST hits than those to the A. thaliana proteins. These sequences belong to a single OrthoMCL cluster, and include a tomato (S. lycopersicum) sequence from the apparent orthologue of the potato (S. tuberosum) StMADS11 gene; all sequences from this cluster were added to the SVP/AGL22-like data set, along with the potato StMADS11 protein (GenBank accession number ACH53556.1).
Sequences for both data sets were aligned using M-Coffee100, via the T-Coffee web server (http://www.tcoffee.org; last accessed 7 December 2016) with the following parameter settings: Mpcma_msa Mmafft_msa Mclustalw_msa Mdialigntx_msa Mpoa_msa Mmuscle_msa Mprobcons_msa Mt_coffee_msa -output = score_html clustalw_aln fasta_aln score_ascii phylip -tree -maxnseq = 150 -maxlen = 2500 -case = upper -seqnos = on -outorder = input -run_name = result -multi_core = 4 -quiet = stdout. Positions in the alignments with consensus scores of <6 from M-Coffee were removed; filtered alignments were then run through the TCS tool101 via the T-Coffee web server and any positions with a reliability score of <6 were removed. Recombination was tested for in the filtered alignments using GARD102. Analyses were run via the Datamonkey server (http://www.datamonkey.org; last accessed 1 June 2016) under the best-fit model of evolution (selected with the corrected Akaike’s information criterion103) with β–Γ rate variation and three rate classes. No breakpoints with significant topological incongruence at P ≤ 0.05 were detected for either data set. Phylogenetic analysis of each data set was conducted using Bayesian inference in MrBayes and maximum likelihood in RAxML; input alignments are provided in Supplementary Data 8. MrBayes (version 3.2.5 (ref. 104)) was run using the mixed amino acid model, to allow models of protein sequence evolution to be fit automatically across the alignments; the following parameter settings were used for each data set: prset aamodellpr = mixed, mcmc nruns = 2, nchains = 4, ngen = 1000000, samplefreq = 1000. Parameter values from both runs for each data set were viewed in TRACER version 1.6 (http://beast.bio.ed.ac.uk/Tracer) to confirm that effective sample sizes of >200 had been obtained for each parameter and stationarity reached. Trees sampled during the first 100,000 generations of each run were discarded as the burn-in; trees and parameter values were summarized in MrBayes using the sumt and sump commands. RAxML (version 8.2.8 (ref. 105)) was run using the option to automatically determine the best protein substitution model, with 1,000 replicates of the rapid bootstrap algorithm; parameter settings were as follows: raxmlHPC -f a -x 13102 -p 29503 -# 1000 -m PROTGAMMAAUTO.
The phylogenetic analysis suggested that FRAEX38873_v2_000173540.4 is a likely orthologue of the A. thaliana SVP/AGL22 gene, or possibly AGL24, whereas FRAEX38873_v2_000048340.1 and FRAEX38873_v2_000048360.1 appear orthologous to the potato StMADS11 gene (Extended Data Fig. 5). These all belong to the SVP/StMADS11 group16 of type II MADS-box genes. FRAEX38873_v2_000261470.1 and FRAEX38873_v2_000199610.1 cluster with the A. thaliana SUPPRESSOR of OVEREXPRESSION of CONSTANS 1(SOC1)-like proteins AGL42, AGL71 and AGL72 (Extended Data Fig. 5). The two other major clades within the phylogenetic tree include the AGL20/SOC1 protein and the AG14 and AGL19 proteins (Extended Data Fig. 5); together, the AGL42/AGL71/AGL72-, AL20- and AGL14/AGL19-containing clades are known as the SOC1/TM3 group of type II MADS-box proteins16,17.
In A. thaliana, AGL42, AGL71 and AGL72 have redundant functions in controlling flowering time and appear to be regulated by AGL20/SOC1 (ref. 20). In turn, AGL20/SOC1 is regulated by both AGL22/SVP and AGL24 (refs 18, 19), which are floral meristem identity genes with redundant functions during early stages of flower development21. The StMADS11 gene does not appear to have a direct orthologue in A. thaliana, but in potato (S. tuberosum) StMADS11 is expressed in vegetative tissues106. Despite their well-known roles in floral regulation, SVP/StMADS11 and SOC1/TM3 proteins are likely to have wider functions. In A. thaliana, it is suggested that AGL22/SVP is also required for age-related resistance, which gives older tissues of plants enhanced pathogen tolerance or resistance24. The B. rapa BrMADS44 gene, which appears orthologous to AGL42, shows differential expression in response to cold and drought stress; some B. rapa genes belonging to the SVP/StMADS11 clade are also differentially expressed in response to these stresses, indicating a potential role in stress resistance22. Furthermore, many genes involved in regulation of flowering time in A. thaliana are involved in controlling phenology in perennial trees species and genes belonging to the SVP/StMADS11 clade have potential roles in growth cessation, bud set and dormancy23.
Metabolomic profiling
To understand whether trees with low and high susceptibility vary in their metabolite profiles as well as their transcriptomes, we undertook untargeted metabolite profiling on a subset of the Danish Test Panel. Untargeted metabolomics has not previously been applied to natural populations but has the potential to identify small molecules (or small-molecule associations) that directly contribute to tolerance or resistance. We compared triplicate samples from five low-susceptibility Danish trees (R-14164C, R-14184A, R-14193A, R-14198B, R-14181) and five high-susceptibility trees (R-14169, R-14127, R-14156 R-14120, 25UTaps).
Three leaflets from each triplicate sample were freeze dried and gently crushed to mix tissue. Approximately 100–150 mg was ground to a fine powder using a TissueLyser (Qiagen), and 10 mg was extracted in 400 μl 80% MeOH containing d5-IAA internal standard at 2.5 ng/ml ([2H5]indole-3-acetic acid; OlChemIm, Czech Republic), centrifuged (10,000 g, 4 °C, 10 min) and the pellet re-extracted in 80% MeOH. The pooled supernatants were filtered through a 0.2 μm syringe filter (Phenomenex, UK).
These leaf extracts (5 μl) were analysed using a Polaris C18 1.8 μm, 2.1 mm × 250 mm reverse-phase analytical column (Agilent Technologies, Palo Alto, California, USA) and samples resolved on an Agilent 1200 series Rapid Resolution HPLC system coupled to a quadrupole time-of-flight QToF 6520 mass spectrometer (Agilent Technologies, Palo Alto, California, USA). Buffers were as follows: positive ion mode; mobile phase A (5% acetonitrile, 0.1% formic acid), mobile phase B (95% acetonitrile with 0.1% formic acid); negative ion mode; mobile phase A (5% acetonitrile with 1 mM ammonium fluoride), mobile phase B (95% acetonitrile). The following gradient was used: 0–10 min, 0% B; 10–30 min, 0–100% B; 30–40 min, 100% B. The flow rate was 0.25 ml/min and the column temperature was held at 35 °C throughout. The source conditions for electrospray ionization were as follows: gas temperature was 325 °C with a drying gas flow rate of 9 l/min and a nebulizer pressure of 35 pounds per square inch gauge. The capillary voltage was 3.5 kV in both positive and negative ion mode. The fragmentor voltage was 115 V and skimmer 70 V. Scanning was performed using the autoMS/MS function at four scans per second for precursor ion surveying and three scans per second for MS/MS with a sloped collision energy of 3.5 V per 100 Da with an offset of 5 V.
Positive and negative ion data were converted into mzData using the export option in Agilent MassHunter. Peak identification and alignment was performed using the Bioconductor R package xcms107 and features were detected using the centWave method108 for high-resolution liquid chromatography/mass spectrometry data in centroid mode at 30 p.p.m. Changes from the default parameters were mzdiff = 0.01, peakwidth = 10-80, noise = 1000, prefilter = 3,500. Peaks were matched across samples using the density method with a bw = 5 and mzwid = 0.025 and retention time correlated using the obiwarp algorithm with profStep = 0.5. Missing peak data were filled in the peaklists generated from the ADB low-susceptibility ash leaf samples compared with the peaklists generated from the ADB susceptible leaves. The resulting peaklists were annotated using the Bioconductor R package, CAMERA109. The peaks were grouped using 0.05% of the width of the full width at half maximum, and groups correlated using a P value of 0.05 and calculating correlation inside and across samples. Isotopes and adducts were annotated using a 10 p.p.m. error.
Statistical analysis and modelling was performed using MetaboAnalyst version 3.0 with the following parameters. Missing values were replaced using a KNN missing value estimation. Data were filtered (40%) to remove non-informative variables using the interquartile range. Samples were normalized using the internal standard d5-IAA (POS: M181T1448; NEG: M179T1382). Data were auto-scaled.
Peaks from the three replicates were aligned with xcms for both positive and negative mode and features tested for practical significance to determine the differences between the tolerant and susceptible genotypes. In addition, PLS-DA was performed using MetaboAnalyst, allowing the discrimination of tolerant and susceptible genotypes on the basis of their metabolic profiles (Fig. 4a).
The individual features (putative metabolites) that contributed to the separation between the different classes were further characterized. We first applied a range of univariate and multivariate statistical tests to determine the importance of these features. This included variable influence on the projection (VIP) values derived from PLS-DA scores, practical significance, t-test, P value, Benjamini and Hochberg false discovery rate P value, effect size and Random Forest analysis, and MS/MS fragmentation network analysis. For example, using Random Forest, significant features were ranked by mean decrease in classification accuracy with 14 out of 15 susceptible samples (out-of-bag error: 0.033; class error 0.07) and 15 out of 15 tolerant samples correctly classified.
For all further analyses we chose to use statistical and practical significance (Response Screening, JMP version 12) to identify features with a practical significance for identification. A combination of k-means clustering was used to group features by patterns of abundance and by retention time. This enabled the clustering of base peaks with their associated isotopes and adducts. Product ions were identified using MS/MS data in Agilent MassHunter Qualitative Analysis version 4.
Identification was not possible for those features with no fragmentation, or lacking significant supporting adducts. Many features of interest were identified but require further work to provide confident attributions, while some features did not provide fragmentation patterns. We thus restricted further identification and characterization to a highly discriminatory class of compounds of the iridoid glycosides and predominantly compounds previously recorded in Oleaceae, summarized in Extended Data Figs 6, 7, 8, 9 and Supplementary Data 9. We validated these identifications using three methods: MS/MS fragmentation networking (Fig. 4c), MS/MS mirror plot (Extended Data Fig. 6) and accurate mass MS/MS product ion structure correlation (Extended Data Fig. 7). The MS/MS fragmentation network was generated after extracting the m/z values of the MS/MS product ions from the discriminatory features using MassHunter Qualitative Analysis Version 4 and visualized using Cytoscape, indicating product ion masses that had been previously reported from fragmentation of iridoid glycosides110. Further validation was performed through a mirror plot comparing the MS/MS spectra of four features (N2–N5) detected in negative mode with an electrospray ionization-time of flight/ion trap-mass spectrometry (ESI-TOF/IT-MS) spectrum of elenolic acid glucoside taken from the literature111. Finally, the accurate masses of MS/MS product ions from four discriminatory features identified in negative mode (N1–N4) were correlated with the structure of the putatively identified compound using MassHunter Molecular Structure Correlator (Agilent).
A timeline for the project may be found in Supplementary Table 14.
URL
Genome website: http://www.ashgenome.org.
Data availability
The reference tree is growing at Earth Trust with accession number 2451S. Trimmed DNA and RNA reads and the final assembly for the 2451S genome sequence, as well as RNA reads for parent tree and raw reads and consensus read mappings of the European diversity panel trees, have been deposited in European Nucleotide Archive under project accession code PRJEB4958 (http://www.ebi.ac.uk/ena/data/view/PRJEB4958). Metabolomic data that support the findings of this study have been deposited in MetaboLights under accession code MTBLS372 (http://www.ebi.ac.uk/metabolights/MTBLS372). All other data are available from the corresponding author upon reasonable request.
Change history
13 February 2017
The Creative Commons license information was updated in the HTML.
References
Poland, T. M. & McCullough, D. G. Emerald ash borer: invasion of the urban forest and the threat to North America’s ash resource. J. Forest. 104, 118–124 (2006)
Pautasso, M., Aas, G., Queloz, V. & Holdenrieder, O. European ash (Fraxinus excelsior) dieback—a conservation biology challenge. Biol. Conserv. 158, 37–49 (2013)
Harper, A. L. et al. Molecular markers for tolerance of European ash (Fraxinus excelsior) to dieback disease identified using associative transcriptomics. Sci. Rep. 6, 19335 (2016)
The Tomato Genome Consortium. The tomato genome sequence provides insights into fleshy fruit evolution. Nature 485, 635–641 (2012)
Ming, R. et al. Genome of the long-living sacred lotus (Nelumbo nucifera Gaertn.). Genome Biol. 14, R41 (2013)
et al.Potato Genome Sequencing Consortium. Genome sequence and analysis of the tuber crop potato. Nature 475, 189–195 (2011)
Kim, S. et al. Genome sequence of the hot pepper provides insights into the evolution of pungency in Capsicum species. Nature Genet. 46, 270–278 (2014)
Hellsten, U. et al. Fine-scale variation in meiotic recombination in Mimulus inferred from population shotgun sequencing. Proc. Natl Acad. Sci. USA 110, 19478–19482 (2013)
Blanc, G. & Wolfe, K. H. Widespread paleopolyploidy in model plant species inferred from age distributions of duplicate genes. Plant Cell 16, 1667–1678 (2004)
Garrison, E. & Marth, G. Haplotype-based variant detection from short-read sequencing. Preprint at https://arxiv.org/abs/1207.3907v2 [q-bio.GN] (2012)
Pritchard, J. K., Stephens, M. & Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 155, 945–959 (2000)
Heuertz, M. et al. Chloroplast DNA phylogeography of European ashes, Fraxinus sp. (Oleaceae): roles of hybridization and life history traits. Mol. Ecol . 15, 2131–2140 (2006)
Wang, J., Street, N. R., Scofield, D. G. & Ingvarsson, P. K. Natural selection and recombination rate variation shape nucleotide polymorphism across the genomes of three related Populus species. Genetics 202, 1185–1200 (2016)
Li, H. & Durbin, R. Inference of human population history from individual whole-genome sequences. Nature 475, 493–496 (2011)
Mazet, O., Rodríguez, W., Grusea, S., Boitard, S. & Chikhi, L. On the importance of being structured: instantaneous coalescence rates and human evolution—lessons for ancestral population size inference? Heredity 116, 362–371 (2016)
Smaczniak, C., Immink, R. G. H., Angenent, G. C. & Kaufmann, K. Developmental and evolutionary diversity of plant MADS-domain factors: insights from recent studies. Development 139, 3081–3098 (2012)
Wells, C. E., Vendramin, E., Jimenez Tarodo, S., Verde, I. & Bielenberg, D. G. A genome-wide analysis of MADS-box genes in peach [Prunus persica (L.) Batsch]. BMC Plant Biol . 15, 41 (2015)
Liu, C. et al. Direct interaction of AGL24 and SOC1 integrates flowering signals in Arabidopsis. Development 135, 1481–1491 (2008)
Li, D. et al. A repressor complex governs the integration of flowering signals in Arabidopsis. Dev. Cell 15, 110–120 (2008)
Dorca-Fornell, C. et al. The Arabidopsis SOC1-like genes AGL42, AGL71 and AGL72 promote flowering in the shoot apical and axillary meristems. Plant J . 67, 1006–1017 (2011)
Gregis, V., Sessa, A., Colombo, L. & Kater, M. M. AGAMOUS-LIKE24 and SHORT VEGETATIVE PHASE determine floral meristem identity in Arabidopsis. Plant J . 56, 891–902 (2008)
Saha, G. et al. Genome-wide identification and characterization of MADS-box family genes related to organ development and stress resistance in Brassica rapa. BMC Genomics 16, 178 (2015)
Ding, J. & Nilsson, O. Molecular regulation of phenology in trees—because the seasons they are a-changin’. Curr. Opin. Plant Biol. 29, 73–79 (2016)
Wilson, D. C., Carella, P., Isaacs, M. & Cameron, R. K. The floral transition is not the developmental switch that confers competence for the Arabidopsis age-related resistance response to Pseudomonas syringae pv. tomato. Plant Mol. Biol. 83, 235–246 (2013)
Jensen, S. R., Franzyk, H. & Wallander, E. Chemotaxonomy of the Oleaceae: iridoids as taxonomic markers. Phytochemistry 60, 213–231 (2002)
Kubo, I., Matsumoto, A. & Takase, I. A multichemical defense mechanism of bitter olive Olea europaea (Oleaceae): is oleuropein a phytoalexin precursor? J. Chem. Ecol. 11, 251–263 (1985)
Eyles, A. et al. Comparative phloem chemistry of Manchurian (Fraxinus mandshurica) and two North American ash species (Fraxinus americana and Fraxinus pennsylvanica). J. Chem. Ecol. 33, 1430–1448 (2007)
Marak, H. B., Biere, A. & Van Damme, J. M. M. Systemic, genotype-specific induction of two herbivore-deterrent iridoid glycosides in Plantago lanceolata L. in response to fungal infection by Diaporthe adunca (Rob.) Niessel. J. Chem. Ecol . 28, 2429–2448 (2002)
Biere, A., Marak, H. B. & van Damme, J. M. M. Plant chemical defense against herbivores and pathogens: generalized defense or trade-offs? Oecologia 140, 430–441 (2004)
Valenta, V., Moser, D., Kuttner, M., Peterseil, J. & Essl, F. A high-resolution map of emerald ash borer invasion risk for southern central Europe. For. Trees Livelihoods 6, 3075–3086 (2015)
FRAXIGEN. Ash Species in Europe: Biological Characteristics and Practical Guidelines for Sustainable Use (Oxford Forestry Institute, Univ. Oxford, 2005)
Doyle, J. J. A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem. Bull . 19, 11–15 (1987)
Obermayer, R., Leitch, I. J., Hanson, L. & Bennett, M. D. Nuclear DNA C-values in 30 species double the familial representation in pteridophytes. Ann. Bot . 90, 209–217 (2002)
Doležel, J., Greilhuber, J. & Suda, J. Estimation of nuclear DNA content in plants using flow cytometry. Nature Protocols 2, 2233–2244 (2007)
Magocˇ, T. & Salzberg, S. L. FLASH: fast length adjustment of short reads to improve genome assemblies. Bioinformatics 27, 2957–2963 (2011)
Leggett, R. M., Clavijo, B. J., Clissold, L., Clark, M. D. & Caccamo, M. NextClip: an analysis and read preparation tool for Nextera Long Mate Pair libraries. Bioinformatics 30, 566–568 (2014)
Boetzer, M., Henkel, C. V., Jansen, H. J., Butler, D. & Pirovano, W. Scaffolding pre-assembled contigs using SSPACE. Bioinformatics 27, 578–579 (2011)
English, A. C. et al. Mind the gap: upgrading genomes with Pacific Biosciences RS long-read sequencing technology. PLoS ONE 7, e47768 (2012)
Marçais, G. & Kingsford, C. A fast, lock-free approach for efficient parallel counting of occurrences of k-mers. Bioinformatics 27, 764–770 (2011)
Gomez-Alvarez, V., Teal, T. K. & Schmidt, T. M. Systematic artifacts in metagenomes from complex microbial communities. ISME J. 3, 1314–1317 (2009)
Novák, P., Neumann, P., Pech, J., Steinhaisl, J. & Macas, J. RepeatExplorer: a Galaxy-based web server for genome-wide characterization of eukaryotic repetitive elements from next-generation sequence reads. Bioinformatics 29, 792–793 (2013)
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990)
Benson, G. Tandem repeats finder: a program to analyze DNA sequences. Nucleic Acids Res . 27, 573–580 (1999)
Camacho, C. et al. BLAST+: architecture and applications. BMC Bioinformatics 10, 421 (2009)
Lamesch, P. et al. The Arabidopsis Information Resource (TAIR): improved gene annotation and new tools. Nucleic Acids Res . 40, D1202–D1210 (2012)
Slater, G. S. C. & Birney, E. Automated generation of heuristics for biological sequence comparison. BMC Bioinformatics 6, 31 (2005)
Wu, T. D. & Watanabe, C. K. GMAP: a genomic mapping and alignment program for mRNA and EST sequences. Bioinformatics 21, 1859–1875 (2005)
Kopylova, E., Noé, L. & Touzet, H. SortMeRNA: fast and accurate filtering of ribosomal RNAs in metatranscriptomic data. Bioinformatics 28, 3211–3217 (2012)
Trapnell, C., Pachter, L. & Salzberg, S. L. TopHat: discovering splice junctions with RNA-seq. Bioinformatics 25, 1105–1111 (2009)
Trapnell, C. et al. Transcript assembly and quantification by RNA-seq reveals unannotated transcripts and isoform switching during cell differentiation. Nature Biotechnol. 28, 511–515 (2010)
Pertea, M. et al. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nature Biotechnol. 33, 290–295 (2015)
Haas, B. J. et al. De novo transcript sequence reconstruction from RNA-seq using the Trinity platform for reference generation and analysis. Nature Protocols 8, 1494–1512 (2013)
Stanke, M., Steinkamp, R., Waack, S. & Morgenstern, B. AUGUSTUS: a web server for gene finding in eukaryotes. Nucleic Acids Res . 32, W309–W312 (2004)
Haas, B. J. et al. Automated eukaryotic gene structure annotation using EVidenceModeler and the Program to Assemble Spliced Alignments. Genome Biol. 9, R7 (2008)
Haas, B. J. et al. Improving the Arabidopsis genome annotation using maximal transcript alignment assemblies. Nucleic Acids Res . 31, 5654–5666 (2003)
Lara, A. J. et al. in Innovations in Hybrid Intelligent Systems 361–368 (Springer, 2007)
Lowe, T. M. & Eddy, S. R. tRNAscan-SE: a program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res . 25, 955–964 (1997)
Lagesen, K. et al. RNAmmer: consistent and rapid annotation of ribosomal RNA genes. Nucleic Acids Res . 35, 3100–3108 (2007)
Kozomara, A. & Griffiths-Jones, S. miRBase: annotating high confidence microRNAs using deep sequencing data. Nucleic Acids Res . 42, D68–D73 (2014)
Stocks, M. B. et al. The UEA sRNA workbench: a suite of tools for analysing and visualizing next generation sequencing microRNA and small RNA datasets. Bioinformatics 28, 2059–2061 (2012)
Lorenz, R. et al. ViennaRNA Package 2.0. Algorithms Mol. Biol. 6, 26 (2011)
Prüfer, K. et al. PatMaN: rapid alignment of short sequences to large databases. Bioinformatics 24, 1530–1531 (2008)
Muñoz-Mérida, A. et al. De novo assembly and functional annotation of the olive (Olea europaea) transcriptome. DNA Res. 20, 93–108 (2013)
Ibarra-Laclette, E. et al. Architecture and evolution of a minute plant genome. Nature 498, 94–98 (2013)
Denoeud, F. et al. The coffee genome provides insight into the convergent evolution of caffeine biosynthesis. Science 345, 1181–1184 (2014)
Jaillon, O. et al. The grapevine genome sequence suggests ancestral hexaploidization in major angiosperm phyla. Nature 449, 463–467 (2007)
Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res . 32, 1792–1797 (2004)
Yang, Z. PAML 4: phylogenetic analysis by maximum likelihood. Mol. Biol. Evol. 24, 1586–1591 (2007)
Maere, S. et al. Modeling gene and genome duplications in eukaryotes. Proc. Natl Acad. Sci. USA 102, 5454–5459 (2005)
Lyons, E., Pedersen, B., Kane, J. & Freeling, M. The value of nonmodel genomes and an example using SynMap Within CoGe to dissect the hexaploidy that predates the rosids. Trop. Plant Biol. 1, 181–190 (2008)
Kiełbasa, S. M., Wan, R., Sato, K., Horton, P. & Frith, M. C. Adaptive seeds tame genomic sequence comparison. Genome Res. 21, 487–493 (2011)
Haas, B. J., Delcher, A. L., Wortman, J. R. & Salzberg, S. L. DAGchainer: a tool for mining segmental genome duplications and synteny. Bioinformatics 20, 3643–3646 (2004)
Li, L., Stoeckert, C. J., Jr & Roos, D. S. OrthoMCL: identification of ortholog groups for eukaryotic genomes. Genome Res. 13, 2178–2189 (2003)
Amborella Genome Project. The Amborella genome and the evolution of flowering plants. Science 342, 1241089 (2013)
The Arabidopsis Genome Initiative. Analysis of the genome sequence of the flowering plant Arabidopsis thaliana. Nature 408, 796–815 (2000)
Young, N. D. et al. The Medicago genome provides insight into the evolution of rhizobial symbioses. Nature 480, 520–524 (2011)
Zimin, A. et al. Sequencing and assembly of the 22-gb loblolly pine genome. Genetics 196, 875–890 (2014)
Tuskan, G. A. et al. The genome of black cottonwood, Populus trichocarpa (Torr. & Gray). Science 313, 1596–1604 (2006)
Enright, A. J., Van Dongen, S. & Ouzounis, C. A. An efficient algorithm for large-scale detection of protein families. Nucleic Acids Res . 30, 1575–1584 (2002)
Heberle, H., Meirelles, G. V., da Silva, F. R., Telles, G. P. & Minghim, R. InteractiVenn: a web-based tool for the analysis of sets through Venn diagrams. BMC Bioinformatics 16, 169 (2015)
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nature Methods 9, 357–359 (2012)
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009)
Etherington, G. J., Ramirez-Gonzalez, R. H. & MacLean, D. bio-samtools 2: a package for analysis and visualization of sequence and alignment data with SAMtools in Ruby. Bioinformatics 31, 2565–2567 (2015)
Cingolani, P. et al. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118; iso-2; iso-3. Fly (Austin) 6, 80–92 (2012)
Ramirez-Gonzalez, R. H., Uauy, C. & Caccamo, M. PolyMarker: A fast polyploid primer design pipeline. Bioinformatics 31, 2038–2039 (2015)
Danecek, P. et al. The variant call format and VCFtools. Bioinformatics 27, 2156–2158 (2011)
Zheng, X. et al. A high-performance computing toolset for relatedness and principal component analysis of SNP data. Bioinformatics 28, 3326–3328 (2012)
Hubisz, M. J., Falush, D., Stephens, M. & Pritchard, J. K. Inferring weak population structure with the assistance of sample group information. Mol. Ecol. Resour . 9, 1322–1332 (2009)
Earl, D. A. & vonHoldt, B. M. STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv. Genet. Resour . 4, 359–361 (2012)
Jakobsson, M. & Rosenberg, N. A. CLUMPP: a cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics 23, 1801–1806 (2007)
Buschiazzo, E., Ritland, C., Bohlmann, J. & Ritland, K. Slow but not low: genomic comparisons reveal slower evolutionary rate and higher dN/dS in conifers compared to angiosperms. BMC Evol. Biol. 12, 8 (2012)
Barbato, M., Orozco-terWengel, P., Tapio, M. & Bruford, M. W. SNeP: a tool to estimate trends in recent effective population size trajectories using genome-wide SNP data. Front. Genet . 6, 109 (2015)
Meglécz, E. et al. QDD version 3.1: a user-friendly computer program for microsatellite selection and primer design revisited: experimental validation of variables determining genotyping success rate. Mol. Ecol. Resour. 14, 1302–1313 (2014)
Rozen, S. & Skaletsky, H. Primer3 on the WWW for general users and for biologist programmers. Methods Mol. Biol. 132, 365–386 (2000)
Bancroft, I. et al. Dissecting the genome of the polyploid crop oilseed rape by transcriptome sequencing. Nature Biotechnol. 29, 762–766 (2011)
Popescu, A.-A., Harper, A. L., Trick, M., Bancroft, I. & Huber, K. T. A novel and fast approach for population structure inference using kernel-PCA and optimization. Genetics 198, 1421–1431 (2014)
Zhang, Z. et al. Mixed linear model approach adapted for genome-wide association studies. Nature Genet. 42, 355–360 (2010)
Lipka, A. E. et al. GAPIT: genome association and prediction integrated tool. Bioinformatics 28, 2397–2399 (2012)
Ruijter, J. M. et al. Amplification efficiency: linking baseline and bias in the analysis of quantitative PCR data. Nucleic Acids Res . 37, e45 (2009)
Di Tommaso, P. et al. T-Coffee: a web server for the multiple sequence alignment of protein and RNA sequences using structural information and homology extension. Nucleic Acids Res . 39, W13–W17 (2011)
Chang, J.-M., Di Tommaso, P. & Notredame, C. TCS: a new multiple sequence alignment reliability measure to estimate alignment accuracy and improve phylogenetic tree reconstruction. Mol. Biol. Evol. 31, 1625–1637 (2014)
Kosakovsky Pond, S. L., Posada, D., Gravenor, M. B., Woelk, C. H. & Frost, S. D. W. GARD: a genetic algorithm for recombination detection. Bioinformatics 22, 3096–3098 (2006)
Sugiura, N. Further analysis of the data by Akaike’s information criterion and the finite corrections. Commun. Stat. Theory Methods 7, 13–26 (1978)
Ronquist, F. et al. MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst. Biol. 61, 539–542 (2012)
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313 (2014)
Carmona, M. J., Ortega, N. & Garcia-Maroto, F. Isolation and molecular characterization of a new vegetative MADS-box gene from Solanum tuberosum L. Planta 207, 181–188 (1998)
Smith, C. A., Want, E. J., O’Maille, G., Abagyan, R. & Siuzdak, G. XCMS: processing mass spectrometry data for metabolite profiling using nonlinear peak alignment, matching, and identification. Anal. Chem. 78, 779–787 (2006)
Tautenhahn, R., Böttcher, C. & Neumann, S. Highly sensitive feature detection for high resolution LC/MS. BMC Bioinformatics 9, 504 (2008)
Kuhl, C., Tautenhahn, R., Böttcher, C., Larson, T. R. & Neumann, S. CAMERA: an integrated strategy for compound spectra extraction and annotation of liquid chromatography/mass spectrometry data sets. Anal. Chem. 84, 283–289 (2012)
Li, C.-M. et al. Structural characterization of iridoid glucosides by ultra-performance liquid chromatography/electrospray ionization quadrupole time-of-flight tandem mass spectrometry. Rapid Commun. Mass Spectrom. 22, 1941–1954 (2008)
Gupta, S. D. Reactive Oxygen Species and Antioxidants in Higher Plants 323 (CRC, 2010)
Acknowledgements
Eurofins MWG provided a discounted service for Illumina and 454 sequencing of the reference genome, funded by Natural Environment Research Council (NERC) Urgency Grant NE/K01112X/1 to R.J.A.B. The associative transcriptomic and metabolomic work was part of the ‘Nornex’ project led by J.A.D. funded jointly by the UK Biotechnology and Biological Sciences Research Council (BBSRC) (BBS/E/J/000CA5323) and the Department for Environment, Food & Rural Affairs. The Earlham Institute, Norwich, UK, sequenced ‘Tree 35’ funded by ‘Nornex’ and the European Diversity Panel funded by the Earlham Institute National Capability in Genomics (BB/J010375/1) grant. W. Crowther assisted with DNA extractions for the KASP assay; The John Innes Centre contributed KASP analyses. J. F. Miranda assisted with RNA extractions and quantitative PCR with reverse transcription (qRT–PCR) at the University of York. H. V. Florance, N. Smirnoff and the Exeter Metabolomics Facility developed metabolomic methods and ran samples, and T. P. Howard helped with statistics. L.J.K. and R.J.A.B. were partly funded by Living with Environmental Change (LWEC) Tree Health and Plant Biosecurity Initiative - Phase 2 grant BB/L012162/1 to R.J.A.B., S.L. and P. Jepson funded jointly by a grant from the BBSRC, Defra, Economic and Social Research Council, the Forestry Commission, NERC and the Scottish Government, under the Tree Health and Plant Biosecurity Initiative. G.W. was funded by Teagasc Walsh Fellowship 2014001 to R.J.A.B. and G.C.D. E.D.C. was funded by a Marie Skłodowska-Curie Individual Fellowship ‘FraxiFam’ (grant agreement 660003) to E.D.C. and R.J.A.B. E.S.A.S. and J.Z. were funded by the Marie Skłodowska-Curie Initial Training Network INTERCROSSING. J.A.D. received a John Innes Foundation fellowship. We thank A. Joecker for supervising E.S.A.S. at Qiagen and for helpful discussions. R.H.R.G. is supported by a Norwich Research Park PhD Studentship and Earlham Institute Funding and Maintenance Grant. This research used Queen Mary’s MidPlus computational facilities, supported by QMUL Research-IT and funded by Engineering and Physical Sciences Research Council grant EP/K000128/1 and NERC EOS Cloud. D.J.S. acknowledges the support of BBSRC grant BB/N021452/1, which partly supported M.G., C.M.S. and D.J.S. during this work.
Author information
Authors and Affiliations
Contributions
R.J.A.B., M.C., D.S., M.G., J.A.D. and I.B. are the lead investigators. R.J.A.B. coordinated the project and directed work on the reference genome. E.S.A.S. assembled the reference genome and organellar genomes, and analysed gene and genome duplications, European population structure and past effective population sizes. L.J.K. extracted high molecular mass DNA for the European Diversity Panel and conducted repetitive element, OrthoMCL and phylogenetic analyses. G.W. conducted SSR analyses. J.Z. extracted high molecular mass DNA and RNA for the reference genome. E.D.C. analysed genome duplication in the reference genome. D.S. and G.K. performed bioinformatic analyses to annotate the reference genome. M.C. conceived and, with R.J.A.B., oversaw the European Diversity Panel sequencing. R.R.-G., E.S.A.S. and M.C. performed SNP calling on the European Diversity Panel, and KASP genotyping. C.U. conducted KASP genotyping. B.J.C. conceived and oversaw the NEXTERA sequencing on the reference tree genome. M.C., J.A.D. and B.J.C. generated the first-pass ‘Tree 35’ Illumina reads included in the European-wide SNP analysis. E.D.K., L.R.N. and L.V.M. generated, selected and collected Danish samples. D.B. generated and J.C. maintained and sampled the reference tree. J.C., D.B., G.C.D. and S.L. generated, selected and collected UK and European Diversity Panel samples. For the associative transcriptomics, I.B. and A.L.H. conceived and planned the study; A.L.H., L.H. and A.F. performed experiments; bioinformatics was executed by Y.L. and Z.H.; and A.L.H. completed the data analysis. For the metabolomics, C.M.S., D.J.S. and M.G. conceived and conducted the analyses; C.M.S. developed methodology; and D.L.S. processed and extracted samples and ran the mass spectrometer.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information: Nature thanks P. Dorrestein, S. Jansson and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Extended data figures and tables
Extended Data Figure 1 Completeness and coherence of annotation models.
a, Assessment of transcript completeness for the F. excelsior gene set. Transcripts were classified as full-length, 5′- end, 3′-end, internal, coding (open reading frame predicted but no BLAST support), unknown (no BLAST support), mis-assembled and putative ncRNA using Full-lengtherNEXT (version 0.0.8); 76.43% of transcript models were identified as complete. b, Coherence in gene length between F. excelsior and M. guttatus proteins. BLAST analysis (1 × 10−5) identified 2,576 proteins that had reciprocal best hits to 2,605 M. guttatus proteins identified as single copy in M. guttatus, S. lycopersicum, S. tuberosum and V. vinifera (Phytozome). A high coherence in gene length was found between F. excelsior and M. guttatus: r > 0.917.
Extended Data Figure 2 Synteny between ash and monkey flower.
Syntenic dotplot between ash (vertical axis) and monkey flower (horizontal axis) showing regions of multiple synteny. Scaffolds equal to approximately 75% of the ash genome assembly for which syntenic blocks were not detected are not shown. For clarity, small scaffold names are omitted.
Extended Data Figure 3 Population structure of F. excelsior in Europe.
a, Results from STRUCTURE; three replicates were run for k = 3, with each replicate using a different set of 8,955 SNPs as input. Numbers refer to samples, whose locations are given in Supplementary Table 11. b, Δk values for three runs of STRUCTURE of each value of k between k = 2 and k = 19; k = 3 has the highest Δk value of 32.91. c, Effective population size history estimated using the SNeP program, with genotype information from all 38 diversity panel samples at 394,885 SNP loci.
Extended Data Figure 4 Prediction of susceptibility of reference tree.
RPKM values for leaf material from the low heterozygosity reference tree 2451S for five CDS models predictive for ADB. These are shown next to expression profiles for the Danish Scoring Panel with the least susceptible and most susceptible expression patterns according to the GEM analysis.
Extended Data Figure 5 Investigation of the function of GEMs for low susceptibility to ADB.
Unrooted maximum likelihood trees from the RAxML analyses. a, Best scoring maximum likelihood tree from the phylogenetic analysis of SVP/AGL22 and StMADS11-like sequences. b, Best scoring maximum likelihood tree for the SOC1-like sequences. Nodes with bootstrap support values of at least 70 from the maximum likelihood analysis and posterior probabilities of at least 0.95 from the Bayesian analysis are indicated with asterisks. F. excelsior sequences are shown in blue; A. thaliana sequences in red. Four-letter taxon codes at the start of sequence names, where present, follow those in Extended Data Table 1. Sequence names are those from the original data files used for the orthoMCL analysis (see Supplementary Table 10), with the exception of the StMADS11 protein from potato, where the GenBank accession number is given. Common names for selected genes/proteins are annotated on the trees. Scale bars indicate the mean number of substitutions per site.
Extended Data Figure 6 MS/MS mirror plot of elenolic acid glucoside (ESI-TOF/IT-MS) compared with four negative mode features (N2, N3, N4 and N5).
The spectra share four product ions in common: m/z 179, 223, 371 and 403 (elenolic acid glucoside molecular ion). These product ions correspond to a loss of a methyl and hydroxyl group (403–371), loss of hexose (403–223), which is followed by a loss of CO2 (223–179). Elenolic acid corresponds to the secoiridoid part of oleuropein-related compounds, suggesting that these four compounds are secoiridoids112.
Extended Data Figure 7 Identification of MS/MS product ions for four iridoid-glycoside-related features observed in negative mode.
Predicted structure for key m/z peaks using Molecular Structure Correlator (Agilent) and the structure of putative identities. Bonds and atoms in black are present in that product ion, whereas grey indicates loss.
Extended Data Figure 8 Identification of iridoid-glycoside-related metabolites in positive mode.
Box-plots showing abundance (log2 transformed) of features in positive mode discriminating between five different genotypes of high- (TOL) and low- (SUS) susceptibility ash trees.
Extended Data Figure 9 Identification of metabolites.
MS/MS fragmentation product ion data of features discriminating between five different genotypes of high- (TOL) and low- (SUS) susceptibility ash trees in positive mode. Corresponding box-plots are presented in Extended Data Fig. 8.
Supplementary information
Supplementary Data 1
Microsatellite marker loci in the ash genome. Primer sequences, locus information, and genome coordinates for 664 SSR sequences in European ash (Fraxinus excelsior), with multiplexing information for 48 markers. (PDF 825 kb)
Supplementary Data 2
Gene expression in the Danish Scored Panel. RPKM values for 182 Danish ash accessions calculated from mapping RNA-seq reads to 40,133 CDS models. (XLSX 96394 kb)
Supplementary Data 3
Associative transcriptomic results in the Danish Scored Panel. Association of 33,205 gene expression markers (GEM) with damages scores for 182 Danish ash accessions. (XLSX 3259 kb)
Supplementary Data 4
Genotypes in the Danish Scored Panel. Genotypes of 182 Danish ash accessions at 395,200 SNP loci obtained following mapping to CDS model reference. (ZIP 14361 kb)
Supplementary Data 5
Details of top 20 GEM markers. BLAST hits and OrthoMCL results for twenty GEMs associated with damage score in 182 Danish Scored Panel accessions. (XLSX 13 kb)
Supplementary Data 6
Marker assay results in the Danish Test Panel and British Screening Panel: qRT-PCR results, cSNP scores and predicted damage scores. (XLSX 47 kb)
Supplementary Data 7
Validation of SNP calls in the European Diversity Panel. KASP calls for 47 randomly selected SNPs from 37 trees in the European Diversity Panel. (XLSX 17 kb)
Supplementary Data 8
Alignments of protein sequences used in phylogenetic analysis. Nexus formatted alignment blocks for the SOC1- and SVP/StMADS11-like datasets. (TXT 15 kb)
Supplementary Data 9
Details of predicted iridoid glycosides discriminating among trees with high and low susceptibility to ADB. (XLSX 45 kb)
Supplementary Data 10
This file contains Supplementary Tables 1-14. (PDF 262 kb)
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International (CC BY 4.0) licence. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons licence, users will need to obtain permission from the licence holder to reproduce the material. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sollars, E., Harper, A., Kelly, L. et al. Genome sequence and genetic diversity of European ash trees. Nature 541, 212–216 (2017). https://doi.org/10.1038/nature20786
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature20786
This article is cited by
-
Identification of full-sibling families from natural single-tree ash progenies based on SSR markers and genome-wide SNPs
Journal of Plant Diseases and Protection (2024)
-
Integrative analysis of green ash phloem transcripts and proteins during an emerald ash borer infestation
BMC Plant Biology (2023)
-
Transcriptome profiling of Fraxinus excelsior genotypes infested by emerald ash borer
Scientific Data (2023)
-
One hundred single-copy nuclear sequence markers for olive variety identification: a case of fingerprinting database construction in China
Molecular Breeding (2023)
-
Computational identification and systematic classification of cytochrome P450 genes in Pogostemon cablin provide insights into flavonoids biosynthesis
Acta Physiologiae Plantarum (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.