Abstract
No large group of recently extinct placental mammals remains as evolutionarily cryptic as the approximately 280 genera grouped as ‘South American native ungulates’. To Charles Darwin1,2, who first collected their remains, they included perhaps the ‘strangest animal[s] ever discovered’. Today, much like 180 years ago, it is no clearer whether they had one origin or several, arose before or after the Cretaceous/Palaeogene transition 66.2 million years ago3, or are more likely to belong with the elephants and sirenians of superorder Afrotheria than with the euungulates (cattle, horses, and allies) of superorder Laurasiatheria4,5,6. Morphology-based analyses have proved unconvincing because convergences are pervasive among unrelated ungulate-like placentals. Approaches using ancient DNA have also been unsuccessful, probably because of rapid DNA degradation in semitropical and temperate deposits. Here we apply proteomic analysis to screen bone samples of the Late Quaternary South American native ungulate taxa Toxodon (Notoungulata) and Macrauchenia (Litopterna) for phylogenetically informative protein sequences. For each ungulate, we obtain approximately 90% direct sequence coverage of type I collagen α1- and α2-chains, representing approximately 900 of 1,140 amino-acid residues for each subunit. A phylogeny is estimated from an alignment of these fossil sequences with collagen (I) gene transcripts from available mammalian genomes or mass spectrometrically derived sequence data obtained for this study. The resulting consensus tree agrees well with recent higher-level mammalian phylogenies7,8,9. Toxodon and Macrauchenia form a monophyletic group whose sister taxon is not Afrotheria or any of its constituent clades as recently claimed5,6, but instead crown Perissodactyla (horses, tapirs, and rhinoceroses). These results are consistent with the origin of at least some South American native ungulates4,6 from ‘condylarths’, a paraphyletic assembly of archaic placentals. With ongoing improvements in instrumentation and analytical procedures, proteomics may produce a revolution in systematics such as that achieved by genomics, but with the possibility of reaching much further back in time.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Accession codes
Data deposits
Raw MS/MS and PEAKS search files have been deposited to the ProteomeXchange with identifier PXD001411. Generated COL1 species consensus sequences will be available in the UniProt Knowledgebase under the accession numbers C0HJN3–C0HJP8.
References
Owen, R. in The Zoology of the Voyage of H. M. S. Beagle, under the Command of Captain Fitzroy, during the Years 1832 to 1836 (ed. Darwin, C. ) Part I, Numbers I–IV (Smith Elder. 1838–40).
Darwin, C. Journal of Researches into the Geology and Natural History of the Various Countries Visited by H.M.S. Beagle: Under the Command of Captain FitzRoy, R.N. from 1832 to 1836 (Henry Colburn, 1839)
Husson, D. et al. Astronomical calibration of the Maastrichtian (Late Cretaceous). Earth Planet. Sci. Lett. 305, 328–340 (2011)
De Muizon, C. & Cifelli, R. L. The “condylarths” (archaic Ungulata, Mammalia) from the early Palaeocene of Tiupampa (Bolivia): implications on the origin of the South American ungulates. Geodiversitas 22, 1–150 (2000)
Agnolin, F. L. & Chimento, N. R. Afrotherian affinities for endemic South American “ungulates”. Mamm. Biol. 76, 101–108 (2011)
O’Leary, M. A. et al. The placental mammal ancestor and the post-K-Pg radiation of placentals. Science 339, 662–667 (2013)
Dos Reis, M. et al. Phylogenomic datasets provide both precision and accuracy in estimating the timescale of placental mammal phylogeny. Proc. R. Soc. B 279, 3491–3500 (2012)
Song, S., Liu, L., Edwards, S. V. & Wu, S. Resolving conflict in eutherian mammal phylogeny using phylogenomics and the multispecies coalescent model. Proc. Natl Acad. Sci. USA 109, 14942–14947 (2012)
Meredith, R. W. et al. Impacts of the Cretaceous Terrestrial Revolution and KPg extinction on mammal diversification. Science 334, 521–524 (2011)
McKenna, M. C., Bell, S. K. & Simpson, G. G. Classification of Mammals above the Species Level (Columbia Univ. Press, 1997)
Simpson, G. G. The beginning of the age of mammals in South America. Part 2. Bull. Am. Mus. Nat. Hist. 137, 1–259 (1967)
Patterson, B. & Pascual, R. The fossil mammal fauna of South America. Q. Rev. Biol. 43, 409–451 (1968)
Cifelli, R. L. in Mammal Phylogeny (eds Szalay, F. S., Novacek, M. J. & McKenna, M. C. ) 195–216 (Springer, 1993)
Horovitz, I. Eutherian mammal systematics and the origins of South American ungulates as based on postcranial osteology. Bull. Carnegie Mus. Nat. Hist. 63–79 (2004)
Asher, R. J. & Lehmann, T. Dental eruption in afrotherian mammals. BMC Biol. 6, 14 (2008)
Sánchez-Villagra, M. R., Narita, Y. & Kuratani, S. Thoracolumbar vertebral number: the first skeletal synapomorphy for afrotherian mammals. Syst. Biodivers. 5, 1–7 (2007)
Van Bocxlaer, I., Roelants, K., Biju, S. D., Nagaraju, J. & Bossuyt, F. Late Cretaceous vicariance in Gondwanan amphibians. PLoS ONE 1, e74 (2006)
Murphy, W. J., Pringle, T. H., Crider, T. A., Springer, M. S. & Miller, W. Using genomic data to unravel the root of the placental mammal phylogeny. Genome Res. 17, 413–421 (2007)
Billet, G. & Martin, T. No evidence for an afrotherian-like delayed dental eruption in South American notoungulates. Naturwissenschaften 98, 509–517 (2011)
Kramarz, A. & Bond, M. Critical revision of the alleged delayed dental eruption in South American “ungulates”. Mamm. Biol. 79, 170–175 (2014)
Van Doorn, N. L. in Encyclopedia of Global Archaeology 7998–8000 (Springer, 2014)
Buckley, M. & Collins, M. J. Collagen survival and its use for species identification in Holocene-lower Pleistocene bone fragments from British archaeological and paleontological sites. Antiqua 1, e1 (2011)
Hamza, V. M. & Vieira, F. P. in Climate Change - Geophysical Foundations and Ecological Effects (eds Blanco, J. & Kheradmand, H. ) Ch. 6, 113–136 (Intech, 2011)
Buckley, M., Collins, M., Thomas-Oates, J. & Wilson, J. C. Species identification by analysis of bone collagen using matrix-assisted laser desorption/ionisation time-of-flight mass spectrometry. Rapid Commun. Mass Spectrom. 23, 3843–3854 (2009)
Asara, J. M., Schweitzer, M. H., Freimark, L. M., Phillips, M. & Cantley, L. C. Protein sequences from mastodon and Tyrannosaurus rex revealed by mass spectrometry. Science 316, 280–285 (2007)
Wilf, P., Rubén Cúneo, N., Escapa, I. H., Pol, D. & Woodburne, M. O. Splendid and seldom isolated: the paleobiogeography of Patagonia. Annu. Rev. Earth Planet. Sci. 41, 561–603 (2013)
Van Valen, L. M. Paleocene dinosaurs or Cretaceous ungulates in South America? Evol. Monogr. 10, 1–79 (1988)
Vizcaino, M., Mikolajewicz, U., Jungclaus, J. & Schurgers, G. Climate modification by future ice sheet changes and consequences for ice sheet mass balance. Clim. Dyn. 34, 301–324 (2010)
Allentoft, M. E. et al. The half-life of DNA in bone: measuring decay kinetics in 158 dated fossils. Proc. R. Soc. B 279, 4724–4733 (2012)
US. Central Intelligence Agency. The World Factbook 2013–14 (Central Intelligence Agency, 2013)
Ma, B. et al. PEAKS: powerful software for peptide de novo sequencing by tandem mass spectrometry. Rapid Commun. Mass Spectrom. 17, 2337–2342 (2003)
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013)
Drummond, A. J. et al. Geneious v4.7. (Geneious, 2010)
Orlando, L. et al. Recalibrating Equus evolution using the genome sequence of an early Middle Pleistocene horse. Nature 499, 74–78 (2013)
Buckley, M. A molecular phylogeny of Plesiorycteropus reassigns the extinct mammalian order ‘Bibymalagasia’. PLoS ONE 8, e59614 (2013)
Terajima, M. et al. Glycosylation and cross-linking in bone type I collagen. J. Biol. Chem. http://dx.doi.org/10.1074/jbc.M113.528513 (2014)
Hudson, D. M., Weis, M. & Eyre, D. R. Insights on the evolution of prolyl 3-hydroxylation sites from comparative analysis of chicken and Xenopus fibrillar collagens. PLoS ONE 6, e19336 (2011)
Hudson, D. M., Werther, R., Weis, M., Wu, J.-J. & Eyre, D. R. Evolutionary origins of C-terminal (GPP)n 3-hydroxyproline formation in vertebrate tendon collagen. PLoS ONE 9, e93467 (2014)
Schweitzer, M. H. et al. Analyses of soft tissue from Tyrannosaurus rex suggest the presence of protein. Science 316, 277–280 (2007)
Buckley, M. et al. Comment on “Protein sequences from mastodon and Tyrannosaurus rex revealed by mass spectrometry”. Science 319, 33 (2008)
Van Doorn, N. L., Wilson, J., Hollund, H., Soressi, M. & Collins, M. J. Site-specific deamidation of glutamine: a new marker of bone collagen deterioration. Rapid Commun. Mass Spectrom. 26, 2319–2327 (2012)
Rohland, N. & Hofreiter, M. Comparison and optimization of ancient DNA extraction. Biotechniques 42, 343–352 (2007)
Leonard, J. A. et al. Animal DNA in PCR reagents plagues ancient DNA research. J. Archaeol. Sci. 34, 1361–1366 (2007)
Brace, S. et al. Population history of the Hispaniolan hutia Plagiodontia aedium (Rodentia: Capromyidae): testing the model of ancient differentiation on a geotectonically complex Caribbean island. Mol. Ecol. 21, 2239–2253 (2012)
Meyer, M. & Kircher, M. Illumina sequencing library preparation for highly multiplexed target capture and sequencing. Cold Spring Harb. Protoc. 2010, http://dx.doi.org/10.1101/pdb.prot5448 (2010)
Dabney, J. & Meyer, M. Length and GC-biases during sequencing library amplification: a comparison of various polymerase-buffer systems with ancient and modern DNA sequencing libraries. Biotechniques 52, 87–94 (2012)
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet.journal 17, 10–12 (2011)
Zuckerkandl, E., Jones, R. T. & Pauling, L. A comparison of animal hemoglobins by tryptic peptide pattern analysis. Proc. Natl Acad. Sci. USA 46, 1349–1360 (1960)
Sarich, V. M. & Wilson, A. C. Rates of albumin evolution in primates. Proc. Natl Acad. Sci. USA 58, 142–148 (1967)
Lanfear, R., Calcott, B., Ho, S. Y. W. & Guindon, S. Partitionfinder: combined selection of partitioning schemes and substitution models for phylogenetic analyses. Mol. Biol. Evol. 29, 1695–1701 (2012)
Ronquist, F. et al. MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst. Biol. 61, 539–542 (2012)
Rambaut, A., Drummond, A. J. & Suchard, M. Tracer v.1. 6. (2013)
Swofford, D. L. PAUP*. Phylogenetic Analysis Using Parsimony (*and Other Methods) v.4.0b10 (Sinauer Associates, 2003)
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313 (2014)
Drummond, A. J., Suchard, M. A., Xie, D. & Rambaut, A. Bayesian phylogenetics with BEAUti and the BEAST 1.7. Mol. Biol. Evol. 29, 1969–1973 (2012)
Benton, M. J., Donoghue, P. C. J. & Asher, R. J. in The Timetree of Life (eds Hedges, B. S. & Kumar, S. ) 35–86 (Oxford Univ. Press, 2009)
Rambaut, A. & Drummond, A. BEAUTi v.1. 4.2. Bayesian evolutionary analysis utility. (2007)
Humphrey, W., Dalke, A. & Schulten, K. VMD – Visual Molecular Dynamics. J. Mol. Graph. 14, 33–38 (1996)
Acknowledgements
We thank the Museo Argentino de Ciencias Naturales “Bernardino Rivadavia”, Buenos Aires (MACN), the Museo de La Plata (MLP), and the Natural History Museum of Denmark, Copenhagen (ZMK), for allowing us to sample fossil specimens in their collections for this project. The American Museum of Natural History and the Copenhagen Zoo provided samples of extant mammals suitable for collagen extraction. Mogens Andersen and Kristian Gregersen of ZMK provided information on specimens in their care. This work was partly supported by SYNTAX award “Barcode of Death”, European Research Council (ERC) Advanced Award CodeX, ERC Consolidator Award GeneFlow, SYNTHESYS FP7 grant agreement 226506, Engineering and Physical Sciences Research Council NE/G012237/1 and National Science Foundation OPP 1142052. J.T.-O. and D.A.A. are members of the York Centre of Excellence in Mass Spectrometry, created thanks to a major capital investment through Science City York, supported by Yorkshire Forward with funds from the Northern Way Initiative.
Author information
Authors and Affiliations
Contributions
R.D.E.M., I.B., and M.J.C. conceived the project and coordinated the writing of the paper with F.W. and J.A.T., with all authors participating. J.N.G., A.K., M.R., E.C., and R.D.E.M. collected fossil and extant mammal samples for protein extraction. M.W., S.B., I.B., J.A.T., J.B., and M.H. conducted DNA analyses. F.W., M.W., P.A., S.K., C.B., C.K., D.A., J.T.-O., R.F., B.K., P.K., J.A.E., E.C., L.O., and M.J.C. performed protein analyses and interpretation of results. J.A.T., I.B., F.W., and M.W. conducted the phylogenetic analyses and constructed trees. S.T.T., J.N.G., M.R., D.M.P., and R.D.E.M. provided the historical, systematic, and palaeontological framework for this study. J.-J.H., E.W., and J.S. provided technical information. Final editing and manuscript preparation was coordinated by M.J.C., R.D.E.M., and I.B.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
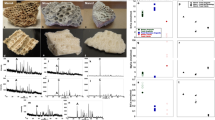
Extended Data Figure 1 Examples of MALDI–TOF–MS and MS/MS product ion spectra.
a, MALDI–TOF–MS ZooMS spectra for Toxodon (upper) and Macrauchenia (lower) were used to screen for samples for the best collagen preservation. b, PEAKS alignment of matching product ion spectra for Macrauchenia MLP 96-V-10-19 (specimen sample number MLP2012.12) highlighting peptides aligning to the sequence GPNGEAGSAGPTGPPGLR. c, d, Annotated PEAKS report of product ion spectra for the same peptide sequence detailed in b for Toxodon (c) and Macrauchenia (d), detailing differences between both genera (gsT and gsA, highlighted) and shared substitutions compared with Equus (gpA for Equus, gpT for Toxodon and Macrauchenia). Note in b that both deamidation (N→D) and variable hydroxylation (P→h) were detected in different peptides covering this region of the sequence.
Extended Data Figure 2 Collagen type I substitution variability for placental mammals (genomic and proteomic data) compared with the dasyurid marsupial Sarcophilus harrisii (Tasmanian devil) as outgroup.
Substitution variability scores range between 0 and 1 and incorporate sequence coverage for a given number of species over a 15-amino-acid moving average (95% standard deviation in lighter tone). Top, along-chain variation in genomic sequence variability (upper red) is similar to proteomic sequence variability (lower blue) both for COL1α1 and for COL1α2 chains. Bottom, molecular surface rendering (via VMD58) of the collagen unit cell taken from coordinates given in Protein Data Bank accession number 3HR2. Colours represent genomic (left) and proteomic (right) sequence variability throughout the structure.
Extended Data Figure 3 Comparison of levels of deamidation for samples in this study with ref. 22 (diamonds).
The Macrauchenia sample was 14C dead, consistent with observed levels of deamidation, which are lower than either Toxodon dated to 12,000 years ago or Equus sp. (Tapalqué; not dated). Dotted lines indicate error ranges on Gln estimation for samples that were not dated or were undateable. The measurement approach used in this study—frequency of deamidation in positions represented in at least seven MS/MS spectra—is different from the approach used in ref. 22, so the absolute values may not be directly comparable.
Extended Data Figure 4 Bayesian constraint tree based on phylogeny published in figure 1 in ref. 6.
See Methods and Supplementary Information section 3.2 for further details and discussion.
Extended Data Figure 5 Maximum clade credibility phylogeny from BEAST molecular dating analysis.
Branch lengths are measured in millions of years; scale axis indicates intervals of 100 Ma. Node labels show 95% highest probability densities for molecular dates (in millions of years). Fossil constraints are provided in Supplementary Table 3. Vertical dashed line indicates Cretaceous/Palaeogene boundary.
Supplementary information
Supplementary Information
This file contains Supplementary Materials, Supplementary Tables 1-5, a Supplementary Discussion and additional references. (PDF 608 kb)
Supplementary Information
This fie contains the MS/MS spectra for residues where Toxodon and Macrauchenia differ. (PDF 12189 kb)
Supplementary Data
This file contains the Amino Acid Sequence data. (TXT 166 kb)
PowerPoint slides
Rights and permissions
About this article
Cite this article
Welker, F., Collins, M., Thomas, J. et al. Ancient proteins resolve the evolutionary history of Darwin’s South American ungulates. Nature 522, 81–84 (2015). https://doi.org/10.1038/nature14249
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature14249
This article is cited by
-
Pleistocene South American native ungulates (Notoungulata and Litopterna) of the historical Roth collections in Switzerland, from the Pampean Region of Argentina
Swiss Journal of Palaeontology (2023)
-
More than 100 years of a mistake: on the anatomy of the atlas of the enigmatic Macrauchenia patachonica
Swiss Journal of Palaeontology (2023)
-
Comparing extraction method efficiency for high-throughput palaeoproteomic bone species identification
Scientific Reports (2023)
-
Anatomy and phylogeny of a new small macraucheniid (Mammalia: Litopterna) from the Bahía Inglesa Formation (late Miocene), Atacama Region, Northern Chile
Journal of Mammalian Evolution (2023)
-
How to weigh a fossil mammal? South American notoungulates as a case study for estimating body mass in extinct clades
Journal of Mammalian Evolution (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.