Abstract
Astrocytes, the most abundant cells in the central nervous system, promote synapse formation and help to refine neural connectivity. Although they are allocated to spatially distinct regional domains during development, it is unknown whether region-restricted astrocytes are functionally heterogeneous. Here we show that postnatal spinal cord astrocytes express several region-specific genes, and that ventral astrocyte-encoded semaphorin 3a (Sema3a) is required for proper motor neuron and sensory neuron circuit organization. Loss of astrocyte-encoded Sema3a leads to dysregulated α-motor neuron axon initial segment orientation, markedly abnormal synaptic inputs, and selective death of α- but not of adjacent γ-motor neurons. In addition, a subset of TrkA+ sensory afferents projects to ectopic ventral positions. These findings demonstrate that stable maintenance of a positional cue by developing astrocytes influences multiple aspects of sensorimotor circuit formation. More generally, they suggest that regional astrocyte heterogeneity may help to coordinate postnatal neural circuit refinement.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Clarke, L. E. & Barres, B. A. Emerging roles of astrocytes in neural circuit development. Nature Rev. Neurosci. 14, 311–321 (2013)
Tsai, H.-H. et al. Regional astrocyte allocation regulates CNS synaptogenesis and repair. Science 337, 358–362 (2012)
Hochstim, C., Deneen, B., Lukaszewicz, A., Zhou, Q. & Anderson, D. J. Identification of positionally distinct astrocyte subtypes whose identities are specified by a homeodomain code. Cell 133, 510–522 (2008)
Muroyama, Y., Fujiwara, Y., Orkin, S. H. & Rowitch, D. H. Specification of astrocytes by bHLH protein SCL in a restricted region of the neural tube. Nature 438, 360–363 (2005)
Rowitch, D. H. & Kriegstein, A. R. Developmental genetics of vertebrate glial-cell specification. Nature 468, 214–222 (2010)
Ge, W.-P., Miyawaki, A., Gage, F. H., Jan, Y.-N. & Jan, L. Y. Local generation of glia is a major astrocyte source in postnatal cortex. Nature 484, 376–380 (2012)
Tien, A.-C. et al. Regulated temporal-spatial astrocyte precursor cell proliferation involves BRAF signalling in mammalian spinal cord. Development 139, 2477–2487 (2012)
Magavi, S., Friedmann, D., Banks, G., Stolfi, A. & Lois, C. Coincident generation of pyramidal neurons and protoplasmic astrocytes in neocortical columns. J. Neurosci. 32, 4762–4772 (2012)
Jessell, T. M. Neuronal specification in the spinal cord: inductive signals and transcriptional codes. Nature Rev. Genet. 1, 20–29 (2000)
Friese, A. et al. Gamma and alpha motor neurons distinguished by expression of transcription factor Err3. Proc. Natl Acad. Sci. USA 106, 13588–13593 (2009)
Arber, S. Motor circuits in action: specification, connectivity, and function. Neuron 74, 975–989 (2012)
Dasen, J. S. & Jessell, T. M. Hox networks and the origins of motor neuron diversity. Curr. Top. Dev. Biol. 88, 169–200 (2009)
Sürmeli, G., Akay, T., Ippolito, G. C., Tucker, P. W. & Jessell, T. M. Patterns of spinal sensory-motor connectivity prescribed by a dorsoventral positional template. Cell 147, 653–665 (2011)
Messersmith, E. K. et al. Semaphorin III can function as a selective chemorepellent to pattern sensory projections in the spinal cord. Neuron 14, 949–959 (1995)
Kolodkin, A. L. et al. Neuropilin is a semaphorin III receptor. Cell 90, 753–762 (1997)
Pasterkamp, R. J. & Giger, R. J. Semaphorin function in neural plasticity and disease. Curr. Opin. Neurobiol. 19, 263–274 (2009)
Cahoy, J. D. et al. A transcriptome database for astrocytes, neurons, and oligodendrocytes: a new resource for understanding brain development and function. J. Neurosci. 28, 264–278 (2008)
Molofsky, A. V. et al. Expression profiling of Aldh1l1-precursors in the developing spinal cord reveals glial lineage-specific genes and direct Sox9-Nfe2l1 interactions. Glia 61, 1518–1532 (2013)
Barros, C. S., Franco, S. J. & Müller, U. Extracellular matrix: functions in the nervous system. Cold Spring Harb. Perspect. Biol. 3, a005108 (2011)
Tissir, F. & Goffinet, A. M. Reelin and brain development. Nature Rev. Neurosci. 4, 496–505 (2003)
Dufour, A. et al. Area specificity and topography of thalamocortical projections are controlled by ephrin/Eph genes. Neuron 39, 453–465 (2003)
Gu, C. et al. Neuropilin-1 conveys semaphorin and VEGF signaling during neural and cardiovascular development. Dev. Cell 5, 45–57 (2003)
Cohen, S. et al. A semaphorin code defines subpopulations of spinal motor neurons during mouse development. Eur. J. Neurosci. 21, 1767–1776 (2005)
Taniguchi, M. et al. Disruption of semaphorin III/D gene causes severe abnormality in peripheral nerve projection. Neuron 19, 519–530 (1997)
Zhuo, L. et al. hGFAP-cre transgenic mice for manipulation of glial and neuronal function in vivo . Genesis 31, 85–94 (2001)
McCall, M. A. et al. Targeted deletion in astrocyte intermediate filament (Gfap) alters neuronal physiology. Proc. Natl Acad. Sci. USA 93, 6361–6366 (1996)
Nishiyama, M. et al. Semaphorin 3A induces CaV2.3 channel-dependent conversion of axons to dendrites. Nature Cell Biol. 13, 676–685 (2011)
Shelly, M. et al. Semaphorin3A regulates neuronal polarization by suppressing axon formation and promoting dendrite growth. Neuron 71, 433–446 (2011)
Duflocq, A., Chareyre, F., Giovannini, M., Couraud, F. & Davenne, M. Characterization of the axon initial segment (AIS) of motor neurons and identification of a para-AIS and a juxtapara-AIS, organized by protein 4.1B. BMC Biol. 9, 66 (2011)
Arber, S. et al. Requirement for the homeobox gene Hb9 in the consolidation of motor neuron identity. Neuron 23, 659–674 (1999)
Huber, A. B. et al. Distinct roles for secreted semaphorin signaling in spinal motor axon guidance. Neuron 48, 949–964 (2005)
Camu, W. & Henderson, C. E. Purification of embryonic rat motoneurons by panning on a monoclonal antibody to the low-affinity NGF receptor. J. Neurosci. Methods 44, 59–70 (1992)
Ashrafi, S. et al. Wnt7A identifies embryonic γ-motor neurons and reveals early postnatal dependence of γ-motor neurons on a muscle spindle-derived signal. J. Neurosci. 32, 8725–8731 (2012)
Brumovsky, P., Watanabe, M. & Hökfelt, T. Expression of the vesicular glutamate transporters-1 and -2 in adult mouse dorsal root ganglia and spinal cord and their regulation by nerve injury. Neuroscience 147, 469–490 (2007)
Brumovsky, P. R. VGLUTs in peripheral neurons and the spinal cord: time for a review. ISRN Neurology 2013, 829753 (2013)
Alvarez, F. J., Villalba, R. M., Zerda, R. & Schneider, S. P. Vesicular glutamate transporters in the spinal cord, with special reference to sensory primary afferent synapses. J. Comp. Neurol. 472, 257–280 (2004)
Mitra, P. & Brownstone, R. M. An in vitro spinal cord slice preparation for recording from lumbar motoneurons of the adult mouse. J. Neurophysiol. 107, 728–741 (2012)
Behar, O., Golden, J. A., Mashimo, H., Schoen, F. J. & Fishman, M. C. Semaphorin III is needed for normal patterning and growth of nerves, bones and heart. Nature 383, 525–528 (1996)
Averill, S., McMahon, S. B., Clary, D. O., Reichardt, L. F. & Priestley, J. V. Immunocytochemical localization of trkA receptors in chemically identified subgroups of adult rat sensory neurons. Eur. J. Neurosci. 7, 1484–1494 (1995)
Ozaki, S. & Snider, W. D. Initial trajectories of sensory axons toward laminar targets in the developing mouse spinal cord. J. Comp. Neurol. 380, 215–229 (1997)
Liu, Y. & Ma, Q. Generation of somatic sensory neuron diversity and implications on sensory coding. Curr. Opin. Neurobiol. 21, 52–60 (2011)
Freeman, M. R. & Rowitch, D. H. Evolving concepts of gliogenesis: a look way back and ahead to the next 25 years. Neuron 80, 613–623 (2013)
Nagai, M. et al. Astrocytes expressing ALS-linked mutated SOD1 release factors selectively toxic to motor neurons. Nature Neurosci. 10, 615–622 (2007)
Chen, A. I., de Nooij, J. C. & Jessell, T. M. Graded activity of transcription factor Runx3 specifies the laminar termination pattern of sensory axons in the developing spinal cord. Neuron 49, 395–408 (2006)
Acknowledgements
We thank J. Flanagan, T. Jessell, J. de Nooj, N. Balaskas, M. Hancock, S. Ohata and R. Krencik for comments on the manuscript and technical suggestions. We are grateful to K. Sabeur, M. Wong and the UCSF Flow Cytometry and Genomics core facilities for expert technical help, A. Kolodkin for Sema3a probe construct, L. Reichardt for the TrkA antibody, J. Dasen for FoxP1 and Scip antibodies, and N. Heintz and J. Dougherty for Aldh1L1-cre mice. A.V.M. is supported by an NIMH Training Grant (5T32MH089920-04) and an APA/Pfizer MD/PhD Psychiatric Research Fellowship. K.W.K is supported by the California Institute for Regenerative Medicine (TG2-01153). S.A.R. is supported by a Ruth L. Kirschstein NRSA FNS081905A. This work was supported by grants from the NINDS (to D.H.R. (R01 NS059893) and J.R.C.), E.M.U. is supported by NIMH (R01MH099595-01), an NIH New Innovator Award (1DP2OD006507-01) and That Man May See. D.H.R. is a HHMI Investigator.
Author information
Authors and Affiliations
Contributions
A.V.M. performed most experiments and data analysis. K.W.K performed electrophysiology under supervision of E.M.U. H.-H.T. contributed to data analysis and experimental design. S.A.R. performed MN purification under supervision of J.R.C. S.M.C performed mouse genotyping. L.M. and S.E.B. performed bioinformatics data processing and analysis. A.V.M. and D.H.R. designed the experiments and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Flow cytometry gating strategy and microarray.
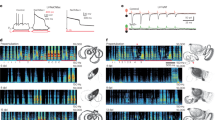
a, Schematic indicating microdissection of Aldh1l1–GFP-positive P7 spinal cord and isolation by flow cytometry using scatter gates, doublet exclusion (not shown) and sorting for GFP-positive cells with live/dead exclusion by DAPI staining. Percentage of Aldh1l1–GFP cells was not significantly different between dorsal and ventral (not shown). b, Summary of differentially expressed genes in astrocytes (AS), whole cord, or both using the analysis parameters indicated. c, Heatmap of all 39 genes differentially expressed between dorsal and ventral cord, highlighting astrocyte-enriched genes with known roles in neural circuit development (red) or extracellular matrix (blue).
Extended Data Figure 2 Coordinate expression of Sema3a and Nrp1 in astrocytes and neurons.
a–c, Sema3a mRNA is expressed in radial glia (RG) and in protoplasmic cells that are NeuN negative throughout the embryonic and early postnatal period. Sema3a was not detected in DRG or in SC white matter (b). d, Sema3a is segregated from Plp-positive oligodendrocytes. e, MN Sema3a expression is detected in α-MN but not γ-MN in cervical SC. f, g, High levels of Nrp1 expression in TrkA+ fibres and cell bodies (white arrowhead) and in MN, but not in PV-positive fibres and cell bodies (yellow arrows). h, Quantification of percentage of Nrp1+ neurons per condition.
Extended Data Figure 3 Fate map of conditional astrocyte deletion lines used in this study.
a, hGFAPcre fate map labels fibrous and a subset of protoplasmic AS but not MN or interneurons in P10 SC. b, Aldh1l1cre fate maps to astrocytes but not to neurons in P10 SC, including α-MN (purple), γ-MN (blue) and interneurons (red).
Extended Data Figure 4 Motor neuron AIS orientation defects in cervical spinal cord.
a, Representative images of cervical SC confocal sections stained to distinguish α− and γ-MN and identify their proximal axon segment (asterisk denotes ventral root). b, Inset shows high-magnification view of representative MN with identifiable AIS and a schematic of their location with respect to the ventral root. c, Overlay of all cervical a-MN angles measured to generate data summarized in Fig. 2c, with positional information preserved, demonstrates that misoriented AIS can be seen at all DV positions.
Extended Data Figure 5 No evidence of abnormal MN cell body positioning with loss of astrocyte-encoded Sema3a.
a, Representative FoxP1 Islet1/2 co-labelling at three rostrocaudal levels in control and mutant animals shows no differences between control and mutant. b, Similar stainings using Scip (a PMC and LMC marker. c, d, No obvious differences in DV or mediolateral boundaries of ChAT+ MN at comparable cervical or lumbar levels at P0 (using Aldh1l1cre to delete Sema3a) and P7 (with hGFAPcre), both time periods where misorientation of AIS is clearly evident.
Extended Data Figure 6 Quantification of ventral interneuron populations after loss of astrocyte-encoded Sema3a.
a, Chx10 staining at E18 and quantification. b, Calbindin staining of Renshaw interneurons at P30 and quantification demonstrates a significant increase at this age. Data are mean ± s.e.m., student’s t-test. Data in a from n = 2 per group, 4 sections per animal; data in b from 4 per group, 4 sections per animal.
Extended Data Figure 7 Additional data and controls for MN electrophysiology.
a, 2 μM strychnine and 20 μM bicuculline block postsynaptic currents (at −55 mV) in a ChAT–GFP+ lumbar MN. b, 20 μM 6,7-dinitroquinoxaline-2,3-dione (DNQX) and 50 μM (2R)-amino-5-phosphonovaleric acid (AP5) block postsynaptic currents (at −75 mV) in a ChAT–GFP+ lumbar MN. c, No difference in input resistance, sIPSC amplitude or sEPSC amplitude between control (Cre-) and hGFAPcre:Sema3afl/fl (fl/fl) MN. n = 5/each; mean ± s.e.m., Student’s t-test.
Extended Data Figure 8 Normal dorsal root ganglia in Aldh1L1cre:Sema3afl/fl mice.
a, No difference in the number of subtype-specific neurons per DRG in control or Aldh1l1cre:Sema3afl/fl mice (n = 3 from 4–5 sections per animal; mean ± s.e.m.; Student’s t-test).
Extended Data Figure 9 Differential expression of regionally heterogeneous astrocyte genes is partly preserved in vitro.
qPCR quantification demonstrates that many regionally heterogeneous microarray genes prospectively identified in vivo remain differentially expressed in vitro after 17 days in culture, including ventral Sema3a. Mean ± s.e.m., n = 3 independent experiments.
Rights and permissions
About this article
Cite this article
Molofsky, A., Kelley, K., Tsai, HH. et al. Astrocyte-encoded positional cues maintain sensorimotor circuit integrity. Nature 509, 189–194 (2014). https://doi.org/10.1038/nature13161
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature13161
This article is cited by
-
Astrocytes in the adult dentate gyrus—balance between adult and developmental tasks
Molecular Psychiatry (2024)
-
Adult expression of Semaphorins and Plexins is essential for motor neuron survival
Scientific Reports (2023)
-
Single-cell transcriptomic landscape of the developing human spinal cord
Nature Neuroscience (2023)
-
Multiplex translaminar imaging in the spinal cord of behaving mice
Nature Communications (2023)
-
Unraveling the lymphatic system in the spinal cord meninges: a critical element in protecting the central nervous system
Cellular and Molecular Life Sciences (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.