Abstract
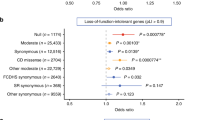
Epileptic encephalopathies are a devastating group of severe childhood epilepsy disorders for which the cause is often unknown1. Here we report a screen for de novo mutations in patients with two classical epileptic encephalopathies: infantile spasms (n = 149) and Lennox–Gastaut syndrome (n = 115). We sequenced the exomes of 264 probands, and their parents, and confirmed 329 de novo mutations. A likelihood analysis showed a significant excess of de novo mutations in the ∼4,000 genes that are the most intolerant to functional genetic variation in the human population (P = 2.9 × 10−3). Among these are GABRB3, with de novo mutations in four patients, and ALG13, with the same de novo mutation in two patients; both genes show clear statistical evidence of association with epileptic encephalopathy. Given the relevant site-specific mutation rates, the probabilities of these outcomes occurring by chance are P = 4.1 × 10−10 and P = 7.8 × 10−12, respectively. Other genes with de novo mutations in this cohort include CACNA1A, CHD2, FLNA, GABRA1, GRIN1, GRIN2B, HNRNPU, IQSEC2, MTOR and NEDD4L. Finally, we show that the de novo mutations observed are enriched in specific gene sets including genes regulated by the fragile X protein (P < 10−8), as has been reported previously for autism spectrum disorders2.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Berg, A. T. et al. Revised terminology and concepts for organization of seizures and epilepsies: report of the ILAE Commission on Classification and Terminology, 2005–2009. Epilepsia 51, 676–685 (2010)
Iossifov, I. et al. De novo gene disruptions in children on the autistic spectrum. Neuron 74, 285–299 (2012)
EPICURE Consortium et al. Genome-wide association analysis of genetic generalized epilepsies implicates susceptibility loci at 1q43, 2p16.1, 2q22.3 and 17q21.32. Hum. Mol. Genet. 21, 5359–5372 (2012)
Heinzen, E. L. et al. Exome sequencing followed by large-scale genotyping fails to identify single rare variants of large effect in idiopathic generalized epilepsy. Am. J. Hum. Genet. 91, 293–302 (2012)
Mulley, J. C. & Mefford, H. C. Epilepsy and the new cytogenetics. Epilepsia 52, 423–432 (2011)
Kasperaviciute, D. et al. Common genetic variation and susceptibility to partial epilepsies: a genome-wide association study. Brain 133, 2136–2147 (2010)
Vissers, L. E. et al. A de novo paradigm for mental retardation. Nature Genet. 42, 1109–1112 (2010)
Neale, B. M. et al. Patterns and rates of exonic de novo mutations in autism spectrum disorders. Nature 485, 242–245 (2012)
Kalscheuer, V. M. et al. Disruption of the serine/threonine kinase 9 gene causes severe X-linked infantile spasms and mental retardation. Am. J. Hum. Genet. 72, 1401–1411 (2003)
Claes, L. et al. De novo mutations in the sodium-channel gene SCN1A cause severe myoclonic epilepsy of infancy. Am. J. Hum. Genet. 68, 1327–1332 (2001)
Saitsu, H. et al. De novo mutations in the gene encoding STXBP1 (MUNC18–1) cause early infantile epileptic encephalopathy. Nature Genet. 40, 782–788 (2008)
Otsuka, M. et al. STXBP1 mutations cause not only Ohtahara syndrome but also West syndrome—result of Japanese cohort study. Epilepsia 51, 2449–2452 (2010)
Veeramah, K. R. et al. De novo pathogenic SCN8A mutation identified by whole-genome sequencing of a family quartet affected by infantile epileptic encephalopathy and SUDEP. Am. J. Hum. Genet. 90, 502–510 (2012)
Kamiya, K. et al. A nonsense mutation of the sodium channel gene SCN2A in a patient with intractable epilepsy and mental decline. J. Neurosci. 24, 2690–2698 (2004)
Tanaka, M., DeLorey, T. M., Delgado-Escueta, A. & Olsen, R. W. in Jasper's Basic Mechanisms of the Epilepsies (eds Noebels, J. L. et al.) (2012)
DeLorey, T. M. et al. Mice lacking the β3 subunit of the GABAA receptor have the epilepsy phenotype and many of the behavioral characteristics of Angelman syndrome. J. Neurosci. 18, 8505–8514 (1998)
Timal, S. et al. Gene identification in the congenital disorders of glycosylation type I by whole-exome sequencing. Hum. Mol. Genet. 21, 4151–4161 (2012)
de Ligt, J. et al. Diagnostic exome sequencing in persons with severe intellectual disability. N. Engl. J. Med. 367, 1921–1929 (2012)
O’Roak, B. J. et al. Sporadic autism exomes reveal a highly interconnected protein network of de novo mutations. Nature 485, 246–250 (2012)
Sanders, S. J. et al. De novo mutations revealed by whole-exome sequencing are strongly associated with autism. Nature 485, 237–241 (2012)
Petrovski, S. et al. Genic intolerance to functional variation and the interpretation of personal genomes. PLoS Gen (in the press) (2013)
Klassen, T. et al. Exome sequencing of ion channel genes reveals complex profiles confounding personal risk assessment in epilepsy. Cell 145, 1036–1048 (2011)
Lemke, J. R. et al. Targeted next generation sequencing as a diagnostic tool in epileptic disorders. Epilepsia 53, 1387–1398 (2012)
Rauch, A. et al. Range of genetic mutations associated with severe non-syndromic sporadic intellectual disability: an exome sequencing study. Lancet 380, 1674–1682 (2012)
Hitomi, Y. et al. Mutations in TNK2 in severe autosomal recessive infantile-onset epilepsy. Ann. Neurol. http://dx.doi.org/doi:10.1002/ana.23934 (2013)
Lee, J. H. et al. De novo somatic mutations in components of the PI3K–AKT3-mTOR pathway cause hemimegalencephaly. Nature Genet. 44, 941–945 (2012)
Gleeson, J. G. et al. Doublecortin, a brain-specific gene mutated in human X-linked lissencephaly and double cortex syndrome, encodes a putative signaling protein. Cell 92, 63–72 (1998)
Fox, J. W. et al. Mutations in filamin 1 prevent migration of cerebral cortical neurons in human periventricular heterotopia. Neuron 21, 1315–1325 (1998)
The EPGP Collaborative. The Epilepsy Phenome/Genome Project. Clin. Trials 10, 568–586 (2013)
Kryukov, G. V., Pennacchio, L. A. & Sunyaev, S. R. Most rare missense alleles are deleterious in humans: implications for complex disease and association studies. Am. J. Hum. Genet. 80, 727–739 (2007)
Kong, A. et al. Rate of de novo mutations and the importance of father’s age to disease risk. Nature 488, 471–475 (2012)
Acknowledgements
We are grateful to the patients, their families, clinical research coordinators and referring physicians for participating in the Epilepsy Phenome/Genome Project (EPGP) and providing the phenotype data and DNA samples used in this study. We thank the following professional and lay organizations for substantial assistance in publicizing EPGP and therefore enabling us to recruit participants effectively: AED Pregnancy Registry, American Epilepsy Society, Association of Child Neurology Nurses, California School Nurses Organization, Child Neurology Society, Citizens United for Research in Epilepsy, Dravet Syndrome Foundation, Epilepsy Alliance of Orange County, Epilepsy Foundation, Epilepsy Therapy Project, Finding a Cure for Epilepsy and Seizures, IDEA League, InfantileSpasms.com, Lennox-Gastaut Syndrome Foundation, PatientsLikeMe, People Against Childhood Epilepsy, PVNH Support & Awareness, and Seizures & Epilepsy Education. We thank the EPGP Administrative Core (C. Freyer, K. Schardein, R.N., M.S., R. Fahlstrom, M.P.H., S. Cristofaro, R.N., B.S.N. and K. McGovern), EPGP Bioinformatics Core (G. Nesbitt, K. McKenna, V. Mays), staff at the Coriell Institute – NINDS Genetics Repository (C. Tarn, A. Scutti), and members of the Duke Center for Human Genome Variation (B. Krueger, J. Bridgers, J. Keebler, H. Shin Kim, E. Campbell, K. Cronin, L. Hong and M. McCall) for their dedication and commitment to this work. We also thank S. Shinnar (Albert Einstein College of Medicine) and N. Risch (University of California, San Francisco) for valuable input into the creation of EPGP and Epi4K, and R. Stewart, K. Gwinn and R. Corriveau from the National Institute of Neurological Disorders and Stroke for their careful oversight and guidance of both EPGP and Epi4K. This work was supported by grants from the National Institute of Neurological Disorders and Stroke (The Epilepsy Phenome/Genome Project NS053998; Epi4K Project 1—Epileptic Encephalopathies NS077364; Epi4K—Administrative Core NS077274; Epi4K—Sequencing, Biostatistics and Bioinformatics Core NS077303 and Epi4K—Phenotyping and Clinical Informatics Core NS077276); Finding a Cure for Epilepsy and Seizures; and the Richard Thalheimer Philanthropic Fund. We would like to acknowledge the following individuals and groups for their contribution of control samples: J. Hoover-Fong, N. Sobreira and D. Valle; The MURDOCK Study Community Registry and Biorepository (D. Murdock); S. Sisodiya; D. Attix; O. Chiba-Falek; V. Shashi; P. Lugar; W. Lowe; S. Palmer; D. Marchuk; Z. Farfel, D. Lancet, E. Pras; Q. Zhao; D. Daskalakis; R. Brown; E. Holtzman; R. Gbadegesin; M. Winn; S. Kerns; and H. Oster. The collection of control samples was funded in part by ARRA 1RC2NS070342, NIAID R56AI098588, the Ellison Medical Foundation New Scholar award AG-NS-0441-08, an award from SAID-Frederick, Inc. (M11-074), and with federal funds by the Center for HIV/AIDS Vaccine Immunology ("CHAVI") under a grant from the National Institute of Allergy and Infectious Diseases, National Institutes of Health (UO1AIO67854).
Author information
Authors and Affiliations
Consortia
Contributions
Initial design of EPGP: B.K.A., O.D., D.D., M.P.E., R.Kuz., D.H.L., R.O., E.H.S. and M.R.W. EPGP patient recruitment and phenotyping: B.A.-K., J.F.B., S.F.B., G.C., D.C., P.Cr., O.D., D.D., M.F., N.B.F., D.F., E.B.G., T.G., S.G., S.R.H., J.H., S.L.H., H.E.K., R.C.K., E.H.K., R.Kup., R.Kuz., D.H.L., S.M.M., P.V.M., E.J.N., J.M.Pao., J.M.Par., K.P., A.P., I.E.S., J.J.S., R.S., J.Si., M.C.S., L.L.T., A.V., E.P.G.V., G.K.V.A., J.L.W. and P.W.-W. Phenotype data analysis: B.A.-K., B.K.A., A.B., G.C., O.D., D.D., J.F., T.G., S.J., A.K., R.C.K., R.Kuz., D.H.L., R.O., J.M.Pao., A.P., I.E.S., R.A.S., E.H.S., J.J.S., J.Su., P.W.-W. and M.R.W. Initial design of Epi4K: S.F.B., P.Co., N.D., D.D., E.E.E., M.P.E., T.G., D.B.G., E.L.H., M.R.J., R.Kuz., D.H.L., A.G.M., H.C.M., T.J.O., R.O., A.P., I.E.S. and E.H.S. Epileptic encephalopathy phenotyping strategy: S.F.B., P.Co., D.D., R.Kuz., D.H.L., R.O., I.E.S. and E.H.S. Encephalopathy phenotyping: D.D., K.B.H., M.R.Z.M., H.C.M., A.P., I.E.S., E.H.S. and C.J.Y. Sequence data analysis and statistical interpretation: A.S.A., D.B.G., Y.Ha., E.L.H., S.E.N., S.P., E.K.R. and E.H.S. Functional evaluation of identified mutations: D.B.G., E.L.H., Y.Hi. and Y.-F.L. Writing of manuscript: A.S.A., S.F.B., D.D., D.B.G., Y.Ha., E.L.H., M.R.J., D.H.L., H.C.M., R.O., A.P., S.P., E.K.R., I.E.S. and E.H.S.
Ethics declarations
Competing interests
The author declare no competing financial interests.
Additional information
Exome sequence data will be available in dbGAP (Epi4K: Gene Discovery in 4,000 Epilepsy Genomes).
Supplementary information
Supplementary Information
This file contains Supplementary Tables 1-15, Supplementary Figures 1-7, Supplementary Methods, Text, Data and Notes and Supplementary References. (PDF 3880 kb)
PowerPoint slides
Rights and permissions
About this article
Cite this article
Epi4K Consortium., Epilepsy Phenome/Genome Project. De novo mutations in epileptic encephalopathies. Nature 501, 217–221 (2013). https://doi.org/10.1038/nature12439
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature12439
This article is cited by
-
NEDD4-2 and the CLC-2 channel regulate neuronal excitability in the pathogenesis of mesial temporal lobe epilepsy
Scientific Reports (2024)
-
Targeting synapse function and loss for treatment of neurodegenerative diseases
Nature Reviews Drug Discovery (2024)
-
De novo truncating variants of TRIM8 and atypical neuro-renal syndrome: a case report and literature review
Italian Journal of Pediatrics (2023)
-
Investigating open reading frames in known and novel transcripts using ORFanage
Nature Computational Science (2023)
-
Germline pathogenic variants in HNRNPU are associated with alterations in blood methylome
European Journal of Human Genetics (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.