Abstract
Worldwide, one person dies every 40 seconds by suicide, a potentially preventable tragedy. A limiting step in our ability to intervene is the lack of objective, reliable predictors. We have previously provided proof of principle for the use of blood gene expression biomarkers to predict future hospitalizations due to suicidality, in male bipolar disorder participants. We now generalize the discovery, prioritization, validation, and testing of such markers across major psychiatric disorders (bipolar disorder, major depressive disorder, schizoaffective disorder, and schizophrenia) in male participants, to understand commonalities and differences. We used a powerful within-participant discovery approach to identify genes that change in expression between no suicidal ideation and high suicidal ideation states (n=37 participants out of a cohort of 217 psychiatric participants followed longitudinally). We then used a convergent functional genomics (CFG) approach with existing prior evidence in the field to prioritize the candidate biomarkers identified in the discovery step. Next, we validated the top biomarkers from the prioritization step for relevance to suicidal behavior, in a demographically matched cohort of suicide completers from the coroner’s office (n=26). The biomarkers for suicidal ideation only are enriched for genes involved in neuronal connectivity and schizophrenia, the biomarkers also validated for suicidal behavior are enriched for genes involved in neuronal activity and mood. The 76 biomarkers that survived Bonferroni correction after validation for suicidal behavior map to biological pathways involved in immune and inflammatory response, mTOR signaling and growth factor regulation. mTOR signaling is necessary for the effects of the rapid-acting antidepressant agent ketamine, providing a novel biological rationale for its possible use in treating acute suicidality. Similarly, MAOB, a target of antidepressant inhibitors, was one of the increased biomarkers for suicidality. We also identified other potential therapeutic targets or biomarkers for drugs known to mitigate suicidality, such as omega-3 fatty acids, lithium and clozapine. Overall, 14% of the top candidate biomarkers also had evidence for involvement in psychological stress response, and 19% for involvement in programmed cell death/cellular suicide (apoptosis). It may be that in the face of adversity (stress), death mechanisms are turned on at a cellular (apoptosis) and organismal level. Finally, we tested the top increased and decreased biomarkers from the discovery for suicidal ideation (CADM1, CLIP4, DTNA, KIF2C), prioritization with CFG for prior evidence (SAT1, SKA2, SLC4A4), and validation for behavior in suicide completers (IL6, MBP, JUN, KLHDC3) steps in a completely independent test cohort of psychiatric participants for prediction of suicidal ideation (n=108), and in a future follow-up cohort of psychiatric participants (n=157) for prediction of psychiatric hospitalizations due to suicidality. The best individual biomarker across psychiatric diagnoses for predicting suicidal ideation was SLC4A4, with a receiver operating characteristic (ROC) area under the curve (AUC) of 72%. For bipolar disorder in particular, SLC4A4 predicted suicidal ideation with an AUC of 93%, and future hospitalizations with an AUC of 70%. SLC4A4 is involved in brain extracellular space pH regulation. Brain pH has been implicated in the pathophysiology of acute panic attacks. We also describe two new clinical information apps, one for affective state (simplified affective state scale, SASS) and one for suicide risk factors (Convergent Functional Information for Suicide, CFI-S), and how well they predict suicidal ideation across psychiatric diagnoses (AUC of 85% for SASS, AUC of 89% for CFI-S). We hypothesized a priori, based on our previous work, that the integration of the top biomarkers and the clinical information into a universal predictive measure (UP-Suicide) would show broad-spectrum predictive ability across psychiatric diagnoses. Indeed, the UP-Suicide was able to predict suicidal ideation across psychiatric diagnoses with an AUC of 92%. For bipolar disorder, it predicted suicidal ideation with an AUC of 98%, and future hospitalizations with an AUC of 94%. Of note, both types of tests we developed (blood biomarkers and clinical information apps) do not require asking the individual assessed if they have thoughts of suicide, as individuals who are truly suicidal often do not share that information with clinicians. We propose that the widespread use of such risk prediction tests as part of routine or targeted healthcare assessments will lead to early disease interception followed by preventive lifestyle modifications and proactive treatment.
Similar content being viewed by others
Introduction
‘Do the difficult things while they are easy and do the great things while they are small’.- Lao Tzu
Predicting suicidal behavior in individuals is one of the hard problems in psychiatry, and in society at large. Improved, objective, and quantitative ways to do it are needed. One cannot always ask individuals if they are suicidal, as desire not to be stopped or future impulsive changes of mind may make their self-report of feelings, thoughts and plans to be unreliable. We had previously provided proof of principle of how first generation blood biomarkers for suicide discovered in male bipolar participants, alone or in combination with clinical symptoms data for anxiety and mood, could have predictive ability for future hospitalizations for suicidality. We now present comprehensive new data for discovery, prioritization, validation, and testing of next-generation broad-spectrum blood biomarkers for suicidal ideation (SI) and behavior, across psychiatric diagnoses. We also describe two clinical information questionnaires in the form of apps, one for affective state (Simplified Affective State Scale, SASS) and one for suicide risk factors (Convergent Functional Information for Suicide, CFI-S), and show their utility in predicting suicidality. Both these instruments do not directly ask about SI. Lastly, we demonstrate how our a priori primary end point, a comprehensive universal predictor for suicide (UP-Suicide), composed of the combination of top biomarkers (from discovery, prioritization and validation), along with CFI-S, and SASS, predicts in independent test cohorts SI and future psychiatric hospitalizations for suicidality.
Materials and methods
Human participants
We present data from four cohorts: one live psychiatric participants discovery cohort; one post-mortem coroner’s office validation cohort; and two live psychiatric participants test cohorts—one for predicting SI and one for predicting future hospitalizations for suicidality (Figure 1).
Cohorts used in study depicting flow of discovery, prioritization, validation and testing of biomarkers from each step.
The live psychiatric participants are part of a larger longitudinal cohort being collected and studied by us. Participants are recruited from the patient population at the Indianapolis VA Medical Center. The participants are recruited largely through referrals from care providers, the use of brochures left in plain sight in public places and mental health clinics, and through word of mouth. All participants understood and signed informed consent forms detailing the research goals, procedure, caveats and safeguards. Participants completed diagnostic assessments by an extensive structured clinical interview—Diagnostic Interview for Genetic Studies—at a baseline visit, followed by up to six testing visits, 3–6 months apart or whenever a hospitalization occurred. At each testing visit, they received a series of psychiatric rating scales, including the Hamilton Rating Scale for Depression-17, which includes a suicidal ideation (SI) rating item (Figure 2), and the blood was drawn. Whole blood (10 ml) was collected in two RNA-stabilizing PAXgene tubes, labeled with an anonymized ID number, and stored at −80 °C in a locked freezer until the time of future processing. Whole-blood (predominantly lymphocyte) RNA was extracted for microarray gene expression studies from the PAXgene tubes, as detailed below. We focused this study on a male population because of the demographics of our catchment area (primarily male in a VA Medical Center), and to minimize any potential gender-related effects on gene expression, which would have decreased the discriminative power of our analysis given our relatively small sample size.
Discovery cohort: longitudinal within-participant analysis. Phchp### is study ID for each participant. V# denotes visit number (1, 2, 3, 4, 5 or 6). (a) Suicidal ideation (SI) scoring. (b) Participants and visits. (c) PhenoChipping: two-way unsupervised hierarchical clustering of all participant visits in the discovery cohort vs 18 quantitative phenotypes measuring affective state and suicidality. A—anxiety items (anxiety, uncertainty, fear, anger, average). M—mood items (mood, motivation, movement, thinking, self-esteem, interest, appetite, average). SASS, simplified affective state scale; STAI-STATE, state trait anxiety inventory, state subscale; YMRS, Young Mania Rating Scale.
Our within-participant discovery cohort, from which the biomarker data were derived, consisted of 37 male participants with psychiatric disorders, with multiple visits in our laboratory, who each had at least one diametric change in SI scores from no SI to high SI from one testing visit to another testing visit. There was one participant with six visits, one participant with five visits, one participant with four visits, 23 participants with three visits each, and 11 participants with two visits each, resulting in a total of 106 blood samples for subsequent microarray studies (Figure 2 and Table 1).
Our post-mortem cohort, in which the top biomarker findings were validated, consisted of a demographically matched cohort of 26 male violent suicide completers obtained through the Marion County coroner’s office (Table 1 and Supplementary Table S2). We required a last observed alive post-mortem interval of 24 h or less, and the cases selected had completed suicide by means other than overdose, which could affect gene expression. Fifteen participants completed suicide by gunshot to head or chest, nine by hanging, one by electrocution and one by slit wrist. Next of kin signed informed consent at the coroner’s office for donation of blood for research. The samples were collected as part of our INBRAIN initiative (Indiana Center for Biomarker Research in Neuropsychiatry).
Our independent test cohort for predicting SI (Table 1) consisted of 108 male participants with psychiatric disorders, demographically matched with the discovery cohort, with one or multiple testing visits in our laboratory, with either no SI, intermediate SI, or high SI, resulting in a total of 223 blood samples in whom whole-genome blood gene expression data were obtained (Table 1 and Supplementary Table S1).
Our test cohort for predicting future hospitalizations (Table 1 and Supplementary Table S1) consisted of male participants in whom whole-genome blood gene expression data were obtained by us at testing visits over the years as part of our longitudinal study. If the participants had multiple testing visits, the visit with the highest marker (or combination of markers) levels was selected for the analyses (so called “high watermark” or index visit). The participants’ subsequent number of psychiatric hospitalizations, with or without suicidality, was tabulated from electronic medical records. All participants had at least 1 year of follow-up or more at our VA Medical Center since the time of the testing visits in the laboratory. Participants were evaluated for the presence of future hospitalizations for suicidality, and for the frequency of such hospitalizations. A hospitalization was deemed to be without suicidality if suicidality was not listed as a reason for admission, and no SI was described in the admission and discharge medical notes. Conversely, a hospitalization was deemed to be because of suicidality if suicidal acts or intent was listed as a reason for admission, and/or SI was described in the admission and discharge medical notes.
Medications
The participants in the discovery cohort were all diagnosed with various psychiatric disorders (Table 1). Their psychiatric medications were listed in their electronic medical records, and documented by us at the time of each testing visit. The participants were on a variety of different psychiatric medications: mood stabilizers; antidepressants; antipsychotics; benzodiazepines; and others (data not shown). Medications can have a strong influence on gene expression. However, our discovery of differentially expressed genes was based on within-participant analyses, which factor out not only genetic background effects but also medication effects, as the participants had no major medication changes between visits. Moreover, there was no consistent pattern in any particular type of medication, or between any change in medications and SI, in the rare instances where there were changes in medications between visits.
Human blood gene expression experiments and analyses
RNA extraction
Whole blood (2.5–5 ml) was collected into each PaxGene tube by routine venipuncture. PaxGene tubes contain proprietary reagents for the stabilization of RNA. RNA was extracted and processed as previously described.1
Microarrays
Biotin-labeled aRNAs were hybridized to Affymetrix HG-U133 Plus 2.0 GeneChips (Affymetrix; with over 40 000 genes and expressed sequence tags), according to the manufacturer’s protocols. Arrays were stained using standard Affymetrix protocols for antibody signal amplification and scanned on an Affymetrix GeneArray 2500 scanner with a target intensity set at 250. Quality-control measures, including 30/50 ratios for glyceraldehyde 3-phosphate dehydrogenase and b-actin, scale factors and background, were within acceptable limits.
Analysis
We have used the participant’s SI scores at the time of blood collection (0—no SI compared with 2 and above—high SI). We looked at gene expression differences between the no SI and the high SI visits, using a within-participant design, then an across participants summation (Figure 2).
Gene expression analyses in the discovery cohort
We analyzed the data in two ways: an absent–present (AP) approach, as in previous work by us on mood biomarkers2 and on psychosis biomarkers,3 and a differential expression (DE) approach, as in previous work by us on suicide biomarkers.1 The AP approach may capture turning on and off of genes, and the DE approach may capture gradual changes in expression. For the AP approach, we used Affymetrix Microarray Suite Version 5.0 (MAS5) to generate Absent (A), Marginal (M) or Present (P) calls for each probeset on the chip (Affymetrix U133 Plus 2.0 GeneChips) for all participants in the discovery cohort. For the DE approach we imported all Affymetrix microarray data as .cel files into Partek Genomic Suites 6.6 software package (Partek Incorporated, St Louis, MI, USA). Using only the perfect match values, we ran a robust multi-array analysis (RMA), background corrected with quantile normalization and a median polish probeset summarization, to obtain the normalized expression levels of all probesets for each chip. RMA was performed independently for each of the six diagnoses used in the study, to avoid potential artefacts due to different ranges of gene expression in different diagnoses.4 Then the participants' normalized data were extracted from these RMAs and assembled for the different cohorts used in the study.
A/P analysis
For the longitudinal within-participant AP analysis, comparisons were made within-participant between sequential visits to identify changes in gene expression from absent to present that track changes in phene expression (SI) from no SI to high SI. For a comparison, if there was a change from absent to present tracking a change from no SI to high SI, or a change from present to absent tracking a change from high SI to no SI, that was given a score of +1 (increased biomarker in high SI). If the change was in opposite direction in the gene vs the phene (SI), that was given a score of −1 (decreased biomarker in High SI). If there was no change in gene expression between visits despite a change of phene expression (SI), or a change in gene expression between visits despite no change in phene expression (SI), that was given a score of 0 (not tracking as a biomarker). If there was no change in gene expression and no change in SI between visits, that was given a score of +1 if there was concordance (P-P with high SI-high SI, or A-A with no SI-no SI), or a score of −1 if there was the opposite (A-A with high SI-high SI, or P-P with no SI-no SI). If the changes were to M (moderate) instead of P, the values used were 0.5 or –0.5. These values were then summed up across the comparisons in each participant, resulting in a overall score for each gene/probeset in each participant. We also used a perfection bonus. If the gene expression perfectly tracked the SI in a participant that had at least two comparisons (three visits), that probeset was rewarded by a doubling of its overall score. Additionally, we used a non-tracking correction. If there was no change in gene expression in any of the comparisons for a particular participant, that overall score for that probeset in that participant was zero.
DE analysis
For the longitudinal within-participant DE analysis, fold changes (FC) in gene expression were calculated between sequential visits within each participant. Scoring methodology was similar to that used above for AP. Probesets that had a FC⩾1.2 were scored+1 (increased in high SI) or −1 (decreased in high SI). FC⩾1.1 were scored +0.5 or −0.5. FC lower than 1.1 were considered no change. The only difference between the DE and the AP analyses was when scoring comparisons where there was no phene expression (SI) change between visits and no change in gene expression between visits (FC lower than 1.1). In that case, the comparison received the same score as the nearest preceding comparison where there was a change in SI from visit to visit. If no preceding comparison with a change in SI was available, then it was given the same score as the nearest subsequent comparison where there was a change in SI. For DE also we used a perfection bonus and a non-tracking correction. If the gene expression perfectly tracked the SI in a participant that had at least two comparisons (three visits), that probeset was rewarded by a doubling of its score. If there was no change in gene expression in any of the comparisons for a particular participant, that overall score for that probeset in that participant was zero.
Internal score
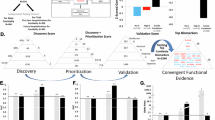
Once scores within each participant were calculated, an algebraic sum across all participants was obtained, for each probeset. Probesets were then given internal points based upon these algebraic sum scores. Probesets with scores above the 33.3% of the distribution (for increased probesets and decreased probesets) received one point, those above 50% of the distribution received two points, and those above 80% of the distribution received four points. For AP analyses, we have 23 probesets which recieved four points, 581 probesets with two points, and 2077 probesets with one point, for a total of 2681 probesets. For DE analyses, we have 31 probesets which received four points, 1294 probesets with two points, and 5839 probesets with one point, for a total of 7164 probesets. The overlap between the two discovery methods is shown in Figure 3. Different probesets may be found by the two methods due to differences in scope (DE capturing genes that are present in both visits of a comparison, that is, PP, but are changed in expression), thresholds (what makes the 33.3% change cut-off across participants varies between methods), and technical detection levels (what is considered in the noise range varies between the methods).
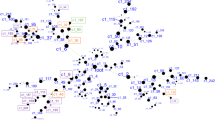
Biomarker discovery, prioritization and validation. (a) Discovery—number of probesets carried forward from the absent–present and differential expression analyses, with an internal score of 1 and above. Red-increased in expression in high suicidal ideation, blue-decreased in expression in high suicidal ideation. (b) Prioritization—convergent functional genomics integration of multiple lines of evidence to prioritize suicide-relevant genes from the discovery step. (c) Validation—top convergent functional genomics genes, with a total score of 4 and above, validated in the cohort of suicide completers. All the genes shown were significantly changed in analysis of variance from no suicidal ideation to high suicidal ideation to suicide completers. *Survived Bonferroni correction. SAT1 (x3) had three different probesets with the same total score of 8.
In total, we identified 9413 probesets with internal convergent functional genomics (CFG) score of 1. Gene names for the probesets were identified using NetAffyx (Affymetrix) and Partek for Affymetrix HG-U133 Plus 2.0 GeneChips, followed by GeneCards to confirm the primary gene symbol. In addition, for those probesets that were not assigned a gene name by NetAffyx or Partek, we used the UCSC Genome Browser to directly map them to known genes, with the following limitations: (1) in case the probeset fell in an intron, that particular gene was assumed to be implicated; and (2) only one gene was assigned to each probeset. Genes were then scored using our manually curated CFG databases as described below (Figure 3).
Convergent functional genomics
Databases
We have established in our laboratory (Laboratory of Neurophenomics, Indiana University School of Medicine, www.neurophenomics.info) manually curated databases of all the human gene expression (post-mortem brain, blood and cell cultures), human genetics (association, copy number variations and linkage) and animal model gene expression and genetic studies published to date on psychiatric disorders. Only the findings deemed significant in the primary publication, by the study authors, using their particular experimental design and thresholds, are included in our databases. Our databases include only primary literature data and do not include review papers or other secondary data integration analyses to avoid redundancy and circularity. These large and constantly updated databases have been used in our CFG cross validation and prioritization (Figure 3). For this study, data from 437 papers on suicide were present in the databases at the time of the CFG analyses.
Human post-mortem brain gene expression evidence
Converging evidence was scored for a gene if there were published reports of human post-mortem data showing changes in expression of that gene or changes in protein levels in brains from participants who died from suicide.
Human blood and other peripheral tissue gene expression data
Converging evidence was scored for a gene if there were published reports of human blood, lymphoblastoid cell lines, cerebrospinal fluid or other peripheral tissue data showing changes in expression of that gene or changes in protein levels in participants who had a history of suicidality or who died from suicide.
Human genetic evidence (association and linkage)
To designate convergence for a particular gene, the gene had to have independent published evidence of association or linkage for suicide. For linkage, the location of each gene was obtained through GeneCards (http://www.genecards.org), and the sex averaged cM location of the start of the gene was then obtained through http://compgen.rutgers.edu/mapinterpolator. For linkage convergence, the start of the gene had to map within 5 cM of the location of a marker linked to the disorder.
CFG scoring
For CFG analysis (Figure 3), the external cross-validating lines of evidence were weighted such that findings in human post-mortem brain tissue, the target organ, were prioritized over peripheral tissue findings and genetic findings, by giving them twice as many points. Human brain expression evidence was given four points, whereas human peripheral evidence was given two points, and human genetic evidence was given a maximum of two points for association, and one point for linkage. Each line of evidence was capped in such a way that any positive findings within that line of evidence result in maximum points, regardless of how many different studies support that single line of evidence, to avoid potential popularity biases. In addition to our external CFG score, we also prioritized genes based upon the initial gene expression analyses used to identify them. Probesets identified by gene expression analyses could receive a maximum of four points. Thus, the maximum possible total CFG score for each gene was 12 points (four points for the internal score and eight points for the external CFG score) (Table 2). The scoring system was decided upon before the analyses were carried out. We sought to give twice as much weight to external score as to internal in order to increase generalizability and avoid fit to cohort of the prioritized genes.5 It has not escaped our attention that other ways of scoring the lines of evidence may give slightly different results in terms of prioritization, if not in terms of the list of genes per se. Nevertheless, we feel this simple scoring system provides a good separation of genes based on gene expression evidence and on independent cross-validating evidence in the field (Figure 3). In the future, with multiple large data sets, machine learning approaches could be used and validated to assign weights to CFG.
Pathway analyses
IPA 9.0 (Ingenuity Systems, www.ingenuity.com, Redwood City, CA, USA), GeneGO MetaCore (Encinitas, CA, USA), and Kyoto Encyclopedia of Genes and Genomes (KEGG) (through the Partek Genomics Suite 6.6 software package) were used to analyze the biological roles, including top canonical pathways, and diseases, of the candidate genes resulting from our work, as well as to identify genes in our data set that are the target of existing drugs (Table 3 and Supplementary Table S3). We ran the analyses together for all the AP and DE probesets with a total CFG score⩾4, then for those of them that showed stepwise change in the suicide completers validation cohort, then for those of them that were nominally significant, and finally for those of them that survived Bonferroni correction.
Validation analyses
For validation of our candidate biomarker genes, we examined which of the top candidate genes (CFG score of 4 or above) were stepwise changed in expression from the no SI group to the high SI group to the suicide completers group. We used an empirical cut-off of 33.3% of the maximum possible CFG score of 12, which also permits the inclusion of potentially novel genes with maximal internal CFG score but no external CFG score. Statistical analyses were performed in SPSS using one-way analysis of variance and Bonferonni corrections.
For the AP analyses, we imported the Affymetrix microarray data files from the participants in the validation cohort of suicide completers into MAS5, alongside the data files from the participants in the discovery cohort.
For the DE analyses, we imported .cel files into Partek Genomic Suites. We then ran a RMA, background corrected with quantile normalization, and a median polish probeset summarization of all the chips from the validation cohort to obtain the normalized expression levels of all probesets for each chip. Partek normalizes expression data into a log base of 2 for visualization purposes. We non-log-transformed expression data by taking 2 to the power of the transformed expression value. We then used the non-log-transformed expression data to compare expression levels of biomarkers in the different groups (Supplementary Figure S1).
Clinical measures
The Simplified Affective State Scale (SASS) is an 11 item scale for measuring mood and anxiety, previously developed and described by us as TASS (Total Affective State Scale).6 The SASS has a set of 11 visual analog scales (7 for mood, 4 for anxiety) that ends up providing a number ranging from 0 to 100 for mood state, and the same for anxiety state. We have now developed an Android app version (Supplementary Figure S2).
CFI-S (Table 4) is a new 22 item scale and Android app (Supplementary Figure S2) for suicide risk, which integrates, in a simple binary fashion (yes-1, no-0), similar to a polygenic risk score, information about known life events, mental health, physical health, stress, addictions and cultural factors that can influence suicide risk.7, 8 For live psychiatric participants, the scale was administered at participant testing visits (n= 57), or scored based on retrospective electronic medical record information and Diagnostic Interview for Genetic Testing (DIGS) information (n=269). For suicide completers (n=35), the scale was based on answers provided by next-of-kin, and corroborated by coroner’s office reports and medical record information. When information was not available for an item, it was not scored (NA).
Combining gene expression and clinical measures
The UP-Suicide construct was decided upon as part of our a priori study design to be broad- spectrum, and combine our top biomarkers from each step (discovery, prioritization, validation) with the phenomic (clinical) markers (SASS and CFI-S). That was our primary end point. Had we done it post hoc with only the markers that showed the best predictive ability in our testing analyses, the results would be even better, but not independent.
Testing analyses
The test cohort for SI and the test cohort for future hospitalizations analyses were assembled out of data that was RMA normalized by diagnosis. Phenomic (clinical) and gene expression markers used for predictions were z scored by diagnosis, to be able to combine different markers into panels and to avoid potential artefacts due to different ranges of phene expression and gene expression in different diagnoses. Markers were combined by computing the average of the increased risk markers minus the average of the decreased risk markers. Predictions were performed using R-studio.
Predicting suicidal ideation
Receiver-operating characteristic (ROC) analyses between marker levels and SI were performed by assigning participants with a HAMD-SI score of 0–1 into the no SI category, and participants with a HAMD-SI score of 2 and greater into the SI category. Additionally, analysis of variance was performed between no (HAMD-SI 0), intermediate (HAMD-SI 1), and high SI participants (HAMD-SI 2 and above) and Pearson R (one-tail) was calculated between HAMD-SI scores and marker levels (Table 5b and Figure 5).
Predicting future hospitalizations for suicidality
We conducted analyses for hospitalizations in the first year following testing, on the participants for which we had at least a year of follow-up data. For each participant in the test cohort for future hospitalizations, the study visit with highest levels for the marker or combination of markers was selected as index visit (or with the lowest levels, in the case of decreased markers). ROC analyses between marker levels and future hospitalizations were performed based on assigning if participants had been hospitalized for suicidality (ideation, attempts) or not following the index testing visit. Additionally, a one-tailed t-test with unequal variance was performed between groups of participants with and without hospitalizations for suicidality. Pearson R (one-tail) correlation was performed between hospitalization frequency (number of hospitalizations for suicidality divided by duration of follow-up) and biomarker score. We also conducted only the correlation analyses for hospitalizations frequency for all future hospitalizations due to suicidality, beyond one year, as this calculation, unlike the ROC and t-test, accounts for the actual length of follow-up, which varied beyond one year from participant to participant.
Results
Discovery of biomarkers for suicidal ideation
We conducted whole-genome gene expression profiling in the blood samples from a longitudinally followed cohort of male participants with psychiatric disorders that predispose to suicidality. The samples were collected at repeated visits, 3–6 months apart. State information about SI was collected from a questionnaire (HAMD) administered at the time of each blood draw (Supplementary Table S1). Out of 217 psychiatric participants (with a total of 531 visits) followed longitudinally in our study, there were 37 participants that switched from a no SI (SI score of 0) to a high SI state (SI score of 2 and above) at different visits, which was our intended discovery group (Figure 2). We used a powerful within-participant design to analyze data from these 37 participants and their 106 visits. A within-participant design factors out genetic variability, as well as some medications, lifestyle, and demographic effects on gene expression, permitting identification of relevant signal with Ns as small as 1.9 Another benefit of a within-participant design may be accuracy/consistency of self-report of psychiatric symptoms (‘phene expression’), similar in rationale to the signal detection benefits it provides in gene expression. The number of participants that met our criteria and were analyzed is small, but comparable to those in human post-mortem brain gene expression studies of suicide. We are indeed treating the blood samples as surrogate tissue for brains, with the caveat that they are not the real target organ. However, with the blood samples from live human participants we have the advantages of in vivo accessibility, better knowledge of the mental state at the time of collection, less technical artifacts and especially of being able to do powerful within-participant analyses from visit to visit.
For discovery, we used two differential expression methodologies: Absent/Present (reflecting on/off of transcription), and Differential Expression (reflecting more subtle gradual changes in expression levels). The genes that tracked SI in each participant were identified in our analyses. We used three thresholds for increased in expression genes and for decreased in expression genes:⩾33.3% (low); ⩾50% (medium); and⩾80% (high) of the maximum scoring increased and decreased gene across participants. Such a restrictive approach was used as a way of minimizing false positives, even at the risk of having false negatives. For example, there were genes on each of the two lists, from AP and DE analyses, that had clear prior evidence for involvement in suicidality, such as OLR110, 11 (32%) and LEPR1, 12 (32%) for AP, and OPRM113, 14 (32%) and CD24 1, 11 (33%) from DE, but were not included in our subsequent analyses because they did not meet our a priori set 33.3% threshold.
Prioritization of biomarkers based on prior evidence in the field
These differentially expressed genes were then prioritized using a Bayesian-like CFG approach (Figure 3) integrating all the previously published human genetic evidence, post-mortem brain gene expression evidence, and peripheral fluids evidence for suicide in the field available at the time of our final analyses (September 2014). This is a way of identifying and prioritizing disease relevant genomic biomarkers, extracting generalizable signal out of potential cohort-specific noise and genetic heterogeneity. We have built in our laboratory manually curated databases of the psychiatric genomic and proteomic literature to date, for use in CFG analyses. The CFG approach is thus a de facto field-wide collaboration. We use in essence, in a Bayesian fashion, the whole body of knowledge in the field to leverage findings from our discovery data sets. Unlike our use of CFG in many previous studies, for the current one we did not use any animal model evidence, as there are to date no clear animal models of self-harm or suicidality published to date.
Validation of biomarkers for behavior in suicide completers
For validation in suicide completers, we used 412 genes that had a CFG score of 4 and above, from AP and DE, reflecting either maximum internal score from discovery or additional external literature cross-validating evidence. Out of these, 208 did not show any stepwise change in suicide completers (non-concordant, NC). As such, they may be involved primarily in ideation and not in behavior (Supplementary Table S6). The remaining 204 genes (49.5%) had levels of expression that were changed stepwise from no SI to high SI to suicide completion. 143 of these genes (34.7%) were nominally significant, and 76 genes (18.4%) survived Bonferroni correction for multiple comparisons (Figure 3 and Supplementary Figure S1). These genes are likely involved in SI and suicidal behavior. (You can have SI without suicidal behavior, but you cannot have suicidal behavior without SI).
Selection of biomarkers for testing of predictive ability
For testing, we decided a priori to select the top scoring increased and decreased biomarkers from each step (discovery, prioritization, validation), so as to avoid potential false negatives in the prioritization step due to lack of prior evidence in the literature, or false negatives in validation step due to possible post-mortem artifacts. The top scoring genes after the discovery step were DTNA and KIF2C from AP, CADM1 and CLIP4 from DE. The top genes after the prioritization with CFG step were SLC4A4 and SKA2 from AP, SAT1 and SKA2 from DE. The top genes after the validation in suicide completers step were IL6 and MBP from AP, JUN and KLHDC3 from DE (Figure 3). Notably, our SAT1 finding is a replication and expansion of our previously reported results identifying SAT1 as a blood biomarker for suicidality in bipolars (Le-Niculescu et al. 2013), and our SKA2 finding is an independent replication of a previous report identifying SKA2 as a blood biomarker for suicidality by Kaminsky and colleagues.15 We also replicated in this larger cohort other top biomarkers from our previous work in bipolar disorder, notably MARCKS and PTEN (Table 2, Supplementary Figure S4). A number of other genes we identified (CADM1, KIF2C, DTNA, CLIP4) are completely novel in terms of their involvement in suicidality.
Biological understanding
We also sought to understand the biology represented by the biomarkers identified by us, and derive some mechanistic and practical insights. We conducted: 1. unbiased biological pathway analyses and hypothesis driven mechanistic queries, 2. overall disease involvement and specific neuropsychiatric disorders queries, and 3. overall drug modulation along with targeted queries for omega-3, lithium and clozapine16 (Table 3, Supplementary Tables S3). Administration of omega-3s in particular may be a mass- deployable therapeutic and preventive strategy.17
The sets of biomarkers identified have biological roles in immune and inflammatory response, growth factor regulation, mTOR signaling, stress, and perhaps overall the switch between cell survival and proliferation vs apoptosis (Table 3 and Supplementary Table S3). 14% of the candidate biomarkers in Supplementary Table S3 have evidence for involvement in psychological stress response, and 19% for involvement in programmed cell death/cellular suicide (apoptosis). An extrapolation can be made and model proposed whereas suicide is a whole body apoptosis (or ‘self-poptosis’) in response to perceived stressful life events.
We also examined evidence for the involvement of these biomarkers for suicidality in other psychiatric disorders, permitting us to address issues of context and specificity (Supplementary Table S3). SKA2, HADHA, SNORA68, RASL11B, CXCL11, HOMEZ, LOC728543, AHCYL1, LDLRAP1, NEAT1 and PAFAH1B2 seem to be relatively specific for suicide, based on the evidence to date in the field. SAT1, IL6, FOXN3 and FKBP5 are less specific for suicide, having equally high evidence for involvement in suicide and in other psychiatric disorders, possibly mediating stress response as a common denominator.11, 18 These boundaries and understanding will likely change as additional evidence in the field accumulates. For example, CADM1, discovered in this work as a top biomarker for suicide, had previous evidence for involvement in other psychiatric disorders, such as autism and bipolar disorder. Interestingly, it was identified in a previous study by us as a blood biomarker increased in expression in low mood states in bipolar participants, and it is increased in expression in the current study in high SI states. Increased expression of CADM1 is associated with decreased cellular proliferation and with apoptosis, and this gene is decreased in expression or silenced in certain types of cancers.
A number of other genes besides CADM1 are changed in opposite direction in suicide in this study vs high mood in our previous mood biomarker study-CHD2, MBP, LPAR1, IGHG1, TEX261 (Supplementary Table S3), suggesting that suicidal participants are in a low mood state. Also, some of the top suicide biomarkers are changed in expression in the same direction as in high psychosis participants in a previous psychosis biomarker study of ours -PIK3C2A, GPM6B, PCBD2, DAB2, IQCH, LAMB1, TEX261 (Supplementary Table S3), suggesting that suicidal participants may be in a psychosis-like state. TEX261 in particular appears in all three studies, decreased in expression in suicide and high hallucinations, and increased in expression in high mood. This protective marker may be an interesting target for future biological studies and drug development. Taken together, the data indicates that suicidality could be viewed as a psychotic dysphoric state, and that TEX261 may be a key biomarker reflecting that state. This molecularly informed view is consistent with the emerging clinical evidence in the field.19
Lastly, we conducted biological pathway analyses on the genes that, after discovery and prioritization, were stepwise changed in suicide completers (n=204) and may be involved in ideation and behavior, vs those that were not stepwise changed (n=208), and that may only be involved in ideation (Supplementary Table S6). The genes involved in ideation map to pathways related to neuronal connectivity (cytoskeleton rearrangement, axonal guidance) and schizophrenia. The genes involved in behavior map to pathways related to neuronal activity (WNT, growth factors) and mood disorders. This is consistent with ideation being related to psychosis, and behavior being related to mood. Of note, clinically, the risk for suicide behavior/completion is higher in mood disorders than in psychotic disorders.
Clinical information
We also developed a simple new 22 item scale and app for suicide risk, Convergent Functional Information for Suicidality (CFI-S), which scores in a simple binary fashion and integrates, information about known life events, mental health, physical health, stress, addictions, and cultural factors that can influence suicide risk.7, 8 Clinical risk predictors and scales are of high interest in the military20 and in the general population at large.21 Our scale builds on those excellent prior achievements, while aiming for comprehensiveness, simplicity and quantification similar to a polygenic risk score. CFI-S is able to distinguish between individuals who committed suicide (coroner’s cases n=35, information obtained from the next-of-kin) and those high-risk participants who did not but had experienced changes in SI (our discovery cohort of psychiatric participants) (Figure 4). We analyzed which items of the CFI-S scale were the most significantly different between high SI live participants and suicide completers. We identified 7 items that were significantly different, 5 of which survived Bonferroni correction: lack of coping skills when faced with stress (P= 3.35e−11), dissatisfaction with current life (P=2.77e−06), lack of hope for the future (4.58e−05), current substance abuse (P=1.25e−04), and acute loss/grief (P= 9.45e−4). It is highly interesting that the top item was inability to cope with stress, which is independently consistent with our biological marker results.
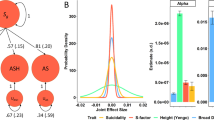
Convergent Functional Information for Suicide (CFI-S) Scale. (a) Validation of scale. Convergent Functional Information for Suicide levels in the discovery cohort and suicide completers. (b) Validation of items. Convergent Functional Information for Suicide was developed independently of any data from this study, by compiling known sociodemographic and clinical risk factors for suicide. It is composed of 22 items that assess the influence of mental health factors, as well as of life satisfaction, physical health, environmental stress, addictions, cultural factors known to influence suicidal behavior, and two demographic factors, age and gender. These 22 items are shown here validated in the discovery cohort and suicide completers in a manner similar to that for biomarkers. Additionally, a student’s t-test was used to evaluate items that were increased in suicide completers when compared to living participants with high suicidal ideation. (c) Predictions. Convergent Functional Information for Suicide predicting SI in the independent test cohort, and predicting future hospitalizations due to suicidality.
We also simplified the wording (and developed a new app for) an 11 item scale for measuring mood and anxiety, the SASS, previously developed and described by us as TASS (Total Affective State Scale).6 The SASS is a set of 11 visual analog scales (7 for mood, 4 for anxiety) that ends up providing a number ranging from 0 to 100 for mood state, and the same for anxiety state.
Testing for predictive ability
The best single biomarker predictor for SI state across all diagnostic groups is SLC4A4 (ROC AUC 0.72, P-value 2.41e−05), the top increased biomarker from our prioritization with CFG of discovery data from AP (Table 5). Within diagnostic groups, the accuracy is even higher. SLC4A4 has very good accuracy at predicting future high SI in bipolar participants (AUC 0.93, P-value 9.45e−06) and good accuracy in schizophrenia participants (AUC 0.76, P-value 0.030). SLC4A4 is a sodium-bicarbonate co-transporter that regulates intracellular pH, and possibly apoptosis. Very little is known to date about its roles in the brain, thus representing a completely novel finding. Brain pH has been reported by Wemmie et al.22 to have a role in pain, fear and panic attacks, which clinically share features with acute SI states.
SKA2, the top decreased biomarker from prioritization with CFG of discovery data from AP and DE, has good accuracy at predicting SI across all diagnostic groups (AUC 0.69, P-value 0.00018), and even better accuracy in bipolar participants (AUC 0.76, P-value 0.0045) and schizophrenia participants (AUC 0.82, P-value 0.011).
The best single biomarker predictor for future hospitalizations for suicidal behavior in the first year across all diagnostic groups was SAT1, the top increased biomarker from the prioritization with CFG of discovery data from DE (AUC 0.55, P-value 0.28). The results across all diagnoses are modest, likely due to the significant variation of markers by diagnostic group (Table 5 and Supplementary Figure S4). This seems to be even more of an issue for trait than for state predictions. Within diagnostic groups, in bipolar disorder, the SAT1 prediction accuracy for future hospitalizations is higher (AUC 0.63, P-value 0.18), consistent with our previous work.1 CADM1 (AUC 0.72, P-value 0.076), SKA2 (AUC 0.71, P-value 0.056), and SLC4A4 (AUC 0.70, P-value 0.08) are even better predictors than SAT1 in bipolar disorder.
CFI-S has very good accuracy (AUC 0.89, P-value 3.53e−13) at predicting SI in psychiatric participants across diagnostic groups (Figure 4c). Within diagnostic groups, in affective disorders, the accuracy is even higher. CFI-S has excellent accuracy at predicting high SI in bipolar participants (AUC 0.97, P-value 1.75e−06) and in depression participants (AUC 0.95, P-value 7.98e−06). CFI-S has good accuracy (AUC 0.71, P-value 0.006) at predicting future hospitalizations for suicidality in the first year, across diagnostic groups.
SASS has very good accuracy (AUC 0.85, 9.96e−11) at predicting SI in psychiatric participants across diagnostic groups. Within diagnostic groups, in bipolar disorder, the accuracy is even higher (AUC 0.87, P-value 0.00011). SASS also has good accuracy (AUC 0.71, P-value 0.008) at predicting future hospitalizations for suicidality in the first year following testing.
Our a priori primary end point was a combined UP-Suicide, composed of the top increased and decreased biomarkers (n=11) from the discovery for ideation (CADM1, CLIP4, DTNA, KIF2C), prioritization with CFG for prior evidence (SAT1, SKA2, SLC4A4), and validation for behavior in suicide completers (IL6, MBP, JUN, KLHDC3) steps, along with CFI-S, and SASS. UP-Suicide is an excellent predictor of SI across all disorders in the independent cohort of psychiatric participants (AUC 0.92, P-value 7.94e−15) (Figure 6). UP-Suicide also has good predictive ability for future psychiatric hospitalizations for suicidality in the first year of follow-up (AUC 0.71, P-value 0.0094). The predictive ability of UP-Suicide is notably higher in affective disorder participants (bipolar, depression) (Table 5 and Figure 5).
Prediction of suicidal ideation by universal predictive measure-suicide. (a) (top left) Receiver-operating curve identifying participants with suicidal ideation against participants with no suicidal ideation or intermediate SI. (top right) Y axis contains the average UP-Suicide scores with standard error of mean for no suicidal ideation, intermediate suicidal ideation and high suicidal ideation. (bottom right) Scatter plot depicting HAMD-SI score on the Y axis and universal predictive measure-suicide score on the X axis with linear trend line. (bottom) Table summarizing descriptive statistics. Analysis of variance was performed between groups with no suicidal ideation, intermediate suicidal ideation and high suicidal ideation. (b) Predictions in test cohort based on thresholds in the discovery cohort - average UP-Suicide scores with standard deviation. (c) Number of participants correctly identified in the test cohort by categories based on thresholds in the discovery cohort. Category 1 means within 1 s.d. above the average of high suicidal ideation participants in the discovery cohort, category 2 means between 1 and 2 s.d. above, and so on. Category 1 means within 1 s.d. below the average of the no suicidal ideation participants in the discovery cohort, category 2 means between 1 and 2 s.d. below and so on.
Testing of universal predictor for suicide (UP-Suicide). UP-Suicide is a combination of our best gene expression biomarkers (top increased and decreased biomarkers from discovery, prioritization by CFG, and validation in suicide completers steps), and phenomic data (CFI-S and SASS). (a) Area Under the Curve (AUC) for the UP-Suicide predicting suicidal ideation and hospitalizations within the first year in all participants, as well as separately in bipolar (BP), major depressive disorder (MDD), schizophrenia (SZ), and schizoaffective (SZA) participants. **Indicates the comparison survived Bonferroni correction for multiple comparisons. *Indicates nominal significance of P<0.05. Bold outline indicates that the UP-Suicide was synergistic to its components, i.e., performed better than the gene expression biomarkers or phenomic data individually. (b) Table containing descriptive statistics for all participants together, as well as separately in BP, MDD, SZ, and SZA. Bold indicates the measure survived Bonferroni correction for 200 comparisons (20 genomic and phenomic markers/combinations × 2 testing cohorts for SI and future hospitalizations in the first year × 5 diagnostic categories–all, BP, MDD, SZA, SZ). We also show Pearson correlation data in the suicidal ideation test cohort for HAMD-SI vs. UP-Suicide, as well as Pearson correlation data in the hospitalization test cohort for frequency of hospitalizations for suicidality in the first year, and for frequency of hospitalizations for suicidality in all future available follow-up interval (which varies among participants, from 1 year to 8.5 years).
Discussion
We carried out systematic studies to identify clinically useful predictors for suicide. Our work focuses on identifying markers involved in SI and suicidal behavior, including suicide completion. Markers involved in behavior may be on a continuum with some of the markers involved in ideation, varying in the degree of expression changes from less severe (ideation) to more severe (behavior). One cannot have suicidal behavior without SI, but it may be possible to have SI without suicidal behavior.
As a first step, we sought to use a powerful but difficult to conduct within-participant design for discovery of blood biomarkers. Such a design is more informative than case-control, case-case, or even identical twins designs. The power of a within-participants longitudinal design for multi-omic discovery was first illustrated by Snyder and colleagues9 in a landmark paper with an n=19. We studied a cohort of male participants with major psychiatric disorders (n=217 participants) followed longitudinally (2–6 testing visits, at 3–6 months interval). In a smaller (n=37) but very valuable subset of these participants, we captured one or more major switches from a no SI state to a high SI state at the time of the different testing visits (Figures 1 and 2).
Second, we conducted whole-genome gene expression discovery studies in the participants that exhibited the switches, using a longitudinal within-participant design, that factors out genetic variability and reduces environmental variability as well. We have demonstrated the power of such a design in our previous work on suicide biomarkers with an n=91. Our current n=37 was four-fold higher, and consequently our power to detect signal was commensurately increased (Figure 2). Genes whose levels of expression tracked SI within each participant were identified.
Third, the lists of top candidate biomarkers for SI from the discovery and prioritization step (genes with a CFG score of 4 and above, reflecting genes that have maximal experimental internal evidence from this study and/or additional external literature cross-validating evidence), were additionally validated for involvement in suicidal behavior in a cohort of demographically matched suicide completers from the coroner’s office (n=26) (Figure 3).
Given that we used two methods (AP, DE), three steps (discovery for ideation, prioritization based on literature evidence, validation for behavior in completers), and two types of markers (increased, decreased), we anticipated having 2 × 3 × 2 =12 top markers. We ended up with 11 due to overlap (Table 2). Of note, 8 of these 11 markers (SAT1, SKA2, SLC4A4, KIF2C, MBP, IL6, JUN and KLHDC3), were significant in validation for behavior in terms of being changed even more in suicide completers, and 5 of them survived Bonferroni correction (SAT1, SLC4A4, MBP, IL6, KLHDC3). The 3 out of 11 markers that were not validated for behavior (DTNA, CLIP4 and CADM1) seemed indeed better in the independent test cohorts at predicting SI than at predicting suicidal behavior (hospitalizations) (Table 5B).
Fourth, we describe a novel, simple and comprehensive phenomic (clinical) risk assessment scale, the CFI-S scale, as well as a companion app to it for use by clinicians and individuals (Supplementary Figure S2). CFI-S was developed independently of any data from this study, by integrating known risk factors for suicide from the clinical literature. It has a total of 20 items (scored in a binary fashion—1 for present, 0 for absent, NA for information not available) that assess the influence of mental health factors, as well as of life satisfaction, physical health, environmental stress, addictions, and cultural factors known to influence suicidal behavior. It also has two demographics risk factors items: age and gender. The result is a simple polyphenic risk score with an absolute range of 0–22, normalized by the number of items on which we had available information, resulting in a score in the range from 0 to 1 (Table 4). We present data validating the CFI-S in our discovery cohort of live psychiatric participants and in suicide completers from the coroner’s office (Figure 4). We acknowledge the possibility of a potential upward bias in next-of-kin reporting post-suicide completion, although each item of the scale was scored factually by a trained rater on its own merits. We believe it is still illustrative and informative to compare the CFI-S in live participants with ideation vs suicide completers, and identify which items are most different (such as inability to cope with stress, which is consistent with biological data from the biomarker side of our study).
Fifth, we have also assessed anxiety and mood, using a visual analog SASS, previously described by us (Niculescu et al. 2006), for which we now have developed an app version (Supplementary Figure S2). Using a PhenoChipping approach6 in our discovery cohort of psychiatric participants, we show that anxiety measures cluster with SI and CFI-S, and mood measures are in the opposite cluster, suggesting that our participants have high SI when they have high anxiety and low mood (Figure 2). We would also like to include in the future measures of psychosis, and of stress, to be more comprehensive.
Sixth, we examined how the biomarkers identified by us are able to predict state (SI) in a larger independent cohort of psychiatric participants (n= 108 participants).
Seventh, we examined whether the biomarkers are able to predict trait (future hospitalizations for suicidal behavior) in psychiatric participants (n=157) in the short term (first year of follow-up) as well as overall (all data for future hospitalizations available for each patient).
Last but not least, we demonstrate how our a priori primary end point, a comprehensive UP-Suicide, composed of the combination of the top increased and decreased biomarkers (n=11) from the discovery, prioritization and validation steps, along with CFI-S and SASS, predicts state (SI) and trait (future psychiatric hospitalizations for suicidality).
The rationale for identifying blood biomarkers as opposed to brain biomarkers is a pragmatic one—the brain cannot be readily accessed in live individuals. Other peripheral fluids, such as cerebrospinal fluid, require more invasive and painful procedures. Nevertheless, it is likely that many of the peripheral blood transcriptomic changes are not necessarily mirroring what is happening in the brain, and vice-versa. The keys to finding peripheral biomarkers4 are, first, to have a powerful discovery approach, such as our within-participant design, that closely tracks the phenotype you are trying to measure and reduces noise. Second, cross-validating and prioritizing the results with other lines of evidence, such as brain gene expression and genetic data, are important in order to establish relevance and generalizability of findings. Third, it is important to validate for behavior in an independent cohort with a robust and relevant phenotype, in this case suicide completers. Fourth, testing for predictive ability in independent/prospective cohorts is a must.
Biomarkers that survive such a rigorous stepwise discovery, prioritization, validation and testing process are likely directly relevant to the disorder studied. As such, we endeavored to study their biology, whether they are involved in other psychiatric disorders or are relatively specific for suicide, and whether they are the modulated by existing drugs in general, and drugs known to treat suicidality in particular. We have identified a series of biomarkers that seem to be changed in opposite direction in suicide vs in treatments with omega-3 fatty acids, lithium, clozapine or MAOIs. These biomarkers could potentially be used to stratify patients to different treatment approaches, and monitor their response (Supplementary Table S4).
We also conducted predictive studies, across all participants and by diagnosis, as a way of assessing how generalizable and how particular to a diagnosis biomarkers are. Different diagnostic groups have different disease biology and are on different medications, which may modify the levels of the biomarkers. We observe a significant variation in the predictive ability of biomarkers by diagnosis, which has important practical applications for future work on diagnostic-specific predictors (Table 5). Of note, a number of biomarkers from the current larger study reproduce our previous work in a smaller, bipolar cohort (SAT1, MARCKS, PTEN, as well as FOXN3, GCOM1, RECK, IL1B, LHFP, ATP6V0E1 and KLHDC3) (Supplementary Table S2). In the current data sets, we have also post hoc carried out biomarker discovery within each diagnosis, which revealed a diversity of top markers, but should be interpreted with caution given the smaller N within each diagnostic group (Supplementary Table S5).
Before any testing, we planned to use a comprehensive combination of genomic data (specifically, the top increased and decreased biomarkers from discovery, prioritization and validation) and phenomic data (specifically, the CFI-S and the SASS) as the primary end point measure, a broad-spectrum universal predictor (UP-Suicide) for state SI and trait future hospitalizations. It has not escaped our attention that certain single biomarkers, particular phenotypic items, or combinations thereof seem to perform better than the UP-Suicide in one or another type of prediction or diagnostic group (see Table 5). However, since such markers and combinations were not chosen by us a priori and such insights derive from testing, we cannot exclude a fit to cohort effect for them and reserve judgement as to their robustness as predictors until further testing in additional independent cohorts, by us and others. What we can put forward for now based on the current work is the UP-Suicide, which seems to be a robust predictor across different scenarios and diagnostic groups.
Overall, our predictive ability for trait future hospitalizations is somewhat less than for state SI (Figure 5, Table 5). However, clinically, events may indeed be driven by state, and the immediate concern is preventing immediate or short term adverse outcomes.
Our study has a number of limitations. All this work was carried out in psychiatric patients, a high-risk group, and it remains to be seen how such predictors apply to non-psychiatric participants. Additionally, the current studies were carried out exclusively in males. Similar work is needed in females, with and without psychiatric disorders. Such work is ongoing in our group. Lastly, for the UP-Suicide testing, the prevalence rate for high SI in our independent test cohort was a relatively low 21% (23 out of 108), and the incidence of future hospitalizations for suicidality was even lower: 7.6% in the first year (12 out of 157), and 21.0% overall (33 out of 157) (Figure 5). Although this is fortunate for the participants enrolled and may reflect the excellence of clinical care they were receiving in our hospital independent of this study, it may bias the predictions. Studies with larger numbers and longer follow-up, currently ongoing, as well as studies in different clinical settings, may provide more generalizability. It is to be noted, however, that the incidence of suicidality in the general population is lower, for example at 1.5% in adolescents in an European cohort23 and estimates of 0.2–2% in the US,24 which underlines the rationale of using a very high-risk group like we did for magnifying and enabling signal detection with a relatively small N.
In conclusion, we have advanced the biological understanding of suicidality, highlighting behavioral and biological mechanisms related to inflammation, mTOR signaling, growth factors, stress response and apoptosis. mTOR signaling has been identified as necessary for the rapid antidepressant response of ketamine.25 The fact that this biological pathway was identified in an unbiased fashion by our work as the top pathway changed in suicide in the validated biomarkers from our analyses (Table 3 and Supplementary Figure S3) is scientifically interesting, and provides a biological rationale for studying ketamine as a potential treatment in acutely suicidal individuals.26 Of equal importance, we developed instruments (biomarkers and apps) for predicting suicidality, that do not require asking the person assessed if they have suicidal thoughts, as individuals who are truly suicidal often do not share that information with people close to them or with clinicians. We propose that the widespread use of such risk prediction tests as part of routine or targeted healthcare assessments will lead to early disease interception followed by preventive lifestyle modifications or treatment. Given the magnitude and urgency of the problem, the importance of efforts to implement such tools cannot be overstated.
References
Le-Niculescu H, Levey DF, Ayalew M, Palmer L, Gavrin LM, Jain N et al. Discovery and validation of blood biomarkers for suicidality. Mol Psychiatry 2013; 18: 1249–1264.
Le-Niculescu H, Kurian SM, Yehyawi N, Dike C, Patel SD, Edenberg HJ et al. Identifying blood biomarkers for mood disorders using convergent functional genomics. Mol Psychiatry 2009; 14: 156–174.
Kurian SM, Le-Niculescu H, Patel SD, Bertram D, Davis J, Dike C et al. Identification of blood biomarkers for psychosis using convergent functional genomics. Mol Psychiatry 2011; 16: 37–58.
Niculescu AB, Levey D, Le-Niculescu H, Niculescu E, Kurian SM, Salomon D . Psychiatric blood biomarkers: avoiding jumping to premature negative or positive conclusions. Mol Psychiatry 2015; 20: 286–288.
Niculescu AB, Le-Niculescu H . Convergent Functional Genomics: what we have learned and can learn about genes, pathways, and mechanisms. Neuropsychopharmacology 2010; 35: 355–356.
Niculescu AB, Lulow LL, Ogden CA, Le-Niculescu H, Salomon DR, Schork NJ et al. PhenoChipping of psychotic disorders: a novel approach for deconstructing and quantitating psychiatric phenotypes. Am J Med Genet B, Neuropsychiatr Genet 2006; 141B: 653–662.
Borges G, Angst J, Nock MK, Ruscio AM, Kessler RC . Risk factors for the incidence and persistence of suicide-related outcomes: a 10-year follow-up study using the National Comorbidity Surveys. J Affect Disord 2008; 105: 25–33.
Nock MK . Future directions for the study of suicide and self-injury. J Clin Child Adolesc Psychol 2012; 41: 255–259.
Chen R, Mias GI, Li-Pook-Than J, Jiang L, Lam HY, Miriami E et al. Personal omics profiling reveals dynamic molecular and medical phenotypes. Cell 2012; 148: 1293–1307.
Darlington TM, Pimentel R, Smith K, Bakian AV, Jerominski L, Cardon J et al. Identifying rare variants for genetic risk through a combined pedigree and phenotype approach: application to suicide and asthma. Transl Psychiatry 2014; 4: e471.
Sequeira A, Morgan L, Walsh DM, Cartagena PM, Choudary P, Li J et al. Gene expression changes in the prefrontal cortex, anterior cingulate cortex and nucleus accumbens of mood disorders subjects that committed suicide. PLoS One 2012; 7: e35367.
Klempan TA, Sequeira A, Canetti L, Lalovic A, Ernst C, ffrench-Mullen J et al. Altered expression of genes involved in ATP biosynthesis and GABAergic neurotransmission in the ventral prefrontal cortex of suicides with and without major depression. Mol Psychiatry 2009; 14: 175–189.
Hishimoto A, Cui H, Mouri K, Nushida H, Ueno Y, Maeda K et al. A functional polymorphism of the micro-opioid receptor gene is associated with completed suicides. J Neural Transm 2008; 115: 531–536.
Gabilondo AM, Meana JJ, Garcia-Sevilla JA . Increased density of mu-opioid receptors in the postmortem brain of suicide victims. Brain Res 1995; 682: 245–250.
Guintivano J, Brown T, Newcomer A, Jones M, Cox O, Maher BS et al. Identification and replication of a combined epigenetic and genetic biomarker predicting suicide and suicidal behaviors. Am J Psychiatry 2014; 171: 1287–1296.
Mann JJ, Currier D . Medication in suicide prevention insights from neurobiology of suicidal behavior. In: Dwivedi Y (ed) The Neurobiological Basis of Suicide. CRC Boca Raton (FL), 2012.
Sublette ME, Hibbeln JR, Galfalvy H, Oquendo MA, Mann JJ . Omega-3 polyunsaturated essential fatty acid status as a predictor of future suicide risk. Am J Psychiatry 2006; 163: 1100–1102.
Oquendo MA, Sullivan GM, Sudol K, Baca-Garcia E, Stanley BH, Sublette ME et al. Toward a biosignature for suicide. Am J Psychiatry 2014; 171: 1259–1277.
Kelleher I, Lynch F, Harley M, Molloy C, Roddy S, Fitzpatrick C et al. Psychotic symptoms in adolescence index risk for suicidal behavior: findings from 2 population-based case-control clinical interview studies. Arch Gen Psychiatry 2012; 69: 1277–1283.
Schoenbaum M, Kessler RC, Gilman SE, Colpe LJ, Heeringa SG, Stein MB et al. Predictors of suicide and accident death in the Army Study to Assess Risk and Resilience in Servicemembers (Army STARRS): results from the Army Study to Assess Risk and Resilience in Servicemembers (Army STARRS). JAMA Psychiatry 2014; 71: 493–503.
Posner K, Brown GK, Stanley B, Brent DA, Yershova KV, Oquendo MA et al. The Columbia-Suicide Severity Rating Scale: initial validity and internal consistency findings from three multisite studies with adolescents and adults. Am J Psychiatry 2011; 168: 1266–1277.
Wemmie JA, Taugher RJ, Kreple CJ . Acid-sensing ion channels in pain and disease. Nat Rev Neuroscience 2013; 14: 461–471.
Wasserman D, Hoven CW, Wasserman C, Wall M, Eisenberg R, Hadlaczky G et al. School-based suicide prevention programmes: the SEYLE cluster-randomised, controlled trial. Lancet 2015; 385: 1536–1544.
Nock MK, Borges G, Bromet EJ, Cha CB, Kessler RC, Lee S . Suicide and suicidal behavior. Epidemiol Rev 2008; 30: 133–154.
Li N, Lee B, Liu RJ, Banasr M, Dwyer JM, Iwata M et al. mTOR-dependent synapse formation underlies the rapid antidepressant effects of NMDA antagonists. Science 2010; 329: 959–964.
Ballard ED, Ionescu DF, Vande Voort JL, Niciu MJ, Richards EM, Luckenbaugh DA et al. Improvement in suicidal ideation after ketamine infusion: relationship to reductions in depression and anxiety. J Psychiatr Res 2014; 58: 161–166.
Pandey GN, Rizavi HS, Ren X, Fareed J, Hoppensteadt DA, Roberts RC et al. Proinflammatory cytokines in the prefrontal cortex of teenage suicide victims. J Psychiatr Res 2012; 46: 57–63.
Hoyo-Becerra C, Huebener A, Trippler M, Lutterbeck M, Liu ZJ, Truebner K et al. Concomitant interferon alpha stimulation and TLR3 activation induces neuronal expression of depression-related genes that are elevated in the brain of suicidal persons. PLoS One 2013; 8: e83149.
Lindqvist D, Janelidze S, Erhardt S, Traskman-Bendz L, Engstrom G, Brundin L . CSF biomarkers in suicide attempters—a principal component analysis. Acta Psychiatr Scand 2011; 124: 52–61.
Bay-Richter C, Linderholm KR, Lim CK, Samuelsson M, Traskman-Bendz L, Guillemin GJ et al. A role for inflammatory metabolites as modulators of the glutamate N-methyl-D-aspartate receptor in depression and suicidality. Brain Behav Immun 2015; 43: 110–117.
Kim YK, Lee SW, Kim SH, Shim SH, Han SW, Choi SH et al. Differences in cytokines between non-suicidal patients and suicidal patients in major depression. Prog Neuropsychopharmacol Biol Psychiatry 2008; 32: 356–361.
Fiori LM, Wanner B, Jomphe V, Croteau J, Vitaro F, Tremblay RE et al. Association of polyaminergic loci with anxiety, mood disorders, and attempted suicide. PloS One 2010; 5: e15146.
Sokolowski M, Ben-Efraim YJ, Wasserman J, Wasserman D . Glutamatergic GRIN2B and polyaminergic ODC1 genes in suicide attempts: associations and gene-environment interactions with childhood/adolescent physical assault. Mol Psychiatry 2013; 18: 985–992.
Perlis RH, Huang J, Purcell S, Fava M, Rush AJ, Sullivan PF et al. Genome-wide association study of suicide attempts in mood disorder patients. Am J Psychiatry 2010; 167: 1499–1507.
Kim S, Choi KH, Baykiz AF, Gershenfeld HK . Suicide candidate genes associated with bipolar disorder and schizophrenia: An exploratory gene expression profiling analysis of post-mortem prefrontal cortex. BMC Genomics 2007; 8: 413.
Labonte B, Suderman M, Maussion G, Lopez JP, Navarro-Sanchez L, Yerko V et al. Genome-wide methylation changes in the brains of suicide completers. Am J Psychiatry 2013; 170: 511–520.
Seder E, Biselli A, Pisano S, Niccolai S, Smith GD, Joo K et al. Longitudinal target-spin asymmetries for deeply virtual compton scattering. Phys Rev Lett 2015; 114: 032001.
Roggenbach J, Muller-Oerlinghausen B, Franke L, Uebelhack R, Blank S, Ahrens B . Peripheral serotonergic markers in acutely suicidal patients. 1. Comparison of serotonergic platelet measures between suicidal individuals, nonsuicidal patients with major depression and healthy subjects. J Neural Transm 2007; 114: 479–487.
Pandey GN, Dwivedi Y, Ren X, Rizavi HS, Roberts RC, Conley RR et al. Altered expression and phosphorylation of myristoylated alanine-rich C kinase substrate (MARCKS) in postmortem brain of suicide victims with or without depression. J Psychiatr Res 2003; 37: 421–432.
Punzi G, Ursini G, Shin JH, Kleinman JE, Hyde TM, Weinberger DR . Increased expression of MARCKS in post-mortem brain of violent suicide completers is related to transcription of a long, noncoding, antisense RNA. Mol Psychiatry 2014; 19: 1057–1059.
Dwivedi Y, Rizavi HS, Zhang H, Roberts RC, Conley RR, Pandey GN . Modulation in activation and expression of phosphatase and tensin homolog on chromosome ten, Akt1, and 3-phosphoinositide-dependent kinase 1: further evidence demonstrating altered phosphoinositide 3-kinase signaling in postmortem brain of suicide subjects. Biol Psychiatry 2010; 67: 1017–1025.
Karege F, Perroud N, Burkhardt S, Fernandez R, Ballmann E, La Harpe R et al. Alterations in phosphatidylinositol 3-kinase activity and PTEN phosphatase in the prefrontal cortex of depressed suicide victims. Neuropsychobiology 2011; 63: 224–231.
Acknowledgements
This work is, in essence, a field-wide collaboration. We acknowledge our debt of gratitude for the efforts and results of the many other groups, cited in our paper, who have conducted and published studies (clinical, genetic and biological) in suicidality. With their arduous and careful work, a convergent approach such as ours is possible. We would particularly like to thank the participants who participated in these studies, their families and their caregivers. Without their contribution, such work to advance the understanding of suicide would not be possible. This work was supported by an NIH Directors’ New Innovator Award (1DP2OD007363) and a VA Merit Award (2I01CX000139) to ABN. Supplementary Information is also available from the Niculescu Laboratory website (www.neurophenomics.info).
Author contributions
ABN designed the study, created the clinical rating scales and wrote the manuscript. DFL, PLP, HL-N, HD, NJ, TBL, RL and EMN analyzed the data. NJ, NPV and FNK performed database work. PLP, JM and GS produced the apps. EB, AJ, SG, HW, DLG and RS organized and conducted testing in psychiatric participants. SC, CH, AB, MY, AS, GES and ABN organized and carried out post-mortem samples collection. TG, NJS, SMK and DRS conducted microarray experiments and provided input on data analyses. All authors discussed the results and commented on the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
ABN is listed as inventor on a patent application being filed by Indiana University. The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Molecular Psychiatry website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Niculescu, A., Levey, D., Phalen, P. et al. Understanding and predicting suicidality using a combined genomic and clinical risk assessment approach. Mol Psychiatry 20, 1266–1285 (2015). https://doi.org/10.1038/mp.2015.112
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/mp.2015.112
This article is cited by
-
Towards precision medicine for anxiety disorders: objective assessment, risk prediction, pharmacogenomics, and repurposed drugs
Molecular Psychiatry (2023)
-
Precision medicine in psychiatry: biomarkers to the forefront
Neuropsychopharmacology (2022)
-
Identification of potential blood biomarkers associated with suicide in major depressive disorder
Translational Psychiatry (2022)
-
Polyphenic risk score shows robust predictive ability for long-term future suicidality
Discover Mental Health (2022)
-
Cerebrospinal fluid concentration of complement component 4A is increased in first episode schizophrenia
Nature Communications (2022)