Abstract
Recent studies have revealed the epigenetic alterations are involved in hepatocarcinogenesis. However, the function of long interspersed nuclear element-1 hypomethylation in hepatocellular carcinomas, and relationship among other clinicopathologic features, and genetic and epigenetic alterations, including CpG island hypermethylation, have not been studied. We determined long interspersed nuclear element-1 methylation, a marker of global methylation, in 57 tumor and nonneoplastic samples, including 24 from high-risk and 33 from low-risk countries. We compared methylation levels of long interspersed nuclear element-1 with eight CpG islands including p16, cyclooxygenase-2, T-type calcium channel, and estrogen receptor genes, and MINT31, MINT1, MINT2, and MINT27, as well as CpG island methylator phenotype and p53 gene mutation. Most hepatocellular carcinomas samples (88%) showed hypomethylation of long interspersed nuclear element-1, with a mean level of global methylation of 58±14 compared to 77±6 in nonneoplastic hepatic tissue (P<0.001). Levels of long interspersed nuclear element-1 hypomethylation differed depending on geographic location (P=0.02), status of hepatitis (P=0.01), hypermethylation of p16, estrogen receptor and MINT2 (P=0.01, 0.002, and 0.045, respectively), CpG island methylator phenotype-positive status (P=0.006), and p53 gene mutation (P=0.04). In conclusion, environmental factors such as geographic location and hepatitis status contribute to hepatocarcinogenesis through global hypomethylation. In hepatocellular carcinomas, hypermethylation of CpG islands, and CpG island methylator phenotype status seems to correlate with levels of long interspersed nuclear element-1 hypomethylation.
Similar content being viewed by others
Main
Hepatocellular carcinoma is one of the most common malignancies in the world and a leading cause of death worldwide. The incidence rate of hepatocellular carcinoma is associated with various risk factors including geographic location and race.1 Hepatocarcinogenesis is a complex, multistep process evolving from normal liver through chronic hepatitis and cirrhosis to hepatocellular carcinoma, but hepatocellular carcinoma also can arise from noncirrhotic liver. This process is known to be influenced by multiple genetic changes such as allelic deletions, chromosomal losses and gains, and DNA rearrangement, and epigenetic alterations.2, 3 Frequent promoter hypermethylation and subsequent loss of protein expression of tumor suppressor genes has also been demonstrated in hepatocellular carcinoma.4
The long interspersed nuclear element (LINE-1), 6 kb in size, is an endogenous mobile repetitive sequence in the genome that constitutes 21% of human genome.5, 6 The level of LINE-1 methylation is thought to represent the amount of global methylation because most 5-methylcytosines lie within the repetitive sequences. CpG sites in a LINE-1 are heavily methylated in the normal state and hypomethylation of LINE promoter might lead to increased expression of LINE element and probably to retrotransposition.5, 6 Retrotransposition, in turn, leads to chromosomal rearrangement. Hypomethylation of LINE-1 is associated with genomic instability and to clinicopathologic features in a variety of tumors.7, 8, 9 Genome-wide hypomethylation has been reported in variety of cancers including hepatocellular carcinomas,10, 11, 12 and a few studies reported global hypomethylation as a poor prognostic marker in hepatocellular carcinoma4, 13 but LINE-1 methylation has not been studied in detail.
We have previously shown that CpG island methylation differs in hepatocellular carcinoma from different geographical regions.15 In this study, we studied if global hypomethylation as assessed by LINE-1 methylation differs in hepatocellular carcinoma from different geographical regions, and is it associated with CpG island methylation. We determined LINE-1 methylation by pyrosequencing from hepatocellular carcinoma and nontumor liver parenchyma and compared the levels of LINE-1 hypomethylation with clinicopathologic features, and methylation status of CpG islands, and CpG island methylator phenotype (CIMP) status.16
Materials and methods
Characteristics of Cell Lines, Specimens, and Patients
Frozen tumor and corresponding nonneoplastic hepatic tissue were obtained from surgical specimens of 57 patients undergoing curative resection for hepatocellular carcinoma,14 including 24 cases from China and Egypt, and 33 from The Johns Hopkins Hospital, Baltimore, MD, USA, respectively. Surveillance Committee (Institutional Review Board) approved this study. Cell lines, including RKO, SW48, and SW480 colon cancer cell lines and Hep3B liver cancer cell lines were grown in recommended culture medium and genomic DNA was extracted.
The patient records were reviewed for clinical information including history of alcohol or other drugs, serologic evidence for hepatitis virus, survival status and follow-up months, and histological parameters were evaluated from the hematoxylin and eosin (H&E)-stained slides. Clinicopathologic parameters evaluated in each case included patient age at initial diagnosis, gender, and tumor stage. This information was unavailable for 24 cases from high-risk countries.
Tissue Preparation and DNA Extraction
DNA from both tumor and nonneoplastic hepatic tissue in microdissected fresh-frozen specimens was extracted using a commercial kit (Qiagen DNA Extraction Kit; Qiagen Inc., Valencia, CA, USA), after H&E-stained slides from a frozen block were reviewed. The tumor cellularity was at least 70% in all samples. All the tumors were typical hepatocellular carcinomas, and fibrolamellar and mixed hepatocellular-cholangiocarcinomas were excluded.
Bisulfite-Pyrosequencing of LINE-1
Bisulfite treatment of DNA was performed as described previously.14 The LINE-1 methylation was quantified by pyrosequencing using the primers and conditions as previously described.16, 17 LINE-1 assay was performed using 10 pmol of forward primer 5′-TTTTGAGTTAGGTGTGGGATATA-3′ and 10 pmol of reverse biotinylated primer 5′-AAAATCAAAAAATTCCCTTTC-3′. The LINE-1 assay was performed in a 25 μl PCR reaction containing 40 ng of bisulfite-treated genomic DNA, 2 mM MgCl2, 0.8 mM dNTP mix, and 1.25 U of Taq polymerase. PCR cycling conditions were 94°C for 30 s, 56°C for 30 s, and 72°C for 30 s for 50 cycles. The PCR product was purified and quantified using the PSQ HS 96 Pyrosequencing System (Pyrosequencing Inc., Westborough, MA, USA). The ratio of C to T nucleotides was evaluated for LINE-1 methylation (examples are shown in Figure 1). DNA from RKO, SW48, and SW480, and Hep3B cell lines as well as SssI-treated DNA were used as controls for optimizing the assay. Deionized water was used as a negative control. The experiments were repeated twice and mean values were taken from these separate experiments. Methylation levels at three consecutive CpG sites were evaluated. Mean methylation levels at the three consecutive CpG sites were statistically correlated with the methylation levels at the first CPG site (data not shown), and methylation levels at the first CpG site were used for all comparisons. This assay has been previously characterized and has been shown to correlate with global DNA methylation measured by other methodologies.16, 17
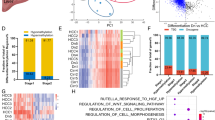
Quantification of LINE-1 methylation levels by pyrosequencing of bisulfite-treated DNA samples. The top panel shows methylation of Hep3b cell line, the middle panel of a nonneoplastic liver parenchyma, and the lower panel from the corresponding hepatocellular carcinoma. The PCR product was purified and methylation was quantified using the PSQ HS 96 Pyrosequencing System (Pyrosequencing Inc.). The program quantifies C or G nucleotides for methylated and A or T for unmethylated or mutated sequences. The shaded regions represent CpG sites used for quantification of methylation at each site, and the percent methylation at each site is shown above the peaks.
The mean LINE-1 methylation in SssI-treated normal samples including placental DNA and normal samples from patients was 85±4% (range 82–97). LINE-1 relative tumor hypomethylation was calculated by subtracting LINE-1 level of patient's nonneoplastic liver sample from the LINE-1 methylation level of patient's tumor sample.
Sequencing of p53 Gene and Combined Bisulfite PCR and Restriction Analysis
The p53 gene mutations and methylation status of eight CpG islands have been previously reported,14 including p16, cyclooxygenase-2 (COX-2), T-type calcium channel (CACNA1G), and estrogen receptor (ER) genes; and MINT31, MINT1, MINT2, and MINT27. A CpG island was considered methylated if methylation density was 15% or more. The tumors were classified CIMP-negative if no CpG island was methylated, CIMP-intermediate if one or two CpG islands were methylated, and CIMP-high if three or more CpG islands were methylated.14, 15
Statistical Analysis
All statistical analyses were performed using SPSS (SPSS Inc., Chicago, IL, USA). Comparisons of categorical variables were made using χ2-test and Fisher's exact test. Comparisons of methylation densities and clinicopathologic features were evaluated by ANOVA test and Student's t-test. All P-values presented are two-sided, and a P-value of less than 0.05 was considered statistically significant.
Results
The selected clinicopathologic features of 57 patients with hepatocellular carcinomas in this study are tabulated in Table 1. As reported previously,14 all the patients from high-risk countries had hepatitis B infection compared to 21% from the low-risk country (P=0.001) and p53 gene mutations were present in 50% of tumors from high-risk countries but in 12% of tumors from low-risk countries (P=0.004).
All cell lines were hypomethylated compared to Sss1-treated normal samples. RKO, SW48, SW480, and Hep3B cell lines had LINE-1 methylation levels of 69, 70, 45, and 70%, respectively. Levels of LINE-1 methylation in normal and tumor samples are shown in Figure 2 and Table 2. Majority of tumor samples (88%) were less methylated compared to corresponding nonneoplastic hepatic tissue at LINE-1. The mean LINE-1 methylation in tumor samples was 58±14 compared with 77±6 in nonneoplastic liver parenchyma samples (P=0.001).
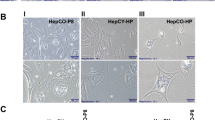
LINE-1 methylation levels in hepatocellular carcinoma samples and corresponding nonneoplastic liver parenchyma samples from high-risk and low-risk countries. The nonneoplastic liver parenchyma sample is above or below the tumor sample from the same patient. The levels of LINE-1 hypomethylation in tumor samples from high-risk country are more than that of tumor samples from low-risk country. There is no significant difference in LINE-1 hypomethylation between nonneoplastic liver parenchyma samples from high-risk and low-risk countries.
The levels of hypomethylation differed according to geographic location of the samples and hepatitis status (Figure 2; Table 2). The mean LINE-1 methylation in tumor samples from high-risk country was lower than that of tumor samples from low-risk country (53 vs 62, P=0.02), but there was no statistical difference in nonneoplastic samples between samples from high-risk and low-risk countries (P=0.8). The mean LINE-1 methylation in tumor samples was lower in hepatitis-positive cases than in hepatitis-negative cases (P=0.01). In patients from low-risk country, the mean LINE-1 methylation in tumor samples was lower in hepatitis-positive cases than in hepatitis-negative cases but was not statistically significant (P=0.26).
Levels of LINE-1 methylation were associated with p53 gene mutations and methylation of several CpG islands and genes (Table 2). Levels of LINE-1 hypomethylation in tumors associated with p53 gene mutation (P=0.04). In patients from low-risk country and high-risk country, the mean LINE-1 methylation in tumor samples with p53 gene mutation were lower than in tumor samples without mutations but were not statistically significant (P=0.39 and 0.24, respectively). Levels of LINE-1 hypomethylation in nontumor liver tissue associated with methylation of CACNAIG (P=0.007), COX-2 (P=0.002), and MINT-1 (P=0.02). In contrast, hypomethylation of tumor correlated with methylation of p16 (P=0.01), ER (P=0.002), MINT-2 (P=0.045), and CIMP status (P=0.006).
The mean relative tumor hypomethylation for LINE-1 was 19±15. Relative tumor hypomethylation levels were associated with multiple clinicopathologic features, and genetic and epigenetic alterations (Table 3), including geographic location (P=0.02), hepatitis status (P=0.02), grade of tumor (P=0.02), ER methylation (P=0.001), COX-2 methylation (P=0.003), CIMP status (P=0.04), and p53 gene mutation status (P=0.03).
Discussion
We studied LINE-1 hypomethylation in hepatocellular carcinomas and nonneoplastic liver parenchyma. We found levels of LINE-1 methylation in tumors were lower in patients from high-risk country, who had hepatitis, and patients whose tumor had p53 gene mutation, and methylation of p16 and ER genes, MINT-2 CpG island, and CIMP positive. In contrast, levels of LINE-1 methylation in nonneoplastic hepatic parenchyma were lower in patients whose tumors had methylation of CACNAIG and COX-2 genes, and MINT-1 CpG island.
Aberrant methylation, consisting of DNA hypomethylation and/or promoter gene CpG hypermethylation, is implicated in the development of a variety of solid tumors. Global loss of methylation in cancer may lead to alterations in the expression of protooncogenes critical to carcinogenesis and may facilitate chromosomal instability, whereas promoter CpG island hypermethylation can lead to transcriptional repression of tumor suppressor genes.18, 19, 20 Previous studies have described global genomic hypomethylation in the several malignancies including carcinomas of ovary,21, 22 colon,17, 23, 24 and prostate25, 26 as well as in hepatocellular carcinomas4, 11, 12 by evaluating genomic repetitive sequences.
In this study and as reported previously,4, 11, 12 most hepatocellular carcinoma samples had global hypomethylation compared to nonneoplastic hepatic tissue. The methylation level of LINE-1 in hepatocellular carcinoma differed significantly depending on hepatitis status and geographic location. As expected, the prevalence of hepatitis B viral infection depends on the geographic location and is much higher in high-risk countries compared to the United States. The association between global methylation and hepatitis status can be explained by hepatitis B virus X protein that can induce altered DNA methyltransferase activity, hypermethylation of specific CpG islands, and global hypomethylation.27, 28 Alternatively, these changes may be due to increased proliferation because of liver injury, as has been demonstrated in a rat model of drug-induced liver injury.29 These findings are further corroborated by our previous study which showed hepatocellular carcinoma from patients with hepatitis or from patients living in high-risk geographic areas have much higher levels of CpG island methylation than the tumors from patients without hepatitis or low-risk areas.14 However, there is an overlap in LINE-1 methylation levels in nonneoplastic liver parenchyma and hepatocellular carcinoma samples from low-risk and high-risk countries suggesting individual variation and heterogeneity in methylation levels in both population. One limitation of this study is that the demographic and certain clinicopathologic features are missing for about 40% of the patients included in this study.
In our study and as previously reported,10 levels of LINE-1 methylation in nonneoplastic liver parenchyma were not associated with the status of hepatitis, cirrhosis or geographical location of samples. Hypomethylation is present at different steps of carcinogenesis in different tumor sites. For example, ovarian tumors showed a stepwise increase in global hypomethylation among benign cystadenomas, low malignant potential tumors and carcinomas.21 In contrast, hypomethylation was present at an earlier stage of carcinogenesis in colorectum30 and stomach31 and was present in colorectal hyperplastic polyps and adenomas and in atrophic gastritis that are considered precursors of invasive carcinomas. These data suggest that hypomethylation is a late event in hepatocarcinogenesis and raise the possibility of utilizing LINE-1 methylation-based assays as potential biomarkers for detection of hepatocellular carcinoma, especially in high-risk countries, as has been reported previously.10, 11 Assessment of LINE-1 methylation in recognized precursors of hepatocellular carcinomas, such as dysplastic nodules, may help in further understanding hepatocarcinogenesis.
In this study, LINE-1 methylation was lower in CIMP-positive tumors and associated with methylation of p16 and ER genes, and MINT-2 CpG island but not with methylation of other genes or CpG islands. In contrast, in colorectal cancers LINE-1 hypomethylation was inversely correlated with CIMP-positive tumors,24 especially CIMP-positive, microsatellite instability-high tumors due to methylation of hMLH1,17 and in prostatic adenocarcinomas LINE-1 hypomethylation was associated with methylation of CpG islands in a variety of genes.26 This suggests that associations among LINE-1 methylation in tumors, CpG island methylation and other genetic and phenotypic features are site specific. Although, these differences between CIMP status and LINE-1 hypomethylation among tumors from different sites may be in part due to different markers of methylation, methodology and criteria used for assessing methylation and CIMP in different studies.
Global hypomethylation is a marker of poor prognosis in several cancers such as chronic myeloid leukemia,32 breast cancer,33 and squamous cell carcinoma of head and neck,34 as well as in hepatocellular carcinoma.4, 11, 13 In hepatocarcinogenesis, global hypomethylation may be promoting progression via chromosomal instability, activation of protooncogenes, and reactivation of transposable elements.4 In patients with hepatocellular carcinoma, serum and tumor LINE-1 hypomethylation is considered a poor prognostic marker involved in the tumor progression and aggressive tumor behavior based on the association with high tumor grade and poor survival rate.4, 13 Recently, it has been reported that the levels of serum LINE-1 hypomethylation correlated significantly with HBsAg status, large tumor size, and advanced tumor stage and as an independent poor prognostic marker in patients who had hepatocellular carcinoma.35
It is still questionable whether gene-specific hypermethylation and global hypomethylation are independent processes or not, however, the present study demonstrated global hypomethylation in hepatocellular carcinoma was associated with gene-specific hypermethylation. In contrast, gene-specific hypermethylation and global hypomethylation appears to be an independent process in colon and urothelial cancers.30, 36 As there are site-specific differences in LINE-1 hypomethylation levels of normal tissue,37 further studies are needed to validate the relationship between global hypomethylation and hypermethylation of specific genes.
In conclusion, our study showed environmental factors such as geographic location and hepatitis status contribute to hepatocarcinogenesis through global hypomethylation, and global hypomethylation in hepatocellular carcinomas is closely associated with methylation of multiple CpG islands.
References
Jemal A, Siegel R, Ward E, et al. Cancer statistics, 2008. CA Cancer J Clin 2008;58:71–96.
Thorgeirsson SS, Grisham JW . Molecular pathogenesis of human hepatocellular carcinoma. Nat Genet 2002;31:339–346.
Herath NI, Leggett BA, MacDonald GA . Review of genetic and epigenetic alterations in hepatocarcinogenesis. J Gastroenterol Hepatol 2006;21:15–21.
Calvisi DF, Ladu S, Gorden A, et al. Mechanistic and prognostic significance of aberrant methylation in the molecular pathogenesis of human hepatocellular carcinoma. J Clin Invest 2007;117:2713–2722.
Ostertag EM, Kazazian Jr HH . Biology of mammalian L1 retrotransposons. Annu Rev Genet 2001;35:501–538.
Kazazian Jr HH, Goodier JL . LINE drive: retrotransposition and genomic instability. Cell 2002;110:277–280.
Lengauer C, Kinzler KW, Vogelstein B . DNA methylation and genetic instability in colorectal cancer cells. Proc Natl Acad Sci USA 1997;94:2103–2105.
Tuck-Muller CM, Narayan A, Tsien F, et al. DNA hypomethylation and unusual chromosome instability in cell lines from ICF syndrome patients. Cytogenet Cell Genet 2000;89:121–128.
Rodriguez J, Frigola J, Vendrell E, et al. Chromosomal instability correlates with genome-wide DNA demethylation in human primary colorectal cancers. Cancer Res 2006;66:8462–8468.
Takai D, Yagi Y, Habib N, et al. Hypomethylation of LINE1 retrotransposon in human hepatocellular carcinomas, but not in surrounding liver cirrhosis. Jpn J Clin Oncol 2000;30:306–309.
Lin CH, Hsieh SY, Sheen IS, et al. Genome-wide hypomethylation in hepatocellular carcinogenesis. Cancer Res 2001;61:4238–4243.
Choi IS, Estecio MR, Nagano Y, et al. Hypomethylation of LINE-1 and Alu in well-differentiated neuroendocrine tumors (pancreatic endocrine tumors and carcinoid tumors). Mod Pathol 2007;20:802–810.
Calvisi DF, Simile MM, Ladu S, et al. Altered methionine metabolism and global DNA methylation in liver cancer: relationship with genomic instability and prognosis. Int J Cancer 2007;121:2410–2420.
Shen L, Ahuja N, Shen Y, et al. DNA methylation and environmental exposures in human hepatocellular carcinoma. J Natl Cancer Inst 2002;94:755–761.
Toyota M, Ahuja N, Ohe-Toyota M, et al. CpG island methylator phenotype in colorectal cancer. Proc Natl Acad Sci USA 1999;96:8681–8686.
Yang AS, Estecio MRH, Doshi K, et al. A simple method for estimating global DNA methylation using bisulfite PCR of repetitive DNA elements. Nucl Acids Res 2004;32:e38.
Estecio MRH, Gharibyan V, Shen L, et al. LINE-1 hypomethylation in cancer is highly variable and inversely correlated with microsatellite instability. PLoS ONE 2007;2:e399.
Feinberg AP, Vogelstein V . Hypomethylation distinguishes genes of some human cancers from their normal counterparts. Nature 1983;301:89–92.
Issa JP . CpG-island methylation in aging and cancer. Curr Top Microbiol Immunol 2000;249:101–118.
Jones PA, Baylin SB . The fundamental role of epigenetic events in cancer. Nat Rev Genet 2002;3:415–428.
Ehrlich M, Woods CB, Yu MC, et al. Quantitative analysis of associations between DNA hypermethylations, hypomethylation, and DNMT RNA levels in ovarian tumors. Oncogene 2006;25:2636–2645.
Pattamadilok J, Huapai N, Rattanatanyong P, et al. LINE-1 hypomethylation level as a potential prognostic factor for epithelial ovarian cancer. Int J Gynecol Cancer 2008;18:711–717.
Suter CM, Martin DI, Ward RL . Hypomethylation of L1 retrotransposons in colorectal cancer and adjacent normal tissue. Int J Colorectal Dis 2004;19:95–101.
Ogino S, Kawasaki T, Nosho K, et al. LINE-1 hypomethylation is inversely associated with microsatellite instability and CpG island methylator phenotype in colorectal cancer. Int J Cancer 2008;122:2767–2773.
Santourlidis S, Florl A, Ackermann R, et al. High frequency of alterations in DNA methylation in adenocarcinoma of the prostate. Prostate 1999;39:166–174.
Cho NY, Kim BH, Choi M, et al. Hypermethylation of CpG island loci and hypomethylation of LINE-1 and Alu repeats in prostate adenocarcinoma and their relationship to clinicopathologic features. J Pathol 2007;211:269–277.
Park IY, Sohn BH, Yu E, et al. Aberrant epigenetic modifications in hepatocarcinogenesis induced by hepatitis B virus X protein. Gastroenterology 2007;132:1476–1494.
Su PF, Lee TC, Lin PJ, et al. Differential DNA methylation associated with hepatitis B virus infection in hepatocellular carcinoma. Int J Cancer 2007;121:1257–1264.
Kovalenko VM, Bagnyukova TV, Sergienko OV, et al. Epigenetic changes in the rat livers induced by pyrazinamide treatment. Toxicol Appl Pharmacol 2007;225:293–299.
Bariol C, Suter C, Cheong K, et al. The relationship between hypomethylation and CpG island methylation in colorectal neoplasia. Am J Pathol 2003;162:1361–1371.
Cravo M, Pinto R, Fidalgo P, et al. Global DNA hypomethylation occurs in the early stages of intestinal type gastric cancer. Gut 1996;39:434–438.
Roman-Gomez J, Jimenez-Velasco A, Agirre X, et al. Promoter hypomethylation of the LINE-1 retrotransposable elements activates sense/antisense transcription and marks the progression of chronic myeloid leukemia. Oncogene 2005;24:7213–7223.
Soares J, Pinto AE, Cunha CV, et al. Global DNA hypomethylation in breast carcinoma: correlation with prognostic factors and tumor progression. Cancer 1999;85:112–118.
Smith IM, Mydlarz WK, Mithani SK, et al. DNA global hypomethylation in squamous cell head and neck cancer associated with smoking, alcohol consumption and stage. Int J Cancer 2007;121:1724–1728.
Tangkijvanich P, Hourpai N, Rattanatanyong P, et al. Serum LINE-1 hypomethylation as a potential prognostic marker for hepatocellular carcinoma. Clin Chim Acta 2002;379:127–133.
Neuhausen A, Florl AR, Grimm MO, et al. DNA methylation alterations in urothelial carcinoma. Cancer Biol Ther 2006;5:993–1001.
Chalitchagorn K, Shuangshoti S, Hourpai N, et al. Distinctive pattern of LINE-1 methylation level in normal tissues and the association with carcinogenesis. Oncogene 2004;23:8841–8846.
Author information
Authors and Affiliations
Corresponding author
Additional information
Disclosure/conflict of interest
There are no disclosures or conflicts of interest.
Rights and permissions
About this article
Cite this article
Kim, MJ., White-Cross, J., Shen, L. et al. Hypomethylation of long interspersed nuclear element-1 in hepatocellular carcinomas. Mod Pathol 22, 442–449 (2009). https://doi.org/10.1038/modpathol.2008.203
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.2008.203
Keywords
This article is cited by
-
Genome-wide tracts of homozygosity and exome analyses reveal repetitive elements with Barrets esophagus/esophageal adenocarcinoma risk
BMC Bioinformatics (2019)
-
Global methylation, oxidative stress and relative telomere length in biliary atresia patients
Scientific Reports (2016)
-
Global DNA hypomethylation in prostate cancer development and progression: a systematic review
Prostate Cancer and Prostatic Diseases (2015)
-
DNA Methylation and Flavonoids in Genitourinary Cancers
Current Pharmacology Reports (2015)
-
Role of estrogen in hepatocellular carcinoma: is inflammation the key?
Journal of Translational Medicine (2014)