Abstract
The normal intestinal flora is required for the development of intestinal inflammation in animal models of inflammatory bowel disease (IBD). In humans, several studies indicated a potential association of Escherichia coli (E. coli) with IBD. In addition, we have shown that T-cell clones of IBD patients cross react toward different enteric bacterial species and thus likely respond to conserved bacterial antigens. Therefore, we hypothesized that highly conserved E. coli proteins might be a reasonable candidate to screen for abnormal T-cell responses in IBD. We used high-throughput techniques for cloning, expression, and purification under native conditions of a set of 271 conserved proteins of E. coli, of which 196 were used for whole blood stimulations to assess peripheral T helper (TH)-cell responses. In addition, because of the association of an adherent-invasive E. coli with Crohn's disease (CD), we included 13 pathogenicity factors of E. coli in the study. We observed that pools of these conserved E. coli proteins less frequently induced interferon-γ (IFNγ) production in peripheral TH cells in patients with CD and ankylosing spondylitis (AS) compared with healthy controls. In addition, lower percentage of patients with CD and AS responded toward single proteins. The reason for the decreased frequency of an in vitro TH-cell IFNγ response toward E. coli proteins in peripheral blood of CD and AS patients, e.g., increased suppression needs to be clarified.
Similar content being viewed by others
Introduction
Crohn's disease (CD) and ulcerative colitis are the two pheno-typic prototypes of inflammatory bowel disease (IBD). CD is characterized by granulomatous inflammation throughout the entire gastrointestinal tract with the terminal ileum mainly affected1 whereas ulcerative colitis is characterized by chronic continuous inflammation in the colorectum without granuloma formation.2 Alhough the etiology of IBD remains unclear, it is well established that genetic factors, environmental factors, and a dysregulated immune response toward commensal bacteria contribute to the pathogenesis.3 Most animal models of IBD are characterized by T helper 1 (TH1) cells4 driven by the enteric bacterial flora.3, 4, 5 In addition to the stimulatory role of the normal intestinal flora, several studies highlighted the association of Escherichia coli (E. coli) and CD. DNA of E. coli was detected in 80% of CD granulomas.6 Antigens of E. coli, Listeria, and Streptococcus were present within the macrophages of CD patients.7 Predominance of E. coli was demonstrated in ileal mucosa8, 9. An adherent-invasive E. coli strain isolated from a chronic lesion of a CD patient was able to survive and replicate in the host-cell cytoplasm after lysis of the endocytic vacuole.10 Moreover, this strain was shown to survive and replicate within macrophages.11 Adherent-invasive E. coli are associated with inflammatory lesions of CD patients.12 Several studies demonstrated the potential role of conserved bacterial antigens in the pathogenesis of IBD and related disorders. T-cell clones of IBD patients were shown to cross-react toward antigens shared by different enteric bacterial species including E. coli13 indicating that conserved antigens of intestinal bacteria could drive chronic inflammation directly or through induction of autoimmunity. In Yersinia-triggered reactive arthritis, which together with IBD and ankylosing spondylitis (AS) belongs to the group of the spondyloarthropathies, a conserved ribosomal protein from Yersinia enterocolitica was identified as an immunodominant antigen.14 In addition, heat-shock protein 60 from Yersinia enterocolitica, Chlamydia trachomatis, Borrelia burgdorferi, and homo sapiens caused proliferative responses with TH1- and TH2-like cytokine-secretion pattern in T-cell clones of a patient with reactive arthritis.15 Yersinia heat-shock protein 60-specific T cells cross-reacted with human heat-shock protein 60.16
On the basis of these findings, we wished to study a potential role of conserved bacterial proteins in the pathogenesis of IBD. The most conserved protein functions are represented in all three biological kingdoms, archaea, prokarya, and eukarya. On account of the potential involvement of E. coli in IBD pathogenesis, we first chose a set of E. coli proteins hypothetically inherited by the Last Universal Common Ancestor (LUCA)17 of the three kingdoms (LUCA proteins). Subsequently, we identified additional conserved proteins between E. coli and humans that were not included in the LUCA set of proteins. We termed these proteins as E. coli-human-homologues. Together, we identified 271 conserved proteins of E. coli. Of these, we accomplished to purify 197 proteins with a minimum amount of 500 μg.18 As most of the highly conserved protein functions are also represented in humans, T-cell reactivity toward these proteins could provide a hint to potential autoimmunity in IBD. In addition, because of the association of an adherent-invasive E. coli with CD,10, 11, 12 we included 13 pathogenicity factors of E. coli in the study. We stimulated whole blood of CD patients, AS patients, and healthy controls with protein pools containing conserved proteins and pathogenicity factors of E. coli, and assessed the CD4+ TH-cell response according to expression of interferon-γ (IFNγ) and CD40 ligand, as CD40 ligand was shown to be an universal marker for the activation of antigen-specific TH cells.19
Results
Expression and purification of recombinant proteins of E. coli
As previously published,18 of 246 LUCA proteins described by Kyrpides et al.,17 we could identify 223 LUCA genes in the E. coli K12 genome. In addition, we identified 48 E. coli-human-homologues proteins. Together, a total of 271 conserved proteins of E. coli could be identified. Of these, we successfully cloned 268 genes (98%) in the expression vector pQTEV2. In small-scale, we could detect the expression of 254 proteins (94%) in the cell lysates. Of these proteins, 249 (98%) yielded purified His-tagged proteins (Figure 1). We observed that several proteins either were not purified or were purified in low yields. We subjected the corresponding genes to the Gateway recombination technology to fuse each of the target proteins with maltose binding protein, glutathione S-transferase, and N-utilizing substance A, respectively. Protein expression and purification revealed that fusion of target proteins to maltose binding protein most efficiently improved protein yields. Fusion to N-utilizing substance A or glutathione S-transferase resulted in little improvement. Treatment of recombinant proteins with polymyxin B-coated magnetic beads revealed less background stimulation of TH cells as compared with untreated proteins. Furthermore, the Limulus amoebocyte lysate test revealed that lipopolysaccharide contents in polymyxin B-treated proteins were below the detection limit. In conclusion, we accomplished to produce 197 conserved proteins with a minimum amount of 500 μg. Besides conserved proteins, we expressed and purified 13 pathogenicity factors of various pathogenic E. coli strains.
Step-by-step efficiency of cloning, small-scale protein expression and purification of 271 conserved proteins of E. coli. A total of 220 purified proteins with high and moderate yields were subjected to large-scale expression. Afterwards, 176 proteins could be purified with a minimum of 500 μg. Overall, 34 target proteins were subjected to Gateway recombination cloning to express fusion proteins with maltose binding protein, glutathione S-transferase, and N-utilizing substance A, respectively. Overall, 21 proteins fused to maltose binding protein were purified with a minimum of 500 μg. ECHH, E. coli-human-homologues; LUCA, Last Common Universal Ancestor
Stimulation of whole blood with pools of E. coli proteins
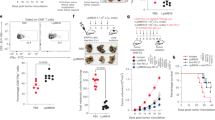
Upon stimulation of whole blood, we observed that the majority of patients with CD and AS lacked peripheral TH cell responses toward E. coli protein pools. Inversely, such responses were present in the majority of healthy controls (Figure 2).
Peripheral TH cell responses (CD40 ligand and IFNγ) upon stimulation of whole blood with pools of conserved E. coli proteins. (a) HC, (b) CD patients, and (c) AS patients. Rectangles in gray indicate TH-cell responses toward protein pools where frequencies of double positive TH cells were at least three-fold increased as compared with the negative control (blood sample supplemented only with anti-CD28 antibody). Few patients with CD and AS responded toward pools. In contrast, higher percentages of healthy controls were responding. AS, ankylosing spondylitis; CD, Crohn's disease; HC, healthy controls; TH, T helper cell.
Identification of stimulatory proteins
To rapidly identify single stimulatory proteins within pools, proteins had been distributed among the pools following a “protein matrix” (Table 1).
To identify proteins driving the immune response in patients with CD, AS, and controls, we first identified protein pools that most frequently caused TH-cell responses in these groups. Whole blood of patients and controls with reactivity to a given protein pool was then restimulated with all proteins contained in that pool, as described above. By that approach nine single stimulatory proteins were identified. Whole blood of patients and healthy controls (10 each) was then stimulated with these conserved proteins to assess the frequencies of responding individuals. Similar to stimulations with protein pools, more healthy controls responded toward these nine proteins than patients with CD and AS (manuscript in preparation).
Discussion
Using a strategy that combines standard restriction cloning with the Gateway recombination system, we established a reliable and efficient approach for the large-scale generation of a large number of evolutionary conserved bacterial proteins, suitable for usage in in vitro T-cell assays.
We observed that pools of these evolutionary conserved E. coli proteins less frequently induced IFNγ production in peripheral TH cells in patients with CD and AS compared with healthy controls. Nine stimulatory conserved proteins were identified from protein pools that most frequently caused TH-cell responses in patients with CD, AS, and healthy controls. Similar to the responses observed against protein pools, lower percentage of patients with CD and AS responded toward these single proteins.
Certainly, the increased and broad reactivity of peripheral blood T cells from healthy controls to evolutionary conserved bacterial proteins is an in vitro phenomenon and should not be interpreted to indicate an ongoing inflammatory process in vivo. This is certainly prevented in vivo by a lack of sufficient stimulatory conditions (e.g., lack of an adequate antigen dose) and/or sufficient suppression. The reason for the decreased frequency of an in vitro TH-cell IFNγ response toward E. coli proteins in peripheral blood of CD and AS patients is not entirely clear. Explanations include that the number of responsive peripheral blood T cells is decreased, e.g., because bacteria reactive T cells enrich in the inflamed tissues of these patients. However, this alone seems unlikely as the observed phenomenon was independent of disease activity. Furthermore, decreased T-cell reactivity might be a consequence of an impaired innate immune response, as recently reported in CD patients by Marks et al.20 or a consequence of increased suppression. Studies to distinguish among these possibilities are currently performed.
Our preliminary data suggest that stimulation with staphylococcal enterotoxin B, a super antigen that cross-links defined Vβ-chains of the T-cell receptor with major histocompatibility complex II,21, 22 also resulted in decreased frequencies of responding peripheral TH cells in patients with CD and AS. In contrast to staphylococcal enterotoxin B, however, we could not observe differences in the frequencies of responding peripheral TH cells toward the control antigens cytomegalovirus pp6523 and tetanus toxoid24 (unpublished data). Whether this indicates a specific abnormality of antibacterial T-cell responses in CD and AS needs further clarification.
Methods
Generation of recombinant proteins. Amplification through PCR and cloning of target genes into the expression vector pQTEV2 and the expression of proteins was performed as described previously. 18 Briefly, to identify clones that provided sufficient amounts of proteins, His-tagged proteins were expressed in small-scale and purified under native conditions through affinity chromatography to a nickel chelate matrix (Ni-NTA; Qiagen, Hilden, Germany). Elution of bound proteins into a denaturing buffer and sodium dodecyl sulfate polyacrylamide gel electrophoresis analysis identified proteins that were purified in sufficient yields. Suitable expression clones were then used for large-scale protein expression and purification to obtain a minimum of 500 μg per protein. In cases that this strategy did not yield in sufficient amounts of proteins, we used the Gateway (Invitrogen, Karlsruhe, Germany) recombination technology25 to fuse the target protein with fusion partners, such as maltose binding protein, glutathione S-transferase, or N-utilizing substance A. These fusion partners may enhance solubility of target proteins.26 Finally, we removed bacterial lipopolysaccharide from protein solutions with polymyxin B-coated magnetic beads (Chemicell, Berlin, Germany). Lipopolysaccharide was reported to interfere with downstream immunological applications27, 28, 29 such as in vitro stimulation of whole blood or peripheral blood mononuclear cells. Pathogenicity factors of E. coli were purified under denatured conditions using 6 M of guanidine hydrochloride to disrupt the bacterial cell wall and to denature proteins. After binding to Ni-NTA, the proteins were washed and eluted with 8 M of urea. Urea was removed by dialysis against 0.1% acetic acid. Subsequently, we removed lipopolysaccharide as described above.
Pooling of E. coli proteins. A total of 196 conserved proteins and 13 pathogenicity factors of E. coli were each adjusted to a final concentration of 500 μg ml−1. We pooled 196 conserved proteins and four pathogenicity factors. Each pool consisted of 10 proteins, resulting in 20 pools. Beside this arrangement, we distributed the proteins in additional 20 “matrix pools.” Thus, each protein was present in two different protein pools. The “protein matrix” containing 100 conserved proteins is illustrated in Table 1. The aim of this approach was the fast identification of potential stimulatory proteins in whole blood stimulations.
Patients and healthy controls. Heparinized whole blood was obtained from 40 CD patients (22 female, 18 male; mean age: 44 years; range: 17–70), 20 AS patients (9 female, 11 male; mean age: 43 years; range: 27–63), and 20 healthy controls (11 female, 9 male; mean age: 37 years; range: 20–61). The study was approved by the local ethics committee.
Stimulation of whole blood with protein pools and flow cytometry analysis. We stimulated whole blood with proteins as described previously.18 Briefly, heparinized blood was stimulated for 6 h at 37 °C and 5% CO2 in the presence of 1 μg ml−1 anti-CD28 antibody (Becton Dickinson, Heidelberg, Germany) with 5 μg ml−1 of E. coli proteins either in pools or as single proteins, 4 μg ml−1 of cytomegalovirus pp65 (Miltenyi, Bergisch Gladbach, Germany), 10 μg ml−1 of tetanus toxoid (Chiron–Behring, Marburg, Germany), and 10 μg ml−1 of staphylococcal enterotoxin B (Sigma, Deisenhofen, Germany) as the positive control. The negative control comprised only blood with anti-CD28 antibody. The last 4 h of stimulation were performed under the presence of 10 μg ml−1 of the secretion inhibitor Brefeldin A (Sigma) to accumulate intracellular cytokines. EDTA (2 mM) was added to the samples and incubated for 10 min at 20 °C. Erythrocytes were lysed and leukocytes were fixed with FACS Lysing Solution (Becton Dickinson) according to the manufacturer's instructions. The cells were permeabilized for subsequent antibody staining with FACS Permeabilizing Solution 2 (Becton Dickinson) according to the manufacturer's instructions. Before addition of following antibodies (Becton Dickinson) the cells were added with 1 mg ml−1 of Beriglobin (Behring) to avoid Fc binding: anti-CD4-PerCP, anti-CD40 ligand-PE, and anti-IFNγ-antigen presenting cell. The cells were incubated for 30 min at 20 °C in the dark, then washed with fluorescence-activated cell sorting buffer (phosphate-buffered saline with 0.5% bovine serum albumin and 0.1% sodium azide) and resuspended in an appropriate volume of fluorescence-activated cell sorting buffer. In the flow cytometry analysis, we included 200,000 TH cells and assessed the frequencies of CD4+ TH cells double positive for CD40 ligand and IFNγ. In the analysis, TH-cell responsiveness toward E. coli protein pools was defined as an at least three-fold increase in frequency of double-positive TH cells compared with the negative control.
Disclosure
Thomas Adam has received grant support from an unnamed source. Joachim Sieper has received consulting fees and lecture fees from Schering Plough, Wyeth, Abbott, and Pfitzer. Joachim Sieper has also received grant support from Schering Plough, Wyeth, and Abbott. Asgar Ergin, Konrad Büssow, Andreas Thiel, and Rainer Duchmann have declared no financial interests.
References
Duchmann, R. & Zeitz, M. Crohn's disease: current pathogenetic paradigms In Mucosal Immunology 3rd edn (Mestecky, J. et al., eds) 1265–1286 ( Elsevier Academic Press, San Diego, 2005 ).
Karp, L.C. & Targan, S.R. Ulcerative colitis: evolving insights into pathogenesis In Mucosal Immunology, 3rd edn (Mestecky, J., et al., eds) 1255–1264 ( Elsevier Academic Press, San Diego, 2005 ).
Lakatos, P.L., Fischer, S., Lakatos, L., Gal, I. & Papp, J. Current concept on the pathogenesis of inflammatory bowel disease-crosstalk between genetic and microbial factors: pathogenic bacteria and altered bacterial sensing or changes in mucosal integrity take “toll”? World J. Gastroenterol. 12, 1829–1841 (2006).
Strober, W., Fuss, I.J. & Blumberg, R.S. The immunology of mucosal models of inflammation. Annu. Rev. Immunol. 20, 495–549 (2002).
Guarner, F. & Malagelada, J.R. Role of bacteria in experimental colitis. Best Pract. Res. Clin. Gastroenterol. 17, 793–804 (2003).
Ryan, P. et al. Bacterial DNA within granulomas of patients with crohn's disease—detection by laser capture microdissection and PCR. Am. J. Gastroenterol. 99, 1539–1543 (2004).
Liu, Y. et al. Immunocytochemical evidence of listeria, escherichia coli, and streptococcus antigens in crohn's disease. Gastroenterology 108, 1396–1404 (1995).
Kotlowski, R., Bernstein, C.N., Sepehri, S. & Krause, D.O. High prevalence of Escherichia coli belonging to the B2+D phylogenetic group in inflammatory bowel disease. Gut 56, 669–675 (2007).
Darfeuille-Michaud, A. et al. Presence of adherent Escherichia coli strains in ileal mucosa of patients with crohn's disease. Gastroenterology 115, 1405–1413 (1998).
Boudeau, J., Glasser, A.L., Masseret, E., Joly, B. & Darfeuille-Michaud, A. Invasive ability of an Escherichia coli strain isolated from the ileal mucosa of a patient with crohn's disease. Infect. Immun. 67, 4499–4509 (1999).
Glasser, A.L. et al. Adherent invasive Escherichia coli strains from patients with Crohn's disease survive and replicate within macrophages without inducing host cell death. Infect. Immun. 69, 5529–5537 (2001).
Barnich, N., Bringer, M.A., Claret, L. & Darfeuille-Michaud, A. Involvement of lipoprotein NlpI in the virulence of adherent invasive Escherichia coli strain LF82 isolated from a patient with crohn's disease. Infect. Immun. 72, 2484–2493 (2004).
Duchmann, R. et al. T cell specificity and cross reactivity towards enterobacteria, bacteroides, bifidobacterium, and antigens from resident intestinal flora in humans. Gut 44, 812–818 (1999).
Mertz, A.K. et al. The evolutionarily conserved ribosomal protein L23 and the cationic urease beta-subunit of Yersinia enterocolitica O:3 belong to the immunodominant antigens in Yersinia-triggered reactive arthritis: implications for autoimmunity. Mol. Med. 1, 44–55 (1994).
Mertz, A.K. et al. Characterization of the synovial T cell response to various recombinant Yersinia antigens in Yersinia enterocolitica-triggered reactive arthritis. Heat-shock protein 60 drives a major immune response. Arthritis Rheum. 41, 315–326 (1998).
Thiel, A. et al. Identification of immunodominant CD4+ T cell epitopes in patients with Yersinia-induced reactive arthritis by cytometric cytokine secretion assay. Arthritis Rheum. 54, 3583–3590 (2006).
Kyrpides, N., Overbeek, R. & Ouzounis, C. Universal protein families and the functional content of the last universal common ancestor. J. Mol. Evol. 49, 413–423 (1999).
Ergin, A. et al. Homologous high-throughput expression and purification of highly conserved E coli proteins. Microb. Cell Fact. 6, 18 (2007).
Frentsch, M. et al. Direct access to CD4+ T cells specific for defined antigens according to CD154 expression. Nat. Med. 11, 1118–1124 (2005).
Marks, D.J. et al. Defective acute inflammation in Crohn's disease: a clinical investigation. Lancet 367, 668–678 (2006).
Choi, Y.W. et al. Interaction of Staphylococcus aureus toxin “superantigens” with human T cells. Proc. Natl. Acad. Sci. USA 86, 8941–8945 (1989).
Hong, S.C., Waterbury, G. & Janeway, C.A. Jr . Different superantigens interact with distinct sites in the Vbeta domain of a single T cell receptor. J. Exp. Med. 183, 1437–1446 (1996).
Beninga, J., Kropff, B. & Mach, M. Comparative analysis of fourteen individual human cytomegalovirus proteins for helper T cell response. J. Gen. Virol. 76 (Part 1), 153–160 (1995).
Fernandez, V., Andersson, J., Andersson, U. & Troye-Blomberg, M. Cytokine synthesis analyzed at the single-cell level before and after revaccination with tetanus toxoid. Eur. J. Immunol. 24, 1808–1815 (1994).
Walhout, A.J. et al. GATEWAY recombinational cloning: application to the cloning of large numbers of open reading frames or ORFeomes. Methods Enzymol. 328, 575–592 (2000).
Cabrita, L.D., Dai, W. & Bottomley, S.P. A family of E coli expression vectors for laboratory scale and high throughput soluble protein production. BMC Biotechnol. 6, 12 (2006).
Bekeredjian-Ding, I. et al. T cell-independent, TLR-induced IL-12p70 production in primary human monocytes. J. Immunol. 176, 7438–7446 (2006).
Schroder, K., Sweet, M.J. & Hume, D.A. Signal integration between IFNgamma and TLR signalling pathways in macrophages. Immunobiology 211, 511–524 (2006).
West, A.P., Koblansky, A.A. & Ghosh, S. Recognition and signaling by toll-like receptors. Annu. Rev. Cell Dev. Biol. 22, 409–437 (2006).
Acknowledgements
We gratefully acknowledge excellent technical assistance of Rebecca Scheer and Peihua Wu and helpful discussions with Kerstin Kapp. This study was funded by DFG grant SFB633-A4.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Ergin, A., Adam, T., Büssow, K. et al. Identification of the predominant antigenic epitopes in intestinal flora in IBD. Mucosal Immunol 1 (Suppl 1), S19–S23 (2008). https://doi.org/10.1038/mi.2008.44
Published:
Issue Date:
DOI: https://doi.org/10.1038/mi.2008.44