Abstract
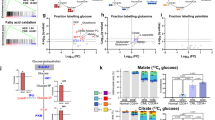
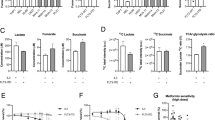
Alterations in metabolic activities are cancer hallmarks that offer a wide range of new therapeutic opportunities. Here we decipher the interplay between mTORC1 activity and glucose metabolism in acute myeloid leukemia (AML). We show that mTORC1 signaling that is constantly overactivated in AML cells promotes glycolysis and leads to glucose addiction. The level of mTORC1 activity determines the sensitivity of AML cells to glycolysis inhibition as switch-off mTORC1 activity leads to glucose-independent cell survival that is sustained by an increase in mitochondrial oxidative phosphorylation. Metabolic analysis identified the pentose phosphate pathway (PPP) as an important pro-survival pathway for glucose metabolism in AML cells with high mTORC1 activity and provided a clear rational for targeting glucose-6-phosphate dehydrogenase (G6PD) in AML. Indeed, our analysis of the cancer genome atlas AML database pinpointed G6PD as a new biomarker in AML, as its overexpression correlated with an adverse prognosis in this cohort. Targeting the PPP using the G6PD inhibitor 6-aminonicotinamide induces in vitro and in vivo cytotoxicity against AML cells and synergistically sensitizes leukemic cells to chemotherapy. Our results demonstrate that high mTORC1 activity creates a specific vulnerability to G6PD inhibition that may work as a new AML therapy.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Warburg O . On the origin of cancer cells. Science 1956; 123: 309–314.
Vander Heiden MGV, Cantley LC, Thompson CB . Understanding the Warburg Effect: the metabolic requirements of cell proliferation. Science 2009; 324: 1029–1033.
Vander Heiden MG . Targeting cancer metabolism: a therapeutic window opens. Nat Rev Drug Discov 2011; 10: 671–684.
Laplante M, Sabatini DM . mTOR signaling in growth control and disease. Cell 2012; 149: 274–293.
Inoki K, Li Y, Zhu T, Wu J, Guan K-L . TSC2 is phosphorylated and inhibited by Akt and suppresses mTOR signalling. Nat Cell Biol 2002; 4: 648–657.
Bar-Peled L, Schweitzer LD, Zoncu R, Sabatini DM . Ragulator is a GEF for the Rag GTPases that signal amino acid levels to mTORC1. Cell 2012; 150: 1196–1208.
Jewell JL, Kim YC, Russell RC, Yu F-X, Park HW, Plouffe SW et al. Differential regulation of mTORC1 by leucine and glutamine. Science 2015; 347: 194–198.
Inoki K, Zhu T, Guan K-L . TSC2 mediates cellular energy response to control cell growth and survival. Cell 2003; 115: 577–590.
Düvel K, Yecies JL, Menon S, Raman P, Lipovsky AI, Souza AL et al. Activation of a metabolic gene regulatory network downstream of mTOR complex 1. Mol Cell 2010; 39: 171–183.
Dibble CC, Manning BD . Signal integration by mTORC1 coordinates nutrient input with biosynthetic output. Nat Cell Biol 2013; 15: 555–564.
Shimobayashi M, Hall MN . Making new contacts: the mTOR network in metabolism and signalling crosstalk. Nat Rev Mol Cell Biol 2014; 15: 155–162.
Zinzalla V, Stracka D, Oppliger W, Hall MN . Activation of mTORC2 by association with the ribosome. Cell 2011; 144: 757–768.
Hagiwara A, Cornu M, Cybulski N, Polak P, Betz C, Trapani F et al. Hepatic mTORC2 activates glycolysis and lipogenesis through Akt, glucokinase, and SREBP1c. Cell Metab 2012; 15: 725–738.
Masui K, Tanaka K, Akhavan D, Babic I, Gini B, Matsutani T et al. mTOR complex 2 controls glycolytic metabolism in glioblastoma through FoxO acetylation and upregulation of c-Myc. Cell Metab 2013; 18: 726–739.
Ciriello G, Miller ML, Aksoy BA, Senbabaoglu Y, Schultz N, Sander C . Emerging landscape of oncogenic signatures across human cancers. Nat Genet 2013; 45: 1127–1133.
Xu Q, Simpson S-E, Scialla TJ, Bagg A, Carroll M . Survival of acute myeloid leukemia cells requires PI3 kinase activation. Blood 2003; 102: 972–980.
Tamburini J, Green AS, Bardet V, Chapuis N, Park S, Willems L et al. Protein synthesis is resistant to rapamycin and constitutes a promising therapeutic target in acute myeloid leukemia. Blood 2009; 114: 1618–1627.
Döhner H, Weisdorf DJ, Bloomfield CD . Acute myeloid leukemia. N Engl J Med 2015; 373: 1136–1152.
Büchner T, Schlenk RF, Schaich M, Döhner K, Krahl R, Krauter J et al. Acute myeloid leukemia (AML): different treatment strategies versus a common standard arm—combined prospective analysis by the German AML Intergroup. J Clin Oncol 2012; 30: 3604–3610.
Konig H, Levis M . Is targeted therapy feasible in acute myelogenous leukemia? Curr Hematol Malig Rep 2014; 9: 118–127.
Hoshii T, Tadokoro Y, Naka K, Ooshio T, Muraguchi T, Sugiyama N et al. mTORC1 is essential for leukemia propagation but not stem cell self-renewal. J Clin Invest 2012; 122: 2114–2129.
Kalaitzidis D, Sykes SM, Wang Z, Punt N, Tang Y, Ragu C et al. mTOR complex 1 plays critical roles in hematopoiesis and PTEN-loss-evoked leukemogenesis. Cell Stem Cell 2012; 11: 429–439.
Magee JA, Ikenoue T, Nakada D, Lee JY, Guan K-L, Morrison SJ . Temporal changes in PTEN and mTORC2 regulation of hematopoietic stem cell self-renewal and leukemia suppression. Cell Stem Cell 2012; 11: 415–428.
Récher C, Beyne-Rauzy O, Demur C, Chicanne G, Santos CD, Mas VM-D et al. Antileukemic activity of rapamycin in acute myeloid leukemia. Blood 2005; 105: 2527–2534.
Willems L, Chapuis N, Puissant A, Maciel TT, Green AS, Jacque N et al. The dual mTORC1 and mTORC2 inhibitor AZD8055 has anti-tumor activity in acute myeloid leukemia. Leukemia 2012; 26: 1195–1202.
Sujobert P, Poulain L, Paubelle E, Zylbersztejn F, Grenier A, Lambert M et al. Co-activation of AMPK and mTORC1 induces cytotoxicity in acute myeloid leukemia. Cell Rep 2015; 11: 1446–1457.
Willems L, Jacque N, Jacquel A, Neveux N, Maciel TT, Lambert M et al. Inhibiting glutamine uptake represents an attractive new strategy for treating acute myeloid leukemia. Blood 2013; 122: 3521–3532.
Jacque N, Ronchetti AM, Larrue C, Meunier G, Birsen R, Willems L et al. Targeting glutaminolysis has antileukemic activity in acute myeloid leukemia and synergizes with BCL-2 inhibition. Blood 2015; 126: 1346–1356.
Wang Y-H, Israelsen WJ, Lee D, Yu VWC, Jeanson NT, Clish CB et al. Cell-state-specific metabolic dependency in hematopoiesis and leukemogenesis. Cell 2014; 158: 1309–1323.
Rossignol R, Gilkerson R, Aggeler R, Yamagata K, Remington SJ, Capaldi RA . Energy substrate modulates mitochondrial structure and oxidative capacity in cancer cells. Cancer Res 2004; 64: 985–993.
Scotland S, Saland E, Skuli N, de Toni F, Boutzen H, Micklow E et al. Mitochondrial energetic and AKT status mediate metabolic effects and apoptosis of metformin in human leukemic cells. Leukemia 2013; 27: 2129–2138.
Lagadinou ED, Sach A, Callahan K, Rossi RM, Neering SJ, Minhajuddin M et al. BCL-2 inhibition targets oxidative phosphorylation and selectively eradicates quiescent human leukemia stem cells. Cell Stem Cell 2013; 12: 329–341.
Lee W-NP, Boros LG, Puigjaner J, Bassilian S, Lim S, Cascante M . Mass isotopomer study of the nonoxidative pathways of the pentose cycle with [1,2-13C2]glucose. Am J Physiol Endocrinol Metab 1998; 274: E843–E851.
Billiard J, Dennison JB, Briand J, Annan RS, Chai D, Colón M et al. Quinoline 3-sulfonamides inhibit lactate dehydrogenase A and reverse aerobic glycolysis in cancer cells. Cancer Metab 2013; 1: 19.
TCGA Research Network. Genomic and epigenomic landscapes of adult de novo acute myeloid leukemia. N Engl J Med 2013; 368: 2059–2074.
Bagger FO, Sasivarevic D, Sohi SH, Laursen LG, Pundhir S, Sønderby CK et al. BloodSpot: a database of gene expression profiles and transcriptional programs for healthy and malignant haematopoiesis. Nucleic Acids Res 2016; 44: D917–D924.
Jung N, Dai B, Gentles AJ, Majeti R, Feinberg AP . An LSC epigenetic signature is largely mutation independent and implicates the HOXA cluster in AML pathogenesis. Nat Commun 2015; 6: 8489.
Hothersall JS, Gordge M, Noronha-Dutra AA . Inhibition of NADPH supply by 6-aminonicotinamide: effect on glutathione, nitric oxide and superoxide in J774 cells. FEBS Lett 1998; 434: 97–100.
Samudio I, Harmancey R, Fiegl M, Kantarjian H, Konopleva M, Korchin B et al. Pharmacologic inhibition of fatty acid oxidation sensitizes human leukemia cells to apoptosis induction. J Clin Invest 2010; 120: 142–156.
Sriskanthadevan S, Jeyaraju DV, Chung TE, Prabha S, Xu W, Skrtic M et al. AML cells have low spare reserve capacity in their respiratory chain that renders them susceptible to oxidative metabolic stress. Blood 2015; 125: 2120–2130.
Goto M, Miwa H, Shikami M, Tsunekawa-Imai N, Suganuma K, Mizuno S et al. Importance of glutamine metabolism in leukemia cells by energy production through TCA cycle and by redox homeostasis. Cancer Invest 2014; 32: 241–247.
Miraki-Moud F, Ghazaly E, Ariza-McNaughton L, Hodby KA, Clear A, Anjos-Afonso F et al. Arginine deprivation using pegylated arginine deiminase has activity against primary acute myeloid leukemia cells in vivo. Blood 2015; 125: 4060–4068.
Herst PM, Howman RA, Neeson PJ, Berridge MV, Ritchie DS . The level of glycolytic metabolism in acute myeloid leukemia blasts at diagnosis is prognostic for clinical outcome. J Leukoc Biol 2011; 89: 51–55.
Chen W-L, Wang J-H, Zhao A-H, Xu X, Wang Y-H, Chen T-L et al. A distinct glucose metabolism signature of acute myeloid leukemia with prognostic value. Blood 2014; 124: 1645–1654.
Choo AY, Kim SG, Vander Heiden MG, Mahoney SJ, Vu H, Yoon S-O et al. Glucose addiction of TSC null cells is caused by failed mTORC1-dependent balancing of metabolic demand with supply. Mol Cell 2010; 38: 487–499.
Barger JF, Gallo CA, Tandon P, Liu H, Sullivan A, Grimes HL et al. S6K1 determines the metabolic requirements for BCR-ABL survival. Oncogene 2013; 32: 453–461.
Cunningham JT, Rodgers JT, Arlow DH, Vazquez F, Mootha VK, Puigserver P . mTOR controls mitochondrial oxidative function through a YY1–PGC-1α transcriptional complex. Nature 2007; 450: 736–740.
Morita M, Gravel S-P, Chénard V, Sikström K, Zheng L, Alain T et al. mTORC1 controls mitochondrial activity and biogenesis through 4E-BP-dependent translational regulation. Cell Metab 2013; 18: 698–711.
Pusapati RV, Daemen A, Wilson C, Sandoval W, Gao M, Haley B et al. mTORC1-dependent metabolic reprogramming underlies escape from glycolysis addiction in cancer cells. Cancer Cell 2016; 29: 548–562.
Zid BM, Rogers AN, Katewa SD, Vargas MA, Kolipinski MC, Lu TA et al. 4E-BP extends lifespan upon dietary restriction by enhancing mitochondrial activity in Drosophila. Cell 2009; 139: 149–160.
Kishton RJ, Barnes CE, Nichols AG, Cohen S, Gerriets VA, Siska PJ et al. AMPK is essential to balance glycolysis and mitochondrial metabolism to control T-ALL cell stress and survival. Cell Metab 2016; 23: 649–662.
Patra KC, Hay N . The pentose phosphate pathway and cancer. Trends Biochem Sci 2014; 39: 347–354.
Kohan AB, Talukdar I, Walsh CM, Salati LM . A role for AMPK in the inhibition of glucose-6-phosphate dehydrogenase by polyunsaturated fatty acids. Biochem Biophys Res Commun 2009; 388: 117–121.
Jiang P, Du W, Wang X, Mancuso A, Gao X, Wu M et al. p53 regulates biosynthesis through direct inactivation of glucose-6-phosphate dehydrogenase. Nat Cell Biol 2011; 13: 310–316.
Hong X, Song R, Song H, Zheng T, Wang J, Liang Y et al. PTEN antagonises Tcl1/hnRNPK-mediated G6PD pre-mRNA splicing which contributes to hepatocarcinogenesis. Gut 2014; 63: 1635–1647.
Kuo W, Lin J, Tang TK . Human glucose-6-phosphate dehydrogenase (G6PD) gene transforms NIH 3T3 cells and induces tumors in nude mice. Int J Cancer 2000; 85: 857–864.
Jiang P, Du W, Yang X . A critical role of glucose-6-phosphate dehydrogenase in TAp73-mediated cell proliferation. Cell Cycle 2013; 12: 3720–3726.
Rao X, Duan X, Mao W, Li X, Li Z, Li Q et al. O-GlcNAcylation of G6PD promotes the pentose phosphate pathway and tumor growth. Nat Commun 2015; 6: 8468.
Acknowledgements
This work was supported by grant from the Institut National du Cancer (Projet Recherche Translationnelle TRANSLA13-087), Ligue Nationale Contre le Cancer (Equipe Labellisée EL2014. No Projet: R14077KK), Plan Cancer BioSys 2014 (Project FLEXAML; J-CP and J-ES), INCa-Cancéropôle GSO (TLP) and from Région Midi-Pyrénées 2014 (Project CRLE; J-CP and J-ES). MetaToul (Metabolomics & Fluxomics Facitilies, Toulouse, France, http://file:///D:/preeditjobs/npg/leu/leu201781/www.metatoul.fr) is gratefully acknowledged for carrying out metabolome analysis. MetaToul is part of the national infrastructure MetaboHUB-ANR-11-INBS-0010 (The French National infrastructure for metabolomics and fluxomics, www.metabohub.fr). MetaToul is supported by grants from the Région Midi-Pyrénées, the European Regional Development Fund, the SICOVAL, the Infrastructures en Biologie Sante et Agronomie (IBiSa, France), the Centre National de la Recherche Scientifique (CNRS) and the Institut National de la Recherche Agronomique (INRA). We thank Dr Olivier Kosmider (Service d’Hématologie Biologique, Hôpital Cochin, Paris) for genotyping AML samples.
Author contributions
LP designed and performed the experiments, analyzed the data and wrote the manuscript; PS designed the experiments, analyzed the data and wrote the manuscript; SB, LS, ML, RB, CG, CD and MF performed experiments; TLP and J-CP performed glucose flux experiments and analyzed the data; FZ and TTM performed in vivo experiments; VC, FV and TF performed bioinformatics analyses. OH, CR, J-ES, FB, PM and MFo analyzed the data; CL and JT analyzed the data and wrote the manuscript, NC and DB designed and supervised the research program, analyzed the data and wrote the manuscript; and all authors approved the final version of the manuscript.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Leukemia website
Rights and permissions
About this article
Cite this article
Poulain, L., Sujobert, P., Zylbersztejn, F. et al. High mTORC1 activity drives glycolysis addiction and sensitivity to G6PD inhibition in acute myeloid leukemia cells. Leukemia 31, 2326–2335 (2017). https://doi.org/10.1038/leu.2017.81
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/leu.2017.81
This article is cited by
-
The role of bone marrow microenvironment (BMM) cells in acute myeloid leukemia (AML) progression: immune checkpoints, metabolic checkpoints, and signaling pathways
Cell Communication and Signaling (2023)
-
Mitochondrial fusion is a therapeutic vulnerability of acute myeloid leukemia
Leukemia (2023)
-
Spliceosome mutations are associated with clinical response in a phase 1b/2 study of the PLK1 inhibitor onvansertib in combination with decitabine in relapsed or refractory acute myeloid leukemia
Annals of Hematology (2023)
-
PDP1 is a key metabolic gatekeeper and modulator of drug resistance in FLT3-ITD-positive acute myeloid leukemia
Leukemia (2023)
-
G6PD promotes cell proliferation and dexamethasone resistance in multiple myeloma via increasing anti-oxidant production and activating Wnt/β-catenin pathway
Experimental Hematology & Oncology (2022)