Abstract
Polymorphisms (A33512C, C21151T and PAT −/+) of the xeroderma pigmentosum group C (XPC) were shown to contribute to genetic susceptibility to cancer. However, association studies on these polymorphisms in cancer have shown conflicting results. Thus, we performed a meta-analysis. Overall, there was no significant association between 33512C (9,091 patients and 11,553 controls) and cancer risk. No significant association was found in stratification analysis by tumor sites and ethnicities except an elevated lung cancer risk under the recessive genetic model in all subjects [P = 0.04, odds ratio (OR) = 1.20, 95% confidence interval (CI) 1.00–1.45, P heterogeneity = 0.88]. There was no significant association between 21151T (5,227 patients and 5,959 controls) and cancer risk in all subjects but an increased cancer risk in Caucasians under the recessive genetic model (P = 0.006, OR = 1.45, 95% CI 1.11–1.90, P heterogeneity = 0.75) and homozygote comparison (P = 0.02, OR = 1.41, 95% CI 1.07–1.81, P heterogeneity = 0.41). It might be that 21151T increases bladder cancer risk under the recessive genetic model (P = 0.02, OR = 1.49, 95% CI 1.06–2.09, P heterogeneity = 0.47) and homozygote comparison (P = 0.02, OR = 1.49, 95% CI 1.05–2.11, P heterogeneity = 0.23). There was no significant association between PAT + (4,600 patients and 4,866 controls) and cancer risk in all subjects. An increased cancer risk in Caucasians was found under the recessive genetic model (P = 0.02, OR = 1.20, 95% CI 1.03–1.40, P heterogeneity = 0.37) and homozygote comparison (P = 0.008, OR = 1.26, 95% CI 1.06–1.50, P heterogeneity = 0.13). The XPC PAT + allele might increase head and neck cancer risk (P = 0.02, OR = 1.29, 95% CI 1.04–1.59, P heterogeneity = 0.15). More studies based on larger, stratified, case–control population, especially studies investigate the combined effect of XPC A33512C, C21151T, and PAT, are required to further evaluate the role of these polymorphisms in different cancers.
Similar content being viewed by others
Introduction
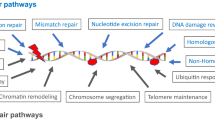
DNA in most cells is regularly damaged by endogenous and exogenous mutagens. DNA repair systems play a critical role in protecting the genome from the insults of cancer-causing agents. Unrepaired damage can result in apoptosis or may lead to unregulated cell growth and then cancer. In humans, more than 70 genes are involved in the four major DNA repair pathways: nucleotide excision repair (NER), base excision repair, mismatch repair, and double-strand-break repair (Hoeijmakers 2001). NER is a versatile repair pathway that can eliminate a wide variety of DNA lesions, including UV-induced photolesions and chemical carcinogen-induced bulky DNA adducts. It is composed of at least two subpathways, global genome repair (GGR) and transcription-coupled repair (TCR) (de Laat et al. 1999). Individuals with decreased NER capacity are at increased risk of cancers. As with many other phenotypic traits, variation in NER capacity may be the result of functional polymorphisms in NER genes. Therefore, it has been hypothesized that inherited polymorphisms in NER genes may modulate susceptibility to cancer.
The xeroderma pigmentosum complementation group C (XPC) protein had been reported to be involved in the early damage recognition and initiation of NER. The XPC protein binds tightly with HR23B (one of two human homologs of Saccharomyces cerevisiae NER factor RAD23), forming the XPC–HR23B complex (Masutani et al. 1997; Sugasawa et al. 1998). Sequence variants of the XPC gene may alter NER capacity and modulate cancer risk. Hollander et al. found deletion of XPC led to lung tumors in mice, and XPC was associated with early events in human lung carcinogenesis (Hollander et al. 2005). Khan et al. discovered an intronic biallelic poly (AT) insertion/deletion polymorphism (PAT) in intron 9 of XPC (Khan et al. 2000). Two nonsynonymous single nucleotide polymorphisms (SNPs), Lys939Gln (an A → C transversion) in exon 15 and Ala499Val (a C → T transition) in exon 8, have also been identified (Hu et al. 2005; Khan et al. 2002).
A large number of molecular epidemiologic studies have been preformed to evaluate the role of XPC polymorphisms in various neoplasms. The Val499Arg (XPC C21151T, rs2228000) and Lys939Gln (XPC A33512C, rs2228001) substitution and a poly (AT) insertion/deletion polymorphism (XPC PAT −/+) in intron 9 are the most thoroughly investigated polymorphism in XPC. However, studies on the XPC A33512C, C21151T and PAT −/+ polymorphisms have shown conflicting results. These polymorphisms might play different roles in different cancers and ethnicities. Even at the same tumor site, considering the possible small effect size of these genetic polymorphisms to cancer and the relatively small sample size in some studies, a small but real association maybe underpowered, which will lead to apparent discrepancy between studies. To assess the association of XPC polymorphisms with the risk of cancer, we conducted a meta-analysis from all eligible case–control studies published to date.
Methods
Identification and eligibility of relevant studies
To identify all studies that examined the association of XPC polymorphisms with cancer, we conducted a computerized literature search of PubMed database (prior to May 2007) using the following keywords and subject terms: “XPC”, “polymorphism” and “cancer”. References of retrieved articles were also screened. Abstracts, case reports, editorials, and review articles were excluded. If an article reported results on different ethnicity subpopulations or tumor sites, each subpopulation or tumor was treated as a separate study in our meta-analysis. Studies included in the meta-analysis had to meet all the following criteria: (1) use an unrelated case–control design, (2) have available genotype frequency, and (3) genotype distribution of control population must be in Hardy–Weinberg equilibrium (HWE).
Data extraction
Data were collected on the genotype of A33512C, C21151T, and PAT −/+ according to different kinds of cancers. First author, year of publication, ethnicity of study population, number of cases and controls, and allele frequency were described (Table 1).
Statistical analysis
The strength of the association between XPC polymorphisms and cancer was measured by odds ratio (OR) corresponding to 95% confidence interval (CI), which was calculated according to the method of Woolf (1955). We examined the association between allele C of XPC A33512C and cancer risk, as well as the dominant genetic model (CC + CA vs. AA), the recessive genetic model (CC vs. CA + AA), homozygote comparison (CC vs. AA), CC vs. CA contrast, and CA vs. AA contrast. The same method was applied to analysis of the C21151T and PAT −/+ polymorphisms. We conducted two models of meta-analysis for dichotomous outcomes in Review-Manager 4.2 software: the fixed-effects model and the random-effects model. A fixed-effects model using the Mantel–Haenszel method assumes that studies are sampled from populations with the same effect size, making an adjustment to the study weights according to the in-study variance. A random-effects model assumes that studies are taken from populations with varying effect sizes and calculates study weights both from in-study and between-study variances, with consideration of the extent of variation, or heterogeneity. A chi-square-based Q statistic test was performed to assess the between-study heterogeneity (Lau et al. 1997). Heterogeneity was considered significant for P < 0.10. A random-effects model (if P < 0.10) or a fixed-effects model (if P > 0.10) was used to pool the results (Petitti 1994). The significance of the pooled OR was determined by the Z test. A P value of <0.05 was considered significant.
Subgroup analysis was stratified by the study characteristics of ethnicity and tumor site, respectively. Tumor sites only investigated once in all the studies were grouped as “other cancers”.
Publication bias was investigated with the funnel plot, in which the standard error of ln(OR) of each study was plotted against its OR. Funnel-plot asymmetry was further assessed by the method of Egger’s linear regression test (Egger et al. 1997). The significance of the intercept was determined by the t test, and a P value of <0.05 was considered significant. Hardy–Weinberg equilibrium was tested by the chi-square test for goodness of fit with a Web program (http://www.ihg.gsf.de/cgi-bin/hw/hwa1.pl). Analyses were performed using the software Stata version 7, ReviewManager 4.2 (Oxford, England). All P values were two-sided.
Results
Study inclusion
Through literature search and selection based on the inclusion criteria, 32 articles (37 studies) were found, and 28 articles (Bai et al. 2007; Blankenburg et al. 2005; Casson et al. 2005; De Ruyck et al. 2007; Festa et al. 2005; Hansen et al. 2007; Hirata et al. 2007; Hu et al. 2005; Huang et al. 2006; Kietthubthew et al. 2006; Lee et al. 2005; Li et al. 2006; Marin et al. 2004; Mechanic et al. 2006; Nelson et al. 2005; Sak et al. 2005, 2006; Sanyal et al. 2004; Shen et al. 2001, 2005; Sugimura et al. 2006; Vogel et al. 2005; Wang et al. 2006; Weiss et al. 2005; Yang et al. 2005; Ye et al. 2006; Zhou et al. 2006; Zhu et al. 2007) (32 studies) met our inclusion criteria, as listed in Table 1. One study of A33512C (Hirata et al. 2006) reported an extremely high variant allele frequency, which may result from wrong allele counting or poor genotyping quality, and was finally excluded from our meta-analysis.
Among the 28 eligible articles, 18 articles (Bai et al. 2007; Blankenburg et al. 2005; Festa et al. 2005; Hansen et al. 2007; Hirata et al. 2007; Hu et al. 2005; Huang et al. 2006; Kietthubthew et al. 2006; Lee et al. 2005; Li et al. 2006; Mechanic et al. 2006; Sak et al. 2005; Sanyal et al. 2004; Vogel et al. 2005; Weiss et al. 2005; Ye et al. 2006; Zhou et al. 2006; Zhu et al. 2007) (23 studies) described A33512C, ten articles (Bai et al. 2007; Hu et al. 2005; Huang et al. 2006; Lee et al. 2005; Li et al. 2006; Sak et al. 2006; Shen et al. 2005; Weiss et al. 2005; Zhou et al. 2006; Zhu et al. 2007) (11 studies) described C21151T, and 13 articles (Blankenburg et al. 2005; Casson et al. 2005; De Ruyck et al. 2007; Kietthubthew et al. 2006; Li et al. 2006; Marin et al. 2004; Nelson et al. 2005; Sak et al. 2005; Shen et al. 2001; Sugimura et al. 2006; Wang et al. 2006; Yang et al. 2005; Zhu et al. 2007) (14 studies) described PAT −/+; 82.1% (23/28) stated that the age and gender status were matched between case and control population. All studies used blood sample for genotyping.
In all the eligible articles, Zhou et al. (2006) provided data on two kinds of cancers: esophageal squamous cell carcinoma (ESCC) and gastric cardiac adenocarcinoma (GcAde). Nelson et al. (2005) provided data on two kinds of cancers: basal cell carcinoma (BCC) and squamous cell carcinoma (SCC). Thus, each type of cancer in these two articles was treated as a separate study in our meta-analysis (Table 2). Mechanic et al. (2006) provided data on subjects of two ethnicities: Africa American and white. Similarly, these two ethnicities were treated as separate studies. Studies providing genotyping data of the population of America were indicated as “mixed” ethnic (Huang et al. 2006; Nelson et al. 2005; Weiss et al. 2005).
Summary statistics
The allele frequencies were calculated for controls from the corresponding genotype distributions (Table 2). The A33512 allele had a higher representation among controls of Asian descent (65.2%, 95% CI 61.1–69.4) than in controls of European descent (61.3%, 95% CI 59.7–62.8). The C21151 allele had a lower representation among controls of Asian descent (69.2%, 95% CI 66.0–72.4) than in controls of European descent (74.5%, 95% CI 69.9–79.1). The PAT− allele had a higher representation among controls of Asian descent (67.0%, 95% CI 61.0–73.1) than in controls of European descent (59.4%, 95% CI 57.4–61.7). The allele frequencies of these three polymorphisms did not show big differences between Asians and Caucasians. Overall, the prevalence of A33512, C21151, and PAT− allele was 63.0%, 72.1%, and 62.4% in controls, respectively (Table 2).
Quantitative synthesis
XPC A33512C
The fixed-effects model was used to pool the result, as the between-study heterogeneity was insignificant. There was no significant association between the 33512C allele and cancer risk in all subjects (P = 0.60, OR = 1.01, 95% CI 0.97–1.05, P heterogeneity = 0.34), as well as in Asians or Caucasians. However, under the recessive genetic model, an elevated but not significant association between CC genotype and cancer risk was found in all subjects (P = 0.05, OR = 1.08, 95% CI 1.00–1.18, P heterogeneity = 0.25); CC genotype showed a significant association with cancer risk in all subjects in analysis of CC vs. CA contrast (P = 0.03, OR = 1.11, 95% CI 1.01–1.20, P heterogeneity = 0.26) (Table 3).
The CC genotype contributed to an elevated risk of lung cancer under both the recessive genetic model (P = 0.04, OR = 1.20, 95% CI 1.00–1.45, P heterogeneity = 0.88, Fig. 1) and homozygote–heterozygote (CC vs. CA) comparison (P = 0.05, OR = 1.21, 95% CI 1.00–1.46, P heterogeneity = 0.96). An increased risk of colorectal cancer was found when the CA genotype was compared with the AA genotype (P = 0.002, OR = 1.32, 95% CI 1.11–1.56, P heterogeneity = 0.14). No evidence of association between 33512C and other cancers was found (Table 3).
Meta-analysis for the xeroderma pigmentosum group C (XPC) A33512C polymorphisms and cancer stratified according to different tumor sites: CC vs. (CA + AA). The study is shown by a point estimate of the odds ratio (OR) and the accompanying 95% confidence interval (CI) using a fixed-effects model. n indicates the total number of CC; N indicates the total number of individuals
XPC C21151T
Significant between-study heterogeneity existed in 11 studies when we compared C21151T C and the T allele in different kinds of cancers (P heterogeneity = 0.03). The random-effects model was used to pool the result. There was no significant association between the 21151T allele and cancer risk (P = 0.74, OR = 0.99, 95% CI 0.90–1.07). An elevated but not significant association between 21151T and cancer risk was found under the recessive genetic model in all subjects (P = 0.06, OR = 1.14, 95% CI 1.00–1.32, P heterogeneity = 0.40), and a significantly elevated association was found in analysis of TT vs. TC contrast in all subjects (P = 0.01, OR = 1.21, 95% CI 1.04–1.40, P heterogeneity = 0.58). In Caucasians, the association was significant under the recessive genetic model (P = 0.006, OR = 1.45, 95% CI 1.11–1.90, P heterogeneity = 0.75, Fig. 3a), homozygote comparison (P = 0.02, OR = 1.41, 95% CI 1.07–1.85, P heterogeneity = 0.54), and TT vs. TC contrast (P = 0.004, OR = 1.52, 95% CI 1.15–2.01, P heterogeneity = 0.94) (Table 3).
In the subgroup analysis for different tumor sites, 21151T had an effect of increasing the bladder cancer risk under the recessive genetic model (P = 0.02, OR = 1.49, 95% CI 1.06–2.09, P heterogeneity = 0.47, Fig. 2a), homozygote comparison (P = 0.02, OR = 1.49, 95% CI 1.05–2.11, P heterogeneity = 0.33), and TT vs. TC contrast (P = 0.03, OR = 1.49, 95% CI 1.05–2.13, P heterogeneity = 0.76) in all subjects. No evidence of association between 21151T and other cancers was found (Table 3).
Meta-analysis for the xeroderma pigmentosum group C (XPC) C21151T and poly (AT) insertion/deletion polymorphism (PAT) −/+ polymorphisms and cancer stratified according to different tumor sites. a C21151T: TT vs. (TC + CC). b PAT −/+: + vs. −. The study is shown by a point estimate of the odds ratio (OR) and the accompanying 95% confidence interval (CI) using a fixed-effects model. n indicates the total number of TT (a) or + (b); N indicates the total number of individuals (a) or alleles (B)
XPC PAT
Significant heterogeneity existed in 14 studies when we compared XPC PAT − and + allele in different kinds of cancers. The random-effects model was used to pool the result. There was no significant association between the PAT + allele and cancer risk in all subjects (P = 0.72, OR = 1.02, 95% CI 0.94–1.11, P heterogeneity = 0.03, Fig. 2b). The PAT + allele appeared to increase the cancer risk under tje recessive genetic model (P = 0.02, OR = 1.20, 95% CI 1.03–1.40, P heterogeneity = 0.37, Fig. 3b) and homozygote comparison (P = 0.008, OR = 1.26, 95% CI 1.06–1.50, P heterogeneity = 0.13) in Caucasians (Table 3).
Meta-analysis for the xeroderma pigmentosum group C (XPC) C21151T and poly (AT) insertion/deletion polymorphism (PAT) −/+ polymorphisms and cancer in Caucasians. a C21151T: TT vs. (TC + CC). b PAT −/+: ++ vs. (+− and − −). The study is shown by a point estimate of the odds ratio (OR) and the accompanying 95% confidence interval (CI) using a fixed-effects model. n indicates the total number of TT (a) or ++ (b); N indicates the total number of individuals
In the subgroup analysis for different tumor sites, PAT + allele contributed to an increased head and neck cancer risk (P = 0.02, OR = 1.29, 95% CI 1.04–1.59, P heterogeneity = 0.15, Fig. 2b), as well as under dominant genetic model and homozygote comparison in all subjects (Table 3).
Publication bias
The funnel plot was applied for comparison of 33512C vs. 33512A in the OR analysis of XPC A33521C, and Egger’s test provided no evidence for funnel-plot asymmetry (t = 1.87, P = 0.076). Similarly, no publication bias was detected for C21151T and PAT −/+ polymorphisms (t = −0.60, P = 0.565; t = 0.64, P = 0.671, respectively; Fig. 4).
Begg’s funnel plot of the Egger’s test of allele comparison for publication bias. a Funnel plot for C vs. A allele comparison in A33512C polymorphism; b funnel plot for T vs. C allele in C21151T polymorphism; c funnel plot for + vs. − allele in poly (AT) insertion/deletion polymorphism (PAT). No asymmetry was found as indicated by the P value of Egger’s test
Discussion
Sanyal et al. first reported that the frequency of the variant C allele of XPC A33512C polymorphism was significantly higher in bladder cancer cases of Caucasian than in controls (P = 0.001, OR = 1.49, 95% CI 1.16–1.92) in 2004 (Sanyal et al. 2004). Thereafter, more and more studies were conducted to further access the association in different tumor sites across different nations. However, the results were fairly confusing rather than conclusive. Most studies could not confirm a significantly increased risk between cancers and 33512C allele. Khan et al. studied the function of the XPC A33512C alteration in an allele-specific post-UV reaction assay in fibroblast cell (Khan et al. 2000). They found that XPC 33512C allele was equally as efficient as A33512 allele, indicating both polymorphisms were fully functional in DNA repair. Our meta-analysis did not reveal a significant association between the 33512C and cancer risk compared with 33512A. However, the CC genotype contributed to an elevated risk of cancer.
No functional data of the XPC C21151T polymorphism was reported. Our meta-analysis indicated 21151T had an effect of increasing the bladder cancer risk under the recessive-genetic model and homozygote comparison, and contributed to an increased cancer risk under the recessive genetic model and homozygote comparison in Caucasians. We found that the cancer risk in Caucasians with XPC PAT + allele increased under the recessive genetic model and homozygote comparison, and XPC PAT + carriers had an elevated head and neck cancer risk under the dominant genetic model and homozygote comparison. The same polymorphism may play different roles in cancer susceptibility in different tumor sites. The XPC PAT + allele might contribute to a higher risk of head and neck cancer but had no effect on the susceptibility of lung cancer, oral cancer, and bladder cancer.
It is interesting that none of the three XPC polymorphisms had a significant effect in Asians, and the variant homozygote genotypes of XPC C21151T and XPC PAT −/+ might increase cancer risk in Caucasians. Due to the difference in different populations, it is necessary to stratify the ethnicity in the tumor-sites analysis. The same polymorphism may play different roles across different ethnicities because of different genetic background.
The PAT + allele is in linkage disequilibrium with the A allele of an intronic SNP (IVS11-6) in intron 11. It appears that IVS11 6A affects alternative splicing and increases the frequency of deletion of exon 12. The XPC splicing isoform without exon 12 had reduced DNA repair activity (Khan et al. 2002). PAT −/+ might not be a causal SNP, and the increased cancer susceptibility of PAT + carriers may arise from the linkage with IVS11-6 A. Blankenburg et al.( 2005) performed a hospital-based case–control study with 294 cutaneous melanoma cases and 375 gender-matched controls. They found XPC intron 9, PAT +, intron 11-6A, and exon 15 33512C polymorphisms were in linkage disequilibrium. The role of PAT + in head and neck cancer still needs further investigation, as the cases and controls involved is too small; analysis of 33512C did not provide data on head and neck cancer. We could not make a comparison between PAT + and 33512C because the studies included in the meta-analysis of these two polymorphisms were different, which should have the same results due to the linkage disequilibrium.
A single polymorphism likely has weak effects on the individual’s phenotype. It may not be measurable except in the context of some supporting environmental factors, such as smoking. We tried to evaluate the effect of smoking on the susceptibility of XPC A33512C polymorphism on cancer risk. Three studies from two articles (Hansen et al. 2007; Zhou et al. 2006) (esophageal squamous cell carcinoma, gastric cardiac adenocarcinoma, colorectal cancer) were recruited for analysis, as their stratification data on smoking were available. We found the cancer risk in smokers carrying the 33512C allele was not higher (P = 0.86, OR = 1.01, 95% CI 0.87–1.18, P heterogeneity = 0.47) than that of the nonsmokers (P = 0.05, OR = 1.18, 95% CI 1.00–1.40, P heterogeneity = 0.83). However, smoking has different effects on different cancer types. It is still necessary to stratify cancer types; the study size was relatively small (975 cancer patients and 1,409 controls). So, this conclusion should be treated as preliminary.
It is necessary to access the combined effect of several polymorphisms, as interaction of different polymorphisms in the same gene or between different genes might contribute to cancer risk. Several articles in our meta-analysis evaluated the combined effect of XPC A33512C, C21151T, and PAT −/+ or two of these three polymorphisms (Blankenburg et al. 2005; Hu et al. 2005; Huang et al. 2006; Li et al. 2006; Sak et al. 2005; Shen et al. 2005; Weiss et al. 2005; Zhu et al. 2007). Hu et al. (2005) examined the combined effect of XPC A33512C and C21151T on lung cancer risk. They found that cases with both 21151CT/TT and 33512AC/CC variant genotypes had a significantly increased lung cancer risk compared with those having both wild-type genotypes (21151CC and 33512AA); smokers with both variant genotypes (21151CT/TT and 33512AC/CC) had the highest lung cancer risk (adjusted OR = 7.36; 95% CI 3.19–17.00) compared with that of nonsmokers. Zhu et al. (2007) evaluated the combined effect of the there polymorphisms and found a protective effect of the haplotype 21151C-PAT− -33512C. We tried to evaluate the combined effect of these polymorphisms on the susceptibility of cancer. Unfortunately, the available data was not compatible. More studies should be carried out to examine the combined effect of these three polymorphisms in different kinds of cancers.
Chance effects, as with false negatives (underpowered studies) and false positives (type I error), together with the true variability among populations, might lead to conflicting conclusions across different studies. In our meta-analysis, false negative and false positive findings would neutralize each other, as a relatively large number of studies were included. However, publication of the findings may depend on the expectation of the researchers. False-negative results may be suppressed and false-positive results magnified (Salanti et al. 2005). Thus, the validity of conclusions in our meta-analysis may be affected. The inclusion of unpublished data is commonly suggested as a means of reducing the impact of false-positive and publication bias. However, in practice, most of the unpublished studies were not available. Although there was no significant publication bias in our meta-analysis, the results may still be affected by the false positive (type I error).
Five genetic contrasts (dominant genetic model, recessive genetic model, and three pairwise comparisons) were considered, as were allelic association; the results under different models were inconsistent. As shown in Table 3, there was an interesting tendency that most significant associations found in our study were under the recessive genetic model, homozygote comparison, and homozygote–heterozygote comparison. The excess of allele homozygotes but not heterozygotes was considered a risk among cancer patients.
In conclusion, our meta-analysis investigated the associations between the three XPC polymorphisms and cancer risk with a total of 12,408 cancer patients and 14,984 controls from 32 case–control studies. Overall, our meta-analysis suggested no significant associations between XPC 33512C, 21151T, and PAT + in Asians. However, 21151TT and PAT ++ might increase cancer risk in Caucasians, which indicated a big difference among different populations. In all subjects, XPC 33512C, 21151T, and PAT + might increase lung cancer, bladder cancer, and head and neck cancer risks under different genetic models, respectively. The present results suggest association only in particular ethnic backgrounds and/or tumor sites, and more studies based on larger, stratified case–control populations are still needed to clarify the different effects of these polymorphisms in Asians and Caucasians. Studies investigating the combined effect of XPC A33512C, C21151T, and PAT will be very important to further evaluate the role of these polymorphisms in different cancers.
References
Bai Y, Xu L, Yang X, Hu Z, Yuan J, Wang F, Shao M, Yuan W, Qian J, Ma H, Wang Y, Liu H, Chen W, Yang L, Jing G, Huo X, Chen F, Liu Y, Jin L, Wei Q, Huang W, Shen H, Lu D, Wu T (2007) Sequence variations in DNA repair gene XPC is associated with lung cancer risk in a Chinese population: a case–control study. BMC Cancer 7:81
Blankenburg S, Konig IR, Moessner R, Laspe P, Thoms KM, Krueger U, Khan SG, Westphal G, Berking C, Volkenandt M, Reich K, Neumann C, Ziegler A, Kraemer KH, Emmert S (2005) Assessment of 3 xeroderma pigmentosum group C gene polymorphisms and risk of cutaneous melanoma: a case–control study. Carcinogenesis 26:1085–1090
Casson AG, Zheng Z, Evans SC, Veugelers PJ, Porter GA, Guernsey DL (2005) Polymorphisms in DNA repair genes in the molecular pathogenesis of esophageal (Barrett) adenocarcinoma. Carcinogenesis 26:1536–1541
de Laat WL, Jaspers NG, Hoeijmakers JH (1999) Molecular mechanism of nucleotide excision repair. Genes Dev 13:768–785
De Ruyck K, Szaumkessel M, De Rudder I, Dehoorne A, Vral A, Claes K, Velghe A, Van Meerbeeck J, Thierens H (2007) Polymorphisms in base-excision repair and nucleotide-excision repair genes in relation to lung cancer risk. Mutat Res 631(2):101–110
Egger M, Davey Smith G, Schneider M, Minder C (1997) Bias in meta-analysis detected by a simple, graphical test. Bmj 315:629–634
Festa F, Kumar R, Sanyal S, Unden B, Nordfors L, Lindholm B, Snellman E, Schalling M, Forsti A, Hemminki K (2005) Basal cell carcinoma and variants in genes coding for immune response, DNA repair, folate and iron metabolism. Mutat Res 574:105–111
Hansen RD, Sorensen M, Tjonneland A, Overvad K, Wallin H, Raaschou-Nielsen O, Vogel U (2007) XPA A23G, XPC Lys939Gln, XPD Lys751Gln and XPD Asp312Asn polymorphisms, interactions with smoking, alcohol and dietary factors, and risk of colorectal cancer. Mutat Res 619:68–80
Hirata H, Hinoda Y, Matsuyama H, Tanaka Y, Okayama N, Suehiro Y, Zhao H, Urakami S, Kawamoto K, Kawakami T, Igawa M, Naito K, Dahiya R (2006) Polymorphisms of DNA repair genes are associated with renal cell carcinoma. Biochem Biophys Res Commun 342:1058–1062
Hirata H, Hinoda Y, Tanaka Y, Okayama N, Suehiro Y, Kawamoto K, Kikuno N, Majid S, Vejdani K, Dahiya R (2007) Polymorphisms of DNA repair genes are risk factors for prostate cancer. Eur J Cancer 43:231–237
Hoeijmakers JH (2001) Genome maintenance mechanisms for preventing cancer. Nature 411:366–374
Hollander MC, Philburn RT, Patterson AD, Velasco-Miguel S, Friedberg EC, Linnoila RI, Fornace AJ Jr (2005) Deletion of XPC leads to lung tumors in mice and is associated with early events in human lung carcinogenesis. Proc Natl Acad Sci USA 102:13200–13205
Hu Z, Wang Y, Wang X, Liang G, Miao X, Xu Y, Tan W, Wei Q, Lin D, Shen H (2005) DNA repair gene XPC genotypes/haplotypes and risk of lung cancer in a Chinese population. Int J Cancer 115:478–483
Huang WY, Berndt SI, Kang D, Chatterjee N, Chanock SJ, Yeager M, Welch R, Bresalier RS, Weissfeld JL, Hayes RB (2006) Nucleotide excision repair gene polymorphisms and risk of advanced colorectal adenoma: XPC polymorphisms modify smoking-related risk. Cancer Epidemiol Biomarkers Prev 15:306–311
Khan SG, Metter EJ, Tarone RE, Bohr VA, Grossman L, Hedayati M, Bale SJ, Emmert S, Kraemer KH (2000) A new xeroderma pigmentosum group C poly(AT) insertion/deletion polymorphism. Carcinogenesis 21:1821–1825
Khan SG, Muniz-Medina V, Shahlavi T, Baker CC, Inui H, Ueda T, Emmert S, Schneider TD, Kraemer KH (2002) The human XPC DNA repair gene: arrangement, splice site information content and influence of a single nucleotide polymorphism in a splice acceptor site on alternative splicing and function. Nucleic Acids Res 30:3624–3631
Kietthubthew S, Sriplung H, Au WW, Ishida T (2006) Polymorphism in DNA repair genes and oral squamous cell carcinoma in Thailand. Int J Hyg Environ Health 209:21–29
Lau J, Ioannidis JP, Schmid CH (1997) Quantitative synthesis in systematic reviews. Ann Intern Med 127:820–826
Lee GY, Jang JS, Lee SY, Jeon HS, Kim KM, Choi JE, Park JM, Chae MH, Lee WK, Kam S, Kim IS, Lee JT, Jung TH, Park JY (2005) XPC polymorphisms and lung cancer risk. Int J Cancer 115:807–813
Li C, Hu Z, Liu Z, Wang LE, Strom SS, Gershenwald JE, Lee JE, Ross MI, Mansfield PF, Cormier JN, Prieto VG, Duvic M, Grimm EA, Wei Q (2006) Polymorphisms in the DNA repair genes XPC, XPD, and XPG and risk of cutaneous melanoma: a case–control analysis. Cancer Epidemiol Biomarkers Prev 15:2526–2532
Marin MS, Lopez-Cima MF, Garcia-Castro L, Pascual T, Marron MG, Tardon A (2004) Poly (AT) polymorphism in intron 11 of the XPC DNA repair gene enhances the risk of lung cancer. Cancer Epidemiol Biomarkers Prev 13:1788–1793
Masutani C, Araki M, Sugasawa K, van der Spek PJ, Yamada A, Uchida A, Maekawa T, Bootsma D, Hoeijmakers JH, Hanaoka F (1997) Identification and characterization of XPC-binding domain of hHR23B. Mol Cell Biol 17:6915–6923
Mechanic LE, Millikan RC, Player J, de Cotret AR, Winkel S, Worley K, Heard K, Heard K, Tse CK, Keku T (2006) Polymorphisms in nucleotide excision repair genes, smoking and breast cancer in African Americans and whites: a population-based case–control study. Carcinogenesis 27:1377–1385
Nelson HH, Christensen B, Karagas MR (2005) The XPC poly-AT polymorphism in nonmelanoma skin cancer. Cancer Lett 222:205–209
Petitti DB (1994) Meta-analysis, decision analysis, and cost-effectiveness analysis. Oxford University Press, Oxford
Sak SC, Barrett JH, Paul AB, Bishop DT, Kiltie AE (2005) The polyAT, intronic IVS11–6 and Lys939Gln XPC polymorphisms are not associated with transitional cell carcinoma of the bladder. Br J Cancer 92:2262–2265
Sak SC, Barrett JH, Paul AB, Bishop DT, Kiltie AE (2006) Comprehensive analysis of 22 XPC polymorphisms and bladder cancer risk. Cancer Epidemiol Biomarkers Prev 15:2537–2541
Salanti G, Sanderson S, Higgins JP (2005) Obstacles and opportunities in meta-analysis of genetic association studies. Genet Med 7:13–20
Sanyal S, Festa F, Sakano S, Zhang Z, Steineck G, Norming U, Wijkstrom H, Larsson P, Kumar R, Hemminki K (2004) Polymorphisms in DNA repair and metabolic genes in bladder cancer. Carcinogenesis 25:729–734
Shen H, Sturgis EM, Khan SG, Qiao Y, Shahlavi T, Eicher SA, Xu Y, Wang X, Strom SS, Spitz MR, Kraemer KH, Wei Q (2001) An intronic poly (AT) polymorphism of the DNA repair gene XPC and risk of squamous cell carcinoma of the head and neck: a case–control study. Cancer Res 61:3321–3325
Shen M, Berndt SI, Rothman N, Demarini DM, Mumford JL, He X, Bonner MR, Tian L, Yeager M, Welch R, Chanock S, Zheng T, Caporaso N, Lan Q (2005) Polymorphisms in the DNA nucleotide excision repair genes and lung cancer risk in Xuan Wei, China. Int J Cancer 116:768–773
Sugasawa K, Ng JM, Masutani C, Iwai S, van der Spek PJ, Eker AP, Hanaoka F, Bootsma D, Hoeijmakers JH (1998) Xeroderma pigmentosum group C protein complex is the initiator of global genome nucleotide excision repair. Mol Cell 2:223–232
Sugimura T, Kumimoto H, Tohnai I, Fukui T, Matsuo K, Tsurusako S, Mitsudo K, Ueda M, Tajima K, Ishizaki K (2006) Gene-environment interaction involved in oral carcinogenesis: molecular epidemiological study for metabolic and DNA repair gene polymorphisms. J Oral Pathol Med 35:11–18
Vogel U, Overvad K, Wallin H, Tjonneland A, Nexo BA, Raaschou-Nielsen O (2005) Combinations of polymorphisms in XPD, XPC and XPA in relation to risk of lung cancer. Cancer Lett 222:67–74
Wang L, Lin DX, Lu XH, Miao XP (2006) [Polymorphisms of the DNA repair genes XRCC1 and XPC: relationship to pancreatic cancer risk]. Wei Sheng Yan Jiu 35:534–536
Weiss JM, Weiss NS, Ulrich CM, Doherty JA, Voigt LF, Chen C (2005) Interindividual variation in nucleotide excision repair genes and risk of endometrial cancer. Cancer Epidemiol Biomarkers Prev 14:2524–2530
Woolf B (1955) On estimating the relation between blood group and disease. Ann Hum Genet 19:251–253
Yang M, Kang MJ, Choi Y, Kim CS, Lee SM, Park CW, Lee HS, Tae K (2005) Associations between XPC expression, genotype, and the risk of head and neck cancer. Environ Mol Mutagen 45:374–379
Ye W, Kumar R, Bacova G, Lagergren J, Hemminki K, Nyren O (2006) The XPD 751Gln allele is associated with an increased risk for esophageal adenocarcinoma: a population-based case–control study in Sweden. Carcinogenesis 27:1835–1841
Zhou RM, Li Y, Wang N, Zhang XJ, Dong XJ, Guo W (2006) [Correlation of XPC Ala499Val and Lys939Gln polymorphisms to risks of esophageal squamous cell carcinoma and gastric cardiac adenocarcinoma]. Ai Zheng 25:1113–1119
Zhu Y, Lai M, Yang H, Lin J, Huang M, Grossman HB, Dinney CP, Wu X (2007) Genotypes, haplotypes and diplotypes of XPC and risk of bladder cancer. Carcinogenesis 28:698–703
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Zhang, D., Chen, C., Fu, X. et al. A meta-analysis of DNA repair gene XPC polymorphisms and cancer risk. J Hum Genet 53, 18–33 (2008). https://doi.org/10.1007/s10038-007-0215-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10038-007-0215-5
Keywords
This article is cited by
-
Predictive genetic markers in neoadjuvant chemoradiotherapy for locally advanced esophageal cancer: a long way to go. Review of the literature
The Pharmacogenomics Journal (2018)
-
Polymorphisms in the XPC gene affect urinary bladder cancer risk: a case-control study, meta-analyses and trial sequential analyses
Scientific Reports (2016)
-
Potential risk of esophageal squamous cell carcinoma due to nucleotide excision repair XPA and XPC gene variants and their interaction among themselves and with environmental factors
Tumor Biology (2016)
-
In silico dissection of miRNA targetome polymorphisms and their role in regulating miRNA-mediated gene expression in esophageal cancer
Cell Biochemistry and Biophysics (2016)
-
Associations between polymorphisms of the XPC gene and lung cancer susceptibility: a meta-analysis
Tumor Biology (2014)