Abstract
Molecular chaperones assist protein folding, and variations in their encoding genes may be disease-causing in themselves or influence the phenotypic expression of disease-associated or susceptibility-conferring variations in many different genes. We have screened three candidate patient groups for variations in the HSPD1 and HSPE1 genes encoding the mitochondrial Hsp60/Hsp10 chaperone complex: two patients with multiple mitochondrial enzyme deficiency, 61 sudden infant death syndrome cases (MIM: #272120), and 60 patients presenting with ethylmalonic aciduria carrying non-synonymous susceptibility variations in the ACADS gene (MIM: *606885 and #201470). Besides previously reported variations we detected six novel variations: two in the bidirectional promoter region, and one synonymous and three non-synonymous variations in the HSPD1 coding region. One of the non-synonymous variations was polymorphic in patient and control samples, and the rare variations were each only found in single patients and absent in 100 control chromosomes. Functional investigation of the effects of the variations in the promoter region and the non-synonymous variations in the coding region indicated that none of them had a significant impact. Taken together, our data argue against the notion that the chaperonin genes play a major role in the investigated diseases. However, the described variations may represent genetic modifiers with subtle effects.
Similar content being viewed by others
Introduction
In many monogenic diseases, the phenotype is not solely determined by gene variations in the primary target gene, but is subject to modification by so-called modifier genes and environmental influences (Dipple and McCabe 2000; Nadeau 2001). In complex multifactorial diseases, no single gene defect with a large impact is present, but rather a set of gene variants (also termed modifiers), each exerting a small effect in itself, can act together with environmental factors to produce the disease phenotype (Botstein and Risch 2003).
Modifying effects may be brought about by influencing the level of expression of a target gene product, its intracellular trafficking and folding, posttranslational modifications, activity and/or structural maintenance, and turnover. Molecular chaperones assist and supervise intracellular protein folding and are, together with specific proteases, involved in clearance of misfolded and damaged proteins. They typically interact with a large number of different proteins, and it has been proposed that genetic variations in chaperone genes may function as modifiers of amino acid substituting variations involved in many traits/diseases (Slavotinek and Biesecker 2001). Genetic screens of neurodegenerative disease models have indeed shown that molecular chaperones, intracellular proteases and other components involved in protein quality control represent a prominent group of such modifier genes (Fernandez-Funez et al. 2000; Bonini and Fortini 2003; Nollen et al. 2004). It has furthermore been demonstrated that even a simple decrease in the levels of the Hsp90 chaperone may enable a number of otherwise suppressed phenotypic traits to be expressed in natural fruitfly and Arabidopsis populations (Rutherford and Lindquist 1998; Queitsch et al. 2002). A similar buffering function against deleterious variations has also been accomplished by experimentally increasing the levels of the bacterial GroEL/ES chaperone complex (Fares et al. 2002).
The mammalian mitochondrial homolog of the bacterial GroEL/GroES chaperone system consists of Hsp60 and Hsp10. It is involved in folding of a subset of mitochondrial proteins (Dubaquie et al. 1998). Homozygosity for deleterious mutations in the Drosophila melanogaster gene encoding the fly Hsp60 homolog causes death early in embryogenesis establishing that Hsp60 is an essential protein (Perezgasga et al. 1999). We have shown that knock-down of the HSPD1 gene by RNAi in human cells compromises folding of the mitochondrial matrix enzymes short-chain acyl-CoA dehydrogenase (SCAD) and medium-chain acyl-CoA dehydrogenase (MCAD), encoded by the ACADS and ACADM genes respectively, and mitochondrially targeted green fluorescent protein (Corydon et al. 2005). In addition, disease-causing mutant variants in the medium- and short-chain acyl-CoA dehydrogenases have been shown to remain in complex with the Hsp60 chaperonin for prolonged periods of time (Saijo et al. 1994; Pedersen et al. 2003), and elevated chaperonin levels could partially rescue the correct folding of some of them (Andresen et al. 2001; Bross et al. 1993, 1995).
We have recently reported the genomic structure of the human chaperonin genes and variations in these genes in a small set of independent control chromosomes (Hansen et al. 2003). Our genetic investigation of a French family with an autosomal dominant form of spastic paraplegia (SPG13; MIM 605280) identified a missense mutation in the HSPD1 gene (c.292G>A/p.Val98Ile) that segregated with the disease phenotype and was shown to compromise chaperonin function (Hansen et al. 2002).
To further extend the search for disease-associated variations in the HSPD1/HSPE1 genes we have in the present study screened three patient groups, in which an involvement of the Hsp60/Hsp10 chaperonin system has been indicated or suggested.
Primary disease gene group
Two patients reported with multiple mitochondrial enzyme deficiency (Agsteribbe et al. 1993) who displayed decreased activities of a number of mitochondrial metabolic pathways; Hsp60 protein levels in cultured fibroblast cells from one of these patients were shown to be decreased (Briones et al. 1997) suggesting that a defect in the Hsp60 chaperonin might cause deficient folding of various enzymes involved in these pathways.
Modifier gene group
Sixty patients presenting with ethylmalonic aciduria who carried either of two polymorphic SCAD missense variations conferring susceptibility to SCAD deficiency (Corydon et al. 2001); although both variant SCAD proteins are functional, they have been shown to cause prolonged association of the variant SCAD proteins with the Hsp60 chaperonin (Pedersen et al. 2003).
Genetic predisposition group
Sixty-one cases of sudden infant death syndrome (SIDS); RFLP analysis has indicated overrepresentation of a certain HSPD1 genotype in a group of SIDS cases (Rahim et al. 1996), and heat stress has been reported as risk factor for SIDS (Byard and Krous 2003).
In these patients, we sequenced all exons, exon-intron junctions and the bidirectional promoter region. The frequencies of the detected variations were analysed in control individuals, and functional analysis of potential effects of variations in the promoter region on transcription and of non-synonymous variations in the coding region on in vivo chaperonin function was performed.
Materials and methods
Patients
SIDS cases
The 61 patients included in the present study form part of an extensive forensic material comprising cases of sudden, unexpected death in infants and young children (0–10 years old) that have been autopsied at the Institute of Forensic Medicine, University of Aarhus, in the years 1989–1998. A part of the samples has previously been used in an investigation of the possible relationship between fatty acid oxidation disorders and sudden death in infancy (Lundemose et al. 1997). DNA was purified from cultured primary fibroblast cells isolated from Achilles tendon biopsies taken at the autopsy. The ethical committee (Den Videnskabsetiske Komité for Århus Amt) has approved the use of the SIDS material in the present study.
Patients with elevated urinary excretion of ethylmalonic acid and presence of SCAD susceptibility variations (hereafter called EMA/SCAD patients)
Patients referred to metabolic centres in Europe and the USA that displayed elevated levels of ethylmalonic acid in urine (>18 mmol/mol creatinine), and/or other biochemical parameters indicating SCAD deficiency, are routinely submitted to our laboratory for genetic analysis. A large fraction of these patients carry the polymorphic c.511C/T and/or c.625G/A variations in the ACADS gene in both alleles or in combination with rare, likely disease-causing mutations in the other allele. The polymorphic variations are present in homozygous or compound homozygous form in 14% of the population but strongly over-represented in patients indicating that other factors contribute. Based on the finding that interaction of the polymorphic SCAD variant proteins with the Hsp60/Hsp10 chaperone system had been shown to be altered (Pedersen et al. 2003), we investigated the HSPD1 and HSPE1 genes in 60 patients with the following ACADS genotypes (28 c.625A/c.625A, 2 c.625A/c.511T; 2 c.511T/c.511T, 24 c.625A/rare; 4 c.511T/rare).
Patients with multiple mitochondrial enzyme deficiency (MMED)
These were two patients described previously (Briones et al. 1997; Agsteribbe et al. 1993) in whom the activities of a series of mitochondrial enzymes were found to be decreased. Both patients died of their disorder at age 2 days and 4 1/2 years respectively. DNA purified from cultured skin fibroblasts was analysed.
Control individuals
Twenty unrelated control individuals were selected from the Danish Cell bank (Eiberg et al. 1983) including the ten individuals reported earlier (Hansen et al. 2003). To obtain a more precise estimate of the frequencies for rare variations detected in the promoter region (g.3172A>G, g.3191dupC, g.3198C>G) and the amino acid replacing variations in the coding region (c.1136A>G/p.Asp379Gly, c.1676G>A/p.Gly559p.Asp, c.1688G>C/p.Gly563Ala), 30 additional, unrelated control individuals from the Danish Cell bank were typed by sequencing the respective regions of the genome. The frequencies for the c.292G>A/p.Val98Ile variations associated with hereditary spastic paraplegia and the c.551A>G (p.Asn184Ser) variation in 400 unrelated control individuals have been determined previously (Hansen et al. 2002). For the polymorphic variation c.1688G>C/p.Gly563Ala an additional 114 unrelated control individuals were typed using a PCR/RFLP assay.
Sequencing
Screening for variations was performed by direct sequencing in both directions (MMED patients) or one direction (SIDS and SCAD/EMA patient groups) of PCR products spanning exons or the promoter region of the chaperonin genes as described previously (Hansen et al. 2003). For ambiguous stretches in the obtained sequences, amplification and sequencing was repeated. Due to limited availability of material from the SCAD/EMA patient group, repeating sequencing of ambiguous regions was not possible in all cases, but high quality sequences covered >95% of the sequenced regions in this patient group. Sequences were aligned using SEQUENCHER software (Version 3.1.1; Gene Codes Corporation, Mich., USA) and the entire chromatograms were reviewed manually. The presence of rare heterozygous variations was confirmed by repeating PCR amplification and direct sequencing of the respective region in the relevant samples.
Functional analysis of promoter variations
The detected promoter variations were introduced by site-directed mutagenesis (QuikChange; Stratagene, La Jolla, Calif., USA) into the two previously described (Hansen et al. 2003) luciferase promoter reporter plasmids containing a 656-bp long fragment of the intergenic region between the HSPD1 and HSPE1 genes in either orientation. The promoter inserts were sequenced to verify the presence of the respective mutations and thereafter subcloned into another batch of the pGL3 (Promega) parent vector to rule out nucleotide variations introduced into the vector region during the mutagenesis procedure. Promoter activity was analysed after co-transfection of the human embryonic kidney cell line HEK-293 (ATCC) with the luciferase reporter constructs and the CMV-ß-galactosidase vector construct. HEK-293 cells were cultured at 37°C with 5% (v/v) CO2 in DMEM (Gibco) supplemented with 5% heat-inactivated FCS, 100 U/ml penicillin, and 0.1 mg/ml streptomycin (Leo, Denmark). Cells were seeded in 12-well plates 1 day prior to transfection. Each well received 75 ng pGL3 vector and 25 ng ß-galactosidase and carrier DNA (empty pcDNA3.1 vector; Invitrogen) to a total amount of 1.5 μg DNA before treatment with FuGENE6 transfection reagent (Roche Diagnostics). Cells were harvested 48 h post transfection in 200 μl lysis buffer (Tropix). Luciferase and ß-galactosidase activity was measured using a MicroLumatPlus LB96V luminometer (Berthold) in combination with the Luc-screen (Applied Biosystems) and Galacto-light plus kits (Tropix), respectively. Correction for transfection efficiency was done by normalising the luciferase activity to the ß-galactosidase activity.
Analysis of missense variations
The non-synonymous variations in the coding region (c.1136A>G/Asp379Gly, c1676G>A/Gly559Asp, and c.1688G>C/Gly563Ala) were introduced, by site-directed mutagenesis (QuikChange or megaprimer PCR-method), into the prokaryotic expression vector that contains cDNA encoding Hsp10 and the processed form of Hsp60 supplemented with a methionine initiation codon as an operon under control of an IPTG-inducible promoter (Hansen et al. 2002). Presence of the respective variations and absence of mutagenesis-induced PCR errors was established by sequencing and subcloning. The resulting vectors were used in the genetic complementation assay described previously (Hansen et al. 2002; Richardson et al. 2001).
Results
Variations in the coding and promoter regions of the HSPD1 and HSPE1 genes
Sequencing of all HSPD1 and HSPE1 exons, exon/intron junctions and the common bidirectional promoter was performed on DNA from the 61 SIDS cases, 60 SCAD/EMA patients and two patients with multiple mitochondrial enzyme deficiency.
We found that the multiple mitochondrial enzyme deficiency patient described by Briones et al. (1997) was heterozygous for the rare g.3507T allele (formerly named IR 453C>T; nomenclature of sequence variations follows the recommendations of the Human Gene Variation Society (http://www.hgvs.org/mutnomen/) and numbering is based on accession number AJ250915) in the promoter region. This variation has been detected previously in a control individual, and functional analysis revealed no effect on promoter activity (Hansen et al. 2003). This patient was homozygous for the frequent variant (g.3175C) of the g.3175C/G polymorphism (formerly described as IR 121C/G) whereas the MMED patient described by Agsteribbe et al. (1993) was homozygous for the less frequent allele (g.3175G) of the g.3175C/G polymorphism (formerly described as IR 121C/G). Both patients were homozygous for the more frequent alleles of the silent variations in exons 2 and 3 (c.69T and c.273A; Fig. 1). Taken together, this defies the notion that mutations in the HSPD1 and HSPE1 promoter, coding regions and exon/intron junctions cause the phenotype.
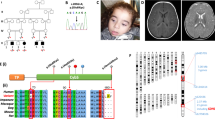
Variations in the coding region of the HSPD1 and HSPE1 chaperonin genes and the bidirectional promoter. The exon and promoter regions are drawn to scale. The position of introns and the direction of transcription of the genes are indicated. Non-synonymous variations in the coding region are indicated above and variations in the promoter and silent variations in the coding region below the drawing. The mutation associated with hereditary spastic paraplegia in a French kindred is boxed with a stippled line. Polymorphic variations are shaded in grey and novel variations detected in the present study are marked with an asterisk. Numbering refers to the genomic and cDNA sequence coordinates from accession number AJ250915
Analysis of the SIDS and SCAD/EMA patients revealed six novel variations (Fig. 1; Tables 1, 2), namely two in the promoter region (g.3172A>G, g.3191dupC) and four in the coding region (one silent: c.27C>G and three amino acid replacing variations: c.1136A>G/p.Asp379Gly, c.1676G>A/p.Gly559Asp, c.1688G>C/p.Gly563Ala; amino acid numbering in this article starts with the initiator methionine; as for Hsp60, the first amino acid of the mature polypeptide corresponding to alanine-27 has commonly been used). No variations were detected in the exon/intron junctions.
With regard to the polymorphic variations, the g.3175C/G variation in the promoter, the c.-15C/T variation in the 5′UTR of the HSPD1 transcript, and the silent c.273G>A variation in the HSPD1 coding region displayed similar frequencies in the patient and control groups (Tables 1, 2). The silent c.69T variant in the coding region was found to be considerably less frequent (8%) in the 20 fully sequenced control individuals as compared to the patient groups (16.4% in the SIDS and 19% in the SCAD/EMA group, respectively). This variation is registered in the dbSNP database (http://www.ncbi.nlm.nih.gov/projects/SNP/; see Table 1) and frequencies of 8 and 22% in a multinational and Japanese population, respectively, have been measured indicating that ethnic differences may be the reason for the discrepancy. The SCAD/EMA patients represent a multinational group whereas the SIDS cohort like the control material is mostly of Danish descent.
With the exception of the polymorphic variation in the coding region (c.1688G>C/p.Gly563Ala), all novel variations detected in this study were only found in one allele of a single patient and absent in the analysed control individuals. The non-synonymous c.551A>G/p.Asn184Ser variation present in one allele of the SIDS cohort was earlier found in a patient with spastic paraplegia, but was absent in affected family members from the same kindred (Hansen et al. 2002). It was also detected in 1 out of 800 alleles from 400 unrelated Danish control individuals, and functional analysis using a bacterial complementation system showed that it had no significant effect.
The overall number of variations detected in our studies in the HSPD1-HSPE1 gene region is 14 (Fig. 1). Three (g.3175C>G, g.3507C>T, c.-15C>T) of the 14 nucleotide variations are localised in hypermutable CpG dinucleotides. Of the 14 variations that we have detected in the present and previous work in altogether 326 chromosomes from patients and control individuals, 5 had previously been reported to the dbSNP database (see Tables 1, 2). Six non-synonymous variations in the HSPD1 and HSPE1 coding region reported in dbSNP have not been observed in any of the 326 chromosomes that we have sequenced. A large number of intron-less HSPD1 and HSPE1 pseudogenes are present in the human genome (see http://www.pseudogene.org and Hansen et al. 2003; Pochon and Mach 1996) and some of the reported variations likely arise from them. It is thus advisable to proceed with caution when using this information, and validate variations in the chaperonin genes by using PCR primers situated in the intron regions.
Based on our sequencing data, the nucleotide diversity in the patient groups is somewhat higher than in the control group (Table 3). In general, the nucleotide diversity in the HSPD1 and HSPE1 coding region in the patient groups and controls is markedly lower than the average gene diversity (1/346) obtained by Cargill and coworkers by screening of 106 potential disease related genes in 114 independent alleles (Cargill et al. 1999). It is likely that the lower diversity in the chaperonin genes reflects their high conservation throughout evolution.
Investigation of the effect of the variations situated in the promoter
The two newly detected variations (g.3172A>G, g.3191dupC) and the previously reported polymorphic g.3175C>G variation in the bidirectional promoter region were analysed using a luciferase reporter assay as described earlier (Hansen et al. 2003). The g.3172A>G variant showed a tendency to slightly increased basal promoter activity in the HSPD1 direction (Fig. 2), whereas the other two variants displayed very similar basal promoter activity in both directions.
Analysis of the effect of the detected variations on activity of the bidirectional promoter. Luciferase reporter plasmids carrying a 656 bp fragment comprising the bidirectional promoter with the indicated promoter variations in both the HSPD1 and HSPE1 orientation were, together with a plasmid carrying a cytomegalo virus promoter driven ß-galactosidase cDNA, used to transfect HEK-293 cells (see Materials and methods for details). Cells were grown at 37°C, harvested after 48 h and luciferase and ß-galactosidase activity were determined. Triplicate transfections were performed with two different DNA preparations of each construct. The quotient of the values obtained from the luciferase and ß-galactosidase measurements normalised to the wild type in the HSPD1 or HSPE1 direction, respectively, are given. As the two data sets had equal variances, the values from both series were combined and common standard deviations (error bars) were calculated. ‘wt’ refers to the sequence carrying the most frequent alleles in all variant positions; the variants carry the indicated variant nucleotide in relation to ‘wt’
Investigation of the effect of the non-synonymous variations in the coding region
The three newly-detected non-synonymous variations in the HSPD1 coding region (p.Asp379Gly, p.Gly559Asp, and p.Gly563Ala) were analysed using the Escherichia coli complementation system used previously (Hansen et al. 2002). In brief, plasmids carrying cDNA encoding the mature part of the respective Hsp60 variant proteins together with the Hsp10 co-chaperonin as an operon were transformed into E. coli B174 cells. The endogenous groESL chaperonin genes were subsequently deleted using phage transduction and homologous recombination. As the bacterial chaperonins are indispensable genes, the ability of the respective expressed Hsp60 variant protein together with wild type Hsp10 to support growth of E. coli cells lacking the endogenous chaperonins was scored as a measure for its functional capacity. All three variants were able to functionally complement the deletion of the E. coli groESL genes suggesting that none of the mutations exerts a major effect on chaperonin function. Furthermore, no apparent differences in growth characteristics at various temperatures could be observed between the transductants carrying plasmids encoding the different Hsp60 variants as compared to transductants with the plasmid encoding wild type Hsp60 (data not shown).
Discussion
Although molecular chaperones have been in focus for more than two decades, and variations in their corresponding genes have repeatedly been suggested as potential modifiers in various disease processes, surprisingly few studies have addressed variations in these genes in humans. Recently, variations in the Hsp90alpha and Hsp90beta genes in 73 Caucasians have been reported (Passarino et al. 2003) and subsequent investigations of their functional effects using a yeast-based assay system revealed that the non-synonymous variant p.Gln488His in the alpha-Hsp90 gene was severely defective (MacLean et al. 2005). Altogether 14 variations in the three highly inducible cytosolic Hsp70 genes HSP70-1, HSP70-2 and HSP70-HOM have been described, and some of them have been shown to be associated with various diseases, such as rheumatoid arthritis, multiple sclerosis, insulin-dependent diabetes mellitus and ageing just to name a few (see Singh et al. 2004 and references therein).
In the present study, we selected a spectrum of diseases where it was indicated that variations in the mitochondrial chaperonin genes could represent disease-associated or disease-modifying factors. In the MMED patients, decreased activities of a series of mitochondrial enzymes were observed. Detailed analysis of one patient showed low levels of Hsp60 protein and a partial deficiency in Hsp60 was suggested as the likely cause (Huckriede et al. 1995; Huckriede and Agsteribbe 1994; Agsteribbe et al. 1993). Our sequencing results of this patient revealed only polymorphic variations in the chaperonin gene coding, intron/exon junction and promoter regions. The patient reported by Briones et al. (1997) displayed heterozygosity for the g.3507C>T variation in the promoter. This variation has been reported previously, and functional analysis of its impact for promoter activity revealed no significant effect (Hansen et al. 2003), arguing against the notion that mutations in the chaperonin genes play a role in these patients.
For the SCAD/EMA patient group, a modifying effect of variations in the chaperonin genes on SCAD enzyme folding was suggested by the observation that the two non-synonymous polymorphic ACADS gene variants c.511T and c.625A, which are strongly overrepresented in the patient group compared to controls (Corydon et al. 1996; Corydon et al. 2001), cause prolonged interaction of their encoded SCAD variant proteins with the Hsp60 chaperone and delayed folding and assembly to the active tetramer (Pedersen et al. 2003). Variations in the chaperonin genes that compromise their function or affect their expression could possibly exacerbate this mild effect and trigger disease. In 60 patients presenting with ethylmalonic aciduria and carrying one of these two rare ACADS variations in at least one allele, we detected two variations in the HSPD1/HSPE1 promoter region (g.3172A>G, g.3191dupC) and two non-synonymous variations in the HSPD1 coding region (c.1136A>G/p.Asp379Gly, c.1676G>A/p.Gly559Asp). These variations were found in one allele of single patients each and could potentially explain the defect in these patients. However, analysis of their potential effects on expression or function revealed no significant impact. Because, in addition, no significant over- or under-representation of the polymorphic variations in the mitochondrial chaperonin genes could be observed, we conclude that variations in the mitochondrial chaperonin genes play no significant role in the SCAD/EMA patient group.
Finally, in the SIDS cohort study, a possible link to the mitochondrial chaperonin genes was suggested by the finding that hyperthermia, which is known to induce HSPD1 and HSPE1 transcription, represents a risk factor (Gozal 1996). In the 61 SIDS cases investigated there was no significant over- or under-representation of polymorphic sequence variations. One patient was heterozygous for the non-synonymous c.551A>G/p.Asn184Ser variation. This variation has been reported earlier, as has the functional analysis of its p.Asn184Ser variant protein in a bacterial complementation assay, which showed no dramatic effect on chaperonin function (Hansen et al. 2002). These results suggest that HSPD1 and HSPE1 gene variations represent no significant factor for the development of SIDS. The MspI RFLP observed with a HSPD1 cDNA probe, and reported to be overrepresented in a SIDS cohort (Rahim et al. 1996), most likely represents a polymorphism that resides in a HSPD1 pseudogene, since the MspI sites in this chromosome region predicted by the complete human genome sequence are not compatible with the chromosomal DNA fragments observed by these authors (data not shown). Furthermore, the statistical significance of the finding has been challenged (Tanner et al. 1997).
The rare variations in the promoter region and the non-synonymous variations in the coding region detected in this report may still exert mild effects contributing to the phenotype in a few patients. Only for the g.3172A variant was a small increase of promoter activity indicated experimentally. The facts that the g.3172A>G variant site, like the g.3175G>C g.3191dupC sites, is situated in a short sequence stretch that is absent in the mouse and rat promoter sequences (Hansen et al. 2003), and that all five so far detected variations in the HSPD1/HSPE1 promoter region are situated distantly from the heat-shock factor binding elements and the CHOP binding motif involved in the heat-shock response and a mitochondria-specific stress-response, respectively (Zhao et al. 2002), argue against a major impact of any of them.
Functional testing of the non-synonymous variations in the coding region using the bacterial complementation assay that is capable of revealing dramatic effects, such as the one exerted by the mutation (c.292G>A; p. Val98Ile) associated with hereditary spastic paraplegia (Hansen et al. 2002), showed no detectable effect, arguing against them having a major impact. Two of the newly detected variations that alter amino acids in Hsp60 are localised in a stretch encoding the Gly-Gly-Met iterations in the carboxy terminus of the protein. Similar sequences are present in the carboxy terminus of many, but not all, type I chaperonins (Brocchieri and Karlin 2000). A four-fold Gly-Gly-Met repeat region is present in E. coli GroEL C-terminus. Its structure is not resolved in the X-ray crystals suggesting a flexible conformation. Deletion of the entire repeat region in GroEL has indicated that it is dispensable, although deletion had some effect on ATPase activity and the ability to suppress temperature-sensitive mutations (Mclennan et al. 1993). Moreover, a recent study showed that replacement of the methionine residues by alanine decelerated folding of a mutant maltose binding protein (Tang et al. 2006).
The variant positions of the p.Asn184Ser and the p.Asp379Gly variations correspond to lysine-160 and glutamic acid-354 in E. coli GroEL, respectively. In the E. coli GroEL structure, both residues are situated at the outer surface of the ring structure. Inspection of alignments shows that the mutated asparagine-184 is not well conserved, but a negatively charged residue at the position corresponding to aspartic acid-379 in human Hsp60 is conserved in the large majority of chaperonins.
In summary, we have described a series of variations in the HSPD1/HSPE1 promoter and coding regions that may exert potential effects on chaperonin expression and function. Although the detected variations do not play a major role in the disease groups investigated, they may still represent modifiers with subtle effects. Although variations in chaperone genes have been used in a large number of association studies, studies on functional tests of the impact of these variations have been reported very rarely. Clearly, functional testing of variations that exert small effects is difficult, and devising appropriate assay systems that can reveal small effects presents a challenge for future research in the field of multifactorial diseases and modifier genes.
References
Agsteribbe E, Huckriede A, Veenhuis M, Ruiters MH, Niezen-Koning KE, Skjeldal OH, Skullerud K, Gupta RS, Hallberg R, van Diggelen OP et al (1993) A fatal, systemic mitochondrial disease with decreased mitochondrial enzyme activities, abnormal ultrastructure of the mitochondria and deficiency of heat shock protein 60. Biochem Biophys Res Commun 193:146–154
Andresen BS, Dobrowolski SF, O'Reilly L, Muenzer J, McCandless SE, Frazier DM, Udvari S, Bross P, Knudsen I, Banas R, Chace DH, Engel P, Naylor EW, Gregersen N (2001) Medium-chain acyl-CoA dehydrogenase (MCAD) mutations identified by MS/MS-based prospective screening of newborns differ from those observed in patients with clinical symptoms: identification and characterization of a new, prevalent mutation that results in mild MCAD deficiency. Am J Hum Genet 68:1408–1418
Bonini NM, Fortini ME (2003) Human neurodegenerative disease modeling using Drosophila. Annu Rev Neurosci 26: 627–656
Botstein D, Risch N (2003) Discovering genotypes underlying human phenotypes: past successes for mendelian disease, future approaches for complex disease. Nat Genet 33(Suppl):228–237
Briones P, Vilaseca MA, Ribes A, Vernet A, Lluch M, Cusi V, Huckriede A, Agsteribbe E (1997) A new case of multiple mitochondrial enzyme deficiencies with decreased amount of heat shock protein 60. J Inherit Metab Dis 20:569–577
Brocchieri L, Karlin S (2000) Conservation among HSP60 sequences in relation to structure, function, and evolution. Protein Sci 9:476–486
Bross P, Andresen BS, Winter V, Kräutle F, Jensen TG, Nandy A, Kølvraa S, Ghisla S, Bolund L, Gregersen N (1993) Co-overexpression of bacterial GroESL chaperonins partly overcomes non-productive folding and tetramer assembly of E. coli-expressed human medium-chain acyl-CoA dehydrogenase (MCAD) carrying the prevalent disease-causing K304E mutation. Biochim Biophys Acta 1182:264–274
Bross P, Jespersen C, Jensen TG, Andresen BS, Kristensen MJ, Winter V, Nandy A, Kräutle F, Ghisla S, Bolund L, Kim JJP, Gregersen N (1995) Effects of two mutations detected in medium chain acyl-CoA dehydrogenase (MCAD)-deficient patients on folding, oligomer assembly, and stability of MCAD enzyme. J Biol Chem 270:10284–10290
Byard RW, Krous HF (2003) Sudden infant death syndrome: overview and update. Pediatr Dev Pathol 6:112–127
Cargill M, Altshuler D, Ireland J, Sklar P, Ardlie K, Patil N, Shaw N, Lane CR, Lim EP, Kalyanaraman N, Nemesh J, Ziaugra L, Friedland L, Rolfe A, Warrington J, Lipshutz R, Daley GQ, Lander ES (1999) Characterization of single-nucleotide polymorphisms in coding regions of human genes. Nat Genet 22:231–238
Corydon MJ, Gregersen N, Lehnert W, Ribes A, Rinaldo P, Kmoch S, Christensen E, Kristensen TJ, Andresen BS, Bross P, Winter V, Martinez G, Neve S, Jensen TG, Bolund L, Kølvraa S (1996) Ethylmalonic aciduria is associated with an amino acid variant of short chain acyl-coenzyme A dehydrogenase. Pediatr Res 39:1059–1066
Corydon MJ, Vockley J, Rinaldo P, Rhead WJ, Kjeldsen M, Winter V, Riggs C, Babovic-Vuksanovic D, Smeitink J, De Jong J, Levy H, Clive SA, Roe C, Matern D, Dasouki M, Gregersen N (2001) Role of common gene variations in the molecular pathogenesis of short- chain acyl-CoA dehydrogenase deficiency. Pediatr Res 49:18–23
Corydon TJ, Hansen J, Bross P, Jensen TG (2005) Down-regulation of Hsp60 expression by RNAi impairs folding of medium-chain acyl-CoA dehydogenase wild-type and disease-associated proteins. Mol Genet Metab 85:260–270
Dipple KM, McCabe ER (2000) Phenotypes of patients with “Simple” Mendelian disorders are complex traits: thresholds, modifiers, and systems dynamics. Am J Hum Genet 66:1729–1735
Dubaquie Y, Looser R, Fünfschilling U, Jenö P, Rospert S (1998) Identification of in vivo substrates of the yeast mitochondrial chaperonins reveals overlapping but non-identical requirement for hsp60 and hsp10. EMBO J 17:5868–5876
Eiberg H, Mohr J, Nielsen LS, Simonsen N (1983) Genetics and linkage relationships of the C3 polymorphism: discovery of C3-Se linkage and assignment of LES-C3-DM-Se-PEPD-Lu synteny to chromosome 19. Clin Genet 24:159–170
Fares MA, Ruiz-Gonzalez MX, Moya A, Elena SF, Barrio E (2002) Endosymbiotic bacteria: GroEL buffers against deleterious mutations. Nature 417:398
Fernandez-Funez P, Nino-Rosales ML, de Gouyon B, She WC, Luchak JM, Martinez P, Turiegano E, Benito J, Capovilla M, Skinner PJ, McCall A, Canal I, Orr HT, Zoghbi HY, Botas J (2000) Identification of genes that modify ataxin-1-induced neurodegeneration. Nature 408:101–106
Gozal D (1996) Deficient heat shock protein expression: a potential mechanism for the sudden infant death syndrome. Med Hypotheses 46:52–54
Hansen JJ, Dürr A, Cournu-Rebeix I, Georgopoulos C, Ang D, Nielsen MN, Davoine CS, Brice A, Fontaine B, Gregersen N, Bross P (2002) Hereditary spastic paraplegia SPG13 is associated with a mutation in the gene encoding the mitochondrial chaperonin Hsp60. Am J Hum Genet 70:1328–1332
Hansen JJ, Bross P, Westergaard M, Nielsen MN, Eiberg H, Børglum AD, Mogensen J, Kristiansen K, Bolund L, Gregersen N (2003) The genomic structure of the human mitochondrial chaperonin genes: HSP60 and HSP10 are localised head to head on chromosome 2 separated by a bidirectional promoter. Hum Genet 112:71–77
Huckriede A, Agsteribbe E (1994) Decreased synthesis and inefficient mitochondrial import of hsp60 in a patient with a mitochondrial encephalomyopathy. Bba-Mol Basis Dis 1227:200–206
Huckriede A, Heikema A, Sjollema K, Briones P, Agsteribbe E (1995) Morphology of the mitochondria in heat shock protein 60 deficient fibroblasts from mitochondrial myopathy patients. Effects of stress conditions. Virchows Archiv 427:159–165
Lundemose JB, Kolvraa S, Gregersen N, Christensen E, Gregersen M (1997) Fatty acid oxidation disorders as primary cause of sudden and unexpected death in infants and young children: an investigation performed on cultured fibroblasts from 79 children who died aged between 0–4 years. Mol Pathol 50:212–217
MacLean MJ, Llordella MM, Bot N, Picard D (2005) A yeast-based assay reveals a functional defect of the Q488H polymorphism in human Hsp90alpha. Biochem Biophys Res Commun 337:133–137
Mclennan NF, Girshovich AS, Lissin NM, Charters Y, Masters M (1993) The strongly conserved carboxyl-terminus glycine methionine motif of the Escherichia coli GroEL chaperonin is dispensable. Mol Microbiol 7:49–58
Nadeau JH (2001) Modifier genes in mice and humans. Nat Rev Genet 2:165–174
Nollen EA, Garcia SM, van HG, Kim S, Chavez A, Morimoto RI, Plasterk RH (2004) Genome-wide RNA interference screen identifies previously undescribed regulators of polyglutamine aggregation. Proc Natl Acad Sci USA 101:6403–6408
Passarino G, Cavalleri GL, Stecconi R, Franceschi C, Altomare K, Dato S, Greco V, Cavalli Sforza LL, Underhill PA, De Benedictis G (2003) Molecular variation of human HSP90alpha and HSP90beta genes in Caucasians. Hum Mutat 21:554–555
Pedersen CB, Bross P, Winter VS, Corydon TJ, Bolund L, Bartlett K, Vockley J, Gregersen N (2003) Misfolding, degradation, and aggregation of variant proteins: the molecular pathogenesis of short chain acyl-CoA dehydrogenase (SCAD) deficiency. J Biol Chem 278:47449–47458
Perezgasga L, Segovia L, Zurita M (1999) Molecular characterization of the 5' control region and of two lethal alleles affecting the hsp60 gene in Drosophila melanogaster. FEBS Lett 456:269–273
Pochon NAM, Mach B (1996) Genetic complexity of the human hsp 60 gene. Int Immunol 8:221–230
Queitsch C, Sangster TA, Lindquist S (2002) Hsp90 as a capacitor of phenotypic variation. Nature 417:618–624
Rahim RA, Boyd PA, Ainslie Patrick WJ, Burdon RH (1996) Human heat shock protein gene polymorphisms and sudden infant death syndrome. Arch Dis Child 75:451–452
Richardson A, Schwager F, Landry SJ, Georgopoulos C (2001) The importance of a mobile loop in regulating chaperonin/ co-chaperonin interaction: humans versus Escherichia coli. J Biol Chem 276:4981–4987
Rutherford SL, Lindquist S (1998) Hsp90 as a capacitor for morphological evolution. Nature 396:336–342
Saijo T, Welch WJ, Tanaka K (1994) Intramitochondrial folding and assembly of medium-chain acyl-CoA dehydrogenase (MCAD)—demonstration of impaired transfer of K304E-variant MCAD from its complex with Hsp60 to the native tetramer. J Biol Chem 269:4401–4408
Singh R, Kolvraa S, Bross P, Gregersen N, Andersen NB, Frederiksen H, Christensen K, Rattan SI (2004) Association between low self-rated health and heterozygosity for -110a > c polymorphism in the promoter region of HSP70-1 in aged Danish twins. Biogerontology 5:169–176
Slavotinek AM, Biesecker LG (2001) Unfolding the role of chaperones and chaperonins in human disease. Trends Genet 17:528–535
Tang YC, Chang HC, Roeben A, Wischnewski D, Wischnewski N, Kerner MJ, Hartl FU, Hayer-Hartl M (2006) Structural features of the GroEL-GroES Nano-cage required for rapid folding of encapsulated protein. Cell 125:903–914
Tanner MS, Sharrard MJ, Rigby AS (1997) Gene polymorphisms and the use of the bonferroni correction factor: when and when not to apply? Arch Dis Child 76:386
Zhao Q, Wang J, Levichkin IV, Stasinopoulos S, Ryan MT, Hoogenraad NJ (2002) A mitochondrial specific stress response in mammalian cells. EMBO J 21:4411–4419
Acknowledgments
This work was supported by grants from the Karen Elise Jensen Foundation, the Danish Medical Research Council, the Danish Centre for Human Genome Research and the Institute of Clinical Medicine, Aarhus University.
Author information
Authors and Affiliations
Corresponding author
Additional information
We dedicate this article to the memory of our colleague Etienne Agsteribbe.
Rights and permissions
About this article
Cite this article
Bross, P., Li, Z., Hansen, J. et al. Single-nucleotide variations in the genes encoding the mitochondrial Hsp60/Hsp10 chaperone system and their disease-causing potential. J Hum Genet 52, 56–65 (2007). https://doi.org/10.1007/s10038-006-0080-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10038-006-0080-7
Keywords
This article is cited by
-
Hitzeschockproteine als forensisch und klinisch bedeutende Stressmarker
Rechtsmedizin (2019)
-
Comparative proteome analysis for identification of differentially abundant proteins in SIDS
International Journal of Legal Medicine (2017)
-
Functional single-nucleotide variant of HSPD1 in sudden infant death syndrome
Pediatric Research (2013)
-
Comparison of the longissimus muscle proteome between obese and lean pigs at 180 days
Mammalian Genome (2013)
-
Gene variants predisposing to SIDS: current knowledge
Forensic Science, Medicine, and Pathology (2011)