Abstract
Knowledge of the spatial scales of diversity is necessary to evaluate the mechanisms driving biodiversity and biogeography in the vast but poorly understood deep sea. The community structure of kinetoplastids, an important group of microbial eukaryotes belonging to the Euglenozoa, from all abyssal plains of the South Atlantic and two areas of the eastern Mediterranean was studied using partial small subunit ribosomal DNA gene clone libraries. A total of 1364 clones from 10 different regions were retrieved. The analysis revealed statistically not distinguishable communities from both the South-East Atlantic (Angola and Guinea Basin) and the South-West Atlantic (Angola and Brazil Basin) at spatial scales of 1000–3000 km, whereas all other communities were significantly differentiated from one another. It seems likely that multiple processes operate at the same time to shape communities of deep-sea kinetoplastids. Nevertheless, constant and homogenous environmental conditions over large spatial scales at abyssal depths, together with high dispersal capabilities of microbial eukaryotes, maintain best the results of statistically indistinguishable communities at larger spatial scales.
Similar content being viewed by others
Introduction
The vast abyssal sea floor covers more than 50% of Earth’s surface and is postulated to have a key role in ecological and biogeochemical processes on a global scale as well as being an untapped reservoir of high genetic and metabolic diversity (Danovaro et al., 2008a). Whereas deep-sea ecosystems are of ‘paramount importance’ (Arístegui et al., 2009) for global oceanic material cycling, only a tiny fraction of the total proportion of the deep sea has been studied up to now (<1%). Most of these studies have focused on vent, ridge or seep ecosystems and not on the vast expanses of the abyssal sea floor, although its extent is orders of magnitude greater (Rex and Etter, 2010).
The local diversity of some deep-sea ecosystems has been well documented, but little is known about deep-sea diversity at greater spatial scales. Danovaro et al. (2008a) reported a clear link between benthic diversity and ecosystem functioning. They estimated that a biodiversity loss of 20–30% could result in a 50–80% reduction of deep-sea ecosystems’ key processes (Danovaro et al., 2008b). Knowledge of the spatial dimension of diversity and hence the need for large-scale community sequencing is necessary to evaluate the mechanisms driving biodiversity and biogeography in this vast but poorly understood ecosystem (Azam and Worden, 2004).
Microbes are important in the function of all ecosystems (Fuhrman, 2009). They dominate the ocean in terms of abundance and metabolic activity (Azam and Malfatti, 2007), and have a disproportional large role in abyssal energy flow (Rex et al., 2006). Among them, microbial eukaryotes are key players in controlling prokaryotic abundance and especially heterotrophic nanoflagellates seem to control efficiently prokaryotic communities in the deep ocean, at least in the mesopelagic zone (Fukuda et al., 2007; Arístegui et al., 2009). The few molecular studies focusing on the diversity of abyssal microbial eukaryotes rather point to specific communities with only minor contributions from epipelagic depths. An important fraction of these communities is comprised of Euglenozoa, a large group of flagellate protozoa characterized by the ultrastructure of their flagella containing a rod (called paraxonemal; Hausmann et al., 2003). This group seems to be more abundant in deep-sea clone libraries than in epipelagic waters (Countway et al., 2007) and has been reported from a wide variety of deep-sea environments (for example, López-García et al., 2001; Stoeck et al., 2003; Scheckenbach et al., 2010).
Kinetoplastids are an ecological important (Moreira et al., 2004) group of euglenozoans. They are characterized by the presence of a DNA-containing granule located within the single mitochondrion associated with the base of the cell’s flagella (the kinetoplast; Hausmann et al., 2003). Kinetoplastids are found in almost all terrestrial and aquatic environments. A recent study of deep-sea euglenozoans revealed the presence of deep-sea specific lineages of diplonemids, a sister group of kinetoplastids (Lara et al., 2008), and studies of deep-sea kinetoplastids reported a disproportional low representation of these deep-sea lineages in public databases (Shah Salani, 2009; Scheckenbach et al., 2010). The ecological relevance of kinetoplastids, together with the potential specific deep water lineages reported, made this group an interesting subject for the study of deep-sea microbial eukaryotes.
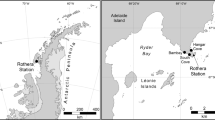
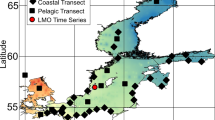
The Mediterranean is separated from the Atlantic by the Gibraltar sill and represents one of the few warm deep-sea regions on Earth, with deep-water temperatures of about 13.5 °C, whereas abyssal bottom water in the Atlantic usually has a temperature of 2–4 °C, which makes it an interesting location for studying deep-sea biogeography. Here, we have studied the community structure and spatial scales of kinetoplastids by rDNA clone libraries from all abyssal plains of the South Atlantic (Angola-, Argentine-, Brazil-, Guinea-, Namibia- and Pernambuco Abyssal Plain; Figure 1), as well as the Ierapetra Basin and the Pliny Trench of the eastern Mediterranean.
Materials and methods
Sampling
Samples were taken using multi-corers from the South-East Atlantic (R/V Meteor cruise 63/2, expedition DIVA2;26 February 2005–31 March 2005; Angola Abyssal Plain, 9° 56′ S, 0° 54′ E; Guinea Abyssal Plain, 0° 0′ S, 2° 25′ W; and Namibia Abyssal Plain, 28° 07′ S, 7° 21′ E; Figure 1), from depths of 5033–5142 m; from the Mediterranean (R/V Meteor cruise 71/2;28 December 2006–15 January 2007), from depths of 4400 m (Ierapetra Basin, 34° 30′ N, 26° 11′ E) and 2670 m (Pliny Trench, 33° 44′ N, 26° 08′ E); and from the South-West Atlantic (R/V Meteor cruise 79/1, expedition DIVA3;10 July 2009–23 August 2009; Argentine, 35° 59′ S, 49° 01′ W; South Brazil Basin, 26° 34′ S, 35° 13′ W; North Brazil Basin, 14° 59′ S, 29° 57′ W; and Pernambuco Abyssal Plain, 3° 57′ S, 28° 05′ W; Figure 1), from depths of 4605–5189 m. Additionally, samples were also taken from the Great Meteor Seamount (29° 57′ N, 28° 35′ W) from depths of 300 m using a box-corer. An overview of the sampling regions is provided in Supplementary Table 1.
From each region of the South-East Atlantic, 10 l of sediment-overlaying water were immediately filtered over 0.2-μm polycarbonate filters (Millipore, Schwalbach, Germany) and stored at −20 °C until further treatment. The following expeditions to the Mediterranean and South-West Atlantic did not enable us to equally sample sediment-overlaying water but solely sediment. Therefore, from the Mediterranean, the upper 2 mm of sediment were filled into 50-ml DNA/RNA and nuclease-free tubes (Sarstedt, Nuembrecht, Germany) together with sediment-overlaying water and stored at −80 °C until further treatment. Owing to the conditions on board of the research vessel during the Mediterranean cruise, a large proportion of the sediment was already suspended and could therefore not be sampled without overlaying water. From the South-West Atlantic and Seamount samples, sediment was filled into 15-ml DNA/RNA and nuclease-free tubes (Sarstedt) and equally stored at −80 °C until further treatment. In addition, 224 ml of sediment-overlaying water from the Brazil Basin were filled in 15-ml DNA/RNA and nuclease-free tubes (Sarstedt) and stored at −80 °C until further treatment.
Genomic DNA extraction, purification, cloning and sequencing
Genomic DNA from filters of the South-East Atlantic was extracted using a phenol/chloroform/CTAB extraction protocol (Clark, 1992) and further purified by using sepharose-4B/PVPP columns (Edel-Hermann et al., 2004). In order to process the larger sampling volume of the Mediterranean, we have chosen to separate the Mediterranean samples into sediment (42 g, Ierapetra Basin; 24 g, Pliny Trench) and water (330 ml, Ierapetra Basin; 198 ml, Pliny Trench) by centrifugation (4000 g at 4 °C for 10 min), and extract genomic DNA from each phase using appropriate methods. The water was filtered over 0.2-μm polycarbonate filters (Millipore). Genomic DNA from these filters was extracted by the same procedure as for the South-East Atlantic samples. The sediment was extracted using the Ultra Clean Soil DNA Isolation kit (MO BIO Laboratories, Carlsbad, CA, USA) as described in the manufacturer’s instructions. As the Mediterranean samples already contained a mixture of sediment and sediment-overlaying water as described above, we have pooled the genomic DNA of both phases after extraction. From the sediment samples of the South-West Atlantic, 2 g were used for DNA isolation using the Ultra Clean Soil DNA Isolation kit (MO BIO Laboratories) following the manufacturer’s instructions. The 224 ml of sediment-overlaying water from the Brazil Basin were equally filtered over 0.2-μm polycarbonate filters (Millipore) and genomic DNA from these filters was extracted by the same procedure as for the South-East Atlantic samples.
The purified DNA was pooled and then used for PCR. The partial 18S rDNA gene (position 668–1196/1231 relative to Bodo saltans, GenBank accession numbers DQ207570, JN629095 and JN630458) was amplified using two primer combinations, For_668 (5′-GCTGTTAAAGGGTTCGTAGTTG-3′) and Rev1_1231 (5′-GGACTACAATGGTMTCTAATCATC-3′), as well as For_668 and Rev2_1196 (5′-CACTTTRGTTCTTGATTGAKGAAGG-3′). Primers were designed manually using the entire kinetoplastid data set of the SILVA database (July 2009; Pruesse et al., 2007) following general rules of primer design in conjunction with the NCBI primer-BLAST tool (http://www.ncbi.nlm.nih.gov/tools/primer-blast/). Primers were tested manually for specificity against several other taxonomic groups retrieved from the SILVA database as well as automatically using BLAST. Primers are kinetoplastid-specific and cover all kinetoplastid sequences present in the SILVA database as of July 2009. PCRs were performed in 50 μl of reaction volume under standard conditions as specified by the manufacturer (Ta=50 °C for Rev1_1231; Ta=51 °C for Rev2_1196). Amplicons were pooled and purified using a PeqGOLD Cycle-Pure Kit (Peqlab, Erlangen, Germany) according to the manufacturer’s instructions. Purified 18S rDNA fragments were cloned into pSC-A cloning vectors and transformed into competent cells using the StrataClone PCR Cloning Kit (Stratagene, Santa Clara, CA, USA). Cloned inserts were amplified by PCR using Dream Taq DNA polymerase (Fermentas, St Leon-Rot, Germany) and sequenced in both directions using the BigDye Terminator v3.1 Cycle Sequencing Kit (Applied Biosystems, Foster City, CA, USA) using the vector primers M13For and M13Rev following the manufacturer’s instructions.
Phylogenetic and statistical analysis
Sequences were assembled and corrected manually. Each sequence was checked for chimeras using the various methods available by Mothur v.1.12.0 (Schloss et al., 2009).
For phylogenetic analysis, multiple alignments were computed using the E-INS-i iterative refinement method of MAFFT v.6.833 (Katoh and Toh, 2010). Maximum likelihood phylogenetic trees were computed from these multiple alignments using RAxML v.7.2.7 with the GTR+Ã model of nucleotide substitutions (Stamatakis, 2006) and the extended majority rule bootstrap convergence criteria. Phylogenetic trees were displayed and midpoint rooting was performed using iTOL (Letunic and Bork, 2007).
For statistical analysis, a p-distance matrix from pairwise p-distances was calculated from pairwise alignments for each possible combination using water and distmat from the EMBOSS package v.6.3.1 (Rice et al., 2000). From the p-distance matrix, a minimum evolution phylogenetic tree was computed using FastME v.2.0.7 (Desper and Gascuel, 2002) with SPR post-processing enabled.
Operational taxonomic unit (OTU)-based analysis were performed using Mothur v.1.12.0 (Schloss et al., 2009). This included clustering of all clones into OTUs, calculation of rarefaction curves and richness estimators (Abundance-based Coverage Estimator (ACE); Chao and Lee, 1992), OTU-based analysis of community structure (θYC; Yue and Clayton, 2005), as well as phylogeny-based weighted UniFrac test (Lozupone and Knight, 2005). The threshold for OTU delineation was estimated to be 1% genetic p-distance (Supplementary Figure 1). The estimation was based on the calculation of the median (Q.5) intraspecific p-distance of 44 morphospecies of kinetoplastids with 459 sequences covering the small subunit rDNA (SSU rDNA) position 500–1300 and being longer than 500 bp, retrieved in January 2010 from the SILVA database (Pruesse et al., 2007). Faith’s phylogenetic diversity (Faith, 1992), mean pairwise distance and mean nearest taxon distance (Webb et al., 2002), and the correlation between species co-occurrence and phylogenetic distance (Cavender-Bares et al., 2004) were calculated using R v.2.12.0 (R Development Core Team, 2010), using the R package picante v.1.2-0 (Kembel et al., 2010). The metric for species co-occurrence used was Jaccard’s index of co-occurrence. For all calculations of null models, the independent swap algorithm was used (Gotelli and Entsminger, 2003). Analysis of molecular variances (Excoiffier et al., 1992) was performed using the R package pegas v.0.3-2 (http://ape.mpl.ird.fr/pegas/). Linear regression was also calculated with R, as was the Mantel test using the R package ade4 (Thioulouse et al., 1997).
Results
Sampling efficiency and diversity estimation
We retrieved a total of 1364 clones with an average length of 526 bp. In addition, 37 clones were identified as chimeric sequences, using the various methods for chimera checking available by Mothur (Schloss et al., 2009). All clones were grouped into OTUs, following a value for OTU delineation of 1% median p-distance, resulting in 317 different OTUs. Rank-abundance curves calculated for each sampling area showed the typical rank-abundance curve usually found by phylogenetic surveys of microbes, with, here, 8% of all OTUs containing 50% of all clones. A high percentage of singletons was found (Table 1). The highest percentage of singletons was found for the Pernambuco Abyssal Plain (69%) and the Angola Abyssal Plain (77%). Rarefaction curves leveled-off for all sampling stations except for the Angola and the Pernambuco Abyssal Plain (data not shown). OUT-based richness estimators (SACE; Table 1) were 4–6 times higher for the Pernambuco and the Angola Abyssal Plain, and about 1.5–2.5 times higher for all other sampling regions. Standardized effect sizes versus null communities of Faith’s phylogenetic diversity showed that the Angola Abyssal Plain and the Ierapetra Basin, and to a lesser extent the Pernambuco Abyssal Plain, had relatively higher phylogenetic diversity than the other sampling regions (Supplementary Figure 2). The mean genetic p-distance between clones and their first BLAST hit was 8% (range=38%) and higher than reported by another environmental study of kinetoplastids using specific primers (n=84, mean=2%, range=16%, position 668–1231; Von der Heyden and Cavalier-Smith, 2005).
Community composition and comparison
The taxonomic community composition was heterogeneous between different sampling regions (Figure 2 and Supplementary Figure 3). The most represented taxa were members of Rhynchomonas, Ichthyobodo and Neobodo (Supplementary Figure 4). Only few other genera were retrieved, all also belonging to the Prokinetoplastina or the order Neobodonida. Whereas most communities were composed of a diverse assemblage of the above named taxa, the community from the Seamount was mainly composed of a single taxa, Dimastigella. No members of the orders Eubodonida and Parabodonida, as well as Tryposomatida, were retrieved, although we were able to successfully amplify several cultured isolates of Eubodonida and Parabodonida using both primer pairs.
Scaled bubble plot showing the relative number of clones for the most abundant OTUs (>4 clones; center of the figure) and for each sampling area (bottom of the figure) arranged according to their taxonomic affiliation as determined by BLAST (right-hand side). The bubble plot was scaled in order to allow a better comparison between different sampling regions and, therefore, the size of the bubbles does not represent absolute numbers. Above, a cladogram shows the relationship between the different sampling regions, as given at the bottom of the figure, based on mean nearest neighbor distances (MNND), with dashed branches for statistically indistinguishable sampling regions (weighted UniFrac test, P>0.05). On the left-hand side, a midpoint rooted maximum likelihood GTR+Ã phylogenetic tree, built using one representative sequence for each OTU of the bubble plot (center of the figure), shows the phylogenetic relationship between the OTUs, with bootstrap support >80 shown as black points. Scale bars are given for both clado/phylograms.
Clear differences in the dominance patterns between the sampling areas are visible (Figure 2; can be deduced from Supplementary Figure 3). The dominant OTUs in the South-West Atlantic mostly belonged to Rhynchomonas and Neobodo designis II, which did not dominate communities from both the South-East Atlantic and the Mediterranean. On the other hand, Prokinetoplastea (Ichthyobodo and Perkinsiella) and N. designis/celer dominated the South-East Atlantic and the Mediterranean, and were mostly rare in the South-West Atlantic.
Using a threshold of 1% p-distance for OTU delineation, the number of OTUs present in two or more sampling regions, that is, OTUs shared between regions, was low. A single OTU was found in 8 of the 10 sampling regions and, consequently, no OTU was present in all sampling regions. Sixty-eight percent of all OTUs were present in one and only 18% in more than two sampling areas. Plotting the number of OTUs against the number of shared sampling regions for a given OTU resulted in a unimodal distribution with a long right-hand tail, as commonly reported for microbial rank-abundance curves. By contrast, plotting the number of shared sampling regions for a given OTU against the number of clones within all OTUs of the respective rank resulted in a bimodal distribution skewed to both sides (Supplementary Figure 5), indicating that abundant OTUs were rather present in several sampling regions. This was in particular apparent for two groups of sampling regions: All three sampling regions from the South-West Atlantic (Argentine and Brazil Basin) and the sampling regions with the highest numbers of clones (Ierapetra Basin, Namibia Abyssal Plain, Pliny Trench). These sampling regions clustered together using OTU-based measures of dissimilarity between the structures of two communities (θYC; Supplementary Figure 4). A cladogram of mean nearest neighbor distances based on cophenetic distances between the sampling regions showed a similar clustering and, in addition, a high similarity between the Angola and the Guinea Abyssal Plain (Figure 2), whose OTUs were more restricted in their distribution (Supplementary Figure 5).
Raising the level for OTU delineation to 2 and 3% p-distance, as commonly found in the literature, did lower the total number of OTUs to 214 (2% p-distance) and 177 (3% p-distance). By contrast, the number of clones present in OTUs found in several sampling regions increased and, hence, the distribution of the number of shared sampling regions for a given OTU against the number of clones within all OTUs of the respective rank shifted toward the right side of the plot (Supplementary Figure 5). However, the maximum number of sampling regions an OTU was present was only marginally raised to nine sampling regions. Yue and Clayton’s measure of dissimilarity resulted in similar sampling regions becoming more similar by increasing the threshold for OTU delineation up to 3% p-distance and, at the same time, dissimilarities became more apparent. However, Yue and Clayton’s θ similarity coefficients within the group encompassing the Mediterranean sampling regions and the Namibia Abyssal Plain did not change significantly: èYC for a threshold of 1% p-distance was in the range of 0.40–0.53, and for a threshold of 3% p-distance in the range of 0.43–0.52.
The differences between the sampling regions, as shown above, were supported by analysis of molecular variances. This test demonstrated that there is a strong biogeographical pattern (P<0.001). Accordingly, Holm’s corrected pairwise analysis of molecular variances (Supplementary Table 2) supported the high similarity of the sampling regions of the South-West Atlantic (Argentine and Brazil Basin; P>0.05) as well as the high similarity of the Angola and Guinea Abyssal Plain (P>0.05). Pairwise weighted UniFrac analysis equally supported these findings (Figure 2 and Supplementary Table 2). Analyses of molecular variances grouped according to habitat, depth, temperature or sampling period were also all significant (P<0.001).
Phylogenetic structure
The maximum likelihood phylogenetic tree of all clones showed phylogenetic structuring of most sampling stations (Supplementary Figure 3). Only clones from the Ierapetra Basin and the Pernambuco Abyssal Plain, as well as to a lesser extent the Pliny Trench, were more evenly spread over the phylogenetic tree. This observed phylogenetic structure is supported by standardized effect sizes of mean pairwise distances versus null communities indicating a tree-wide phylogenetic evenness of these sampling regions (Figure 3). The other sampling regions, on the other hand, showed phylogenetic clustering. Standardized effect sizes of mean nearest taxon distances versus null communities showed that the clones from the Angola Abyssal Plain, and to a lesser extent from the Ierapetra Basin, were more evenly spread at the tips of the phylogenetic tree (Figure 3). The other communities showed phylogenetic clustering closer to the tips of the phylogeny. This result was corroborated by a significant negative correlation between species co-occurrence and phylogenetic distance (corr=−0.053, P<0.001).
Relationship between genetic and geographic distances
The mean percentage of identical (0.00% p-distance) clones within each sampling area in relation to the total number of clones of the according area was 58%, whereas the average percentage of shared identical clones between two sampling regions in relation to the total number of clones within both environments was relatively low (18%). There was, however, no significant linear relationship between the percentage of shared identical clones between two sampling regions in relation to the total number of clones within both environments and geographic distance (r=−0.05, P>0.05). By contrast, there was a significant and positive, although weak, linear relationship between genetic and geographic distance (r=0.07, P<0.001). This significant correlation between genetic and geographic distance was supported by a Mantel test (P<0.001).
Discussion
The ocean is an interconnected geophysical fluid principally allowing extensive gene flow at large geographic scales of organisms with a presumed high dispersal potential, such as microorganisms. It may, therefore, not be surprising to find coherent communities of microbial eukaryotes at a given depth horizon at adjacent regions. However, similar communities (P>0.05) were not found in the range of kilometers or tens of kilometers but in the range of hundreds and thousands of kilometers, as reported here for the South-West Atlantic (Argentine and Brazil Basin) and the South-East Atlantic (Angola and Guinea Basin). This may underline the constant and homogeneous conditions prevailing over large spatial distances at abyssal depths. Two previous studies already reported of similar, and even statistically not distinguishable, communities of microbial eukaryotes at larger scales from the abyssal North-West Atlantic (Countway et al., 2007) and the abyssal South-East Atlantic (Scheckenbach et al., 2010). The latter study equally reported of statistically not distinguishable communities from the Angola and the Guinea Basin, and also of statistically distinct communities from the Namibia Abyssal Plain. Our results from the South-East Atlantic, thus, corroborate the report of Scheckenbach et al. (2010) and are also in agreement with a comparable study of communities of prokaryotes from the same sampling regions also reporting of statistically indistinguishable communities from both the Angola and the Guinea Basin, as well as the Guinea Basin and the Namibia Abyssal Plain (Schauer et al., 2009). Furthermore, as the kinetoplastid-specific clone libraries from the South-East Atlantic, reported here, show the same pattern as clone libraries built with general eukaryotic primers as reported by Scheckenbach et al. (2010), it is admissible to assume that the pattern reported here for both the South-East and South-West Atlantic may be similar for other groups of microbial eukaryotes.
The assumption of McClain and Hardy (2010), that geographical ranges may increase with depth as environmental conditions become more constant and homogeneous, may therefore be plausible. Moreover, as many genetically studied microbial eukaryotes from the deep sea have revealed wide distribution ranges (Pawlowski et al., 2007; Lecroq et al., 2009). In their extensive review on deep-sea biogeography, these authors furthermore noticed that many taxa appear widely distributed across the deep sea, even at specialized environments such as hydrothermal vents (McClain and Hardy, 2010). The fact that we had no linear relationship among the number of identical clones (0.00% p-distance) between two sampling regions in relation to the total number of clones at both sampling regions and the geographical distance between both regions (r=−0.05, P>0.05), may point into that direction. Moreover, most dominant OTUs were equally present in both Mediterranean sampling areas and the Cape Basin (Figure 2; Shah Salani, 2009), supporting the hypothesis that some marine protists are indeed cosmopolitan despite geographic barriers and different ecological parameters. Kouridaki et al. (2010) even reported of communities of prokaryotes from the same depth layer (4000 m) of the North-East Pacific and the Eastern Mediterranean having different trophic states and temperature conditions, but which did not show substantial differences in community composition. Wide dispersal ranges, however, of microbes may be a general characteristic of, at least, marine ecosystems. Cermeño and Falkowski (2009) reported that marine diatoms were not limited by dispersal. Wide dispersal ranges may, to some extent, explain the large areas covered by coherent communities and the finding that a particular percentage of identical clones is shared among even geographically distantly related regions, but not the substantial differences among most sampling regions studied.
A central tenet of community ecology is that strong environmental gradients shape ecosystems by controlling the spatial and temporal distribution of species. In this regard, recent evidence suggests a link between marine microbial community structure and water masses (Agogué et al., 2008; Varela et al., 2008; Galand et al., 2009; Kirchman et al., 2010). Galand et al. (2010) reported of a strong association between the large-scale distribution of microbial communities from the deep Arctic and the hydro-geography of the arctic water masses. Differences in community structure would thus reflect the differences in environmental parameters among water masses, as also suggested by studies of marine foraminiferans (De Vargas et al., 1999; Darling et al., 2000). Schauer et al. (2009), who studied the same sampling regions of the South-East Atlantic, attributed the differences reported, at least partially, to environmental differences of the different water masses. The distinction among some communities studied may, therefore, be attributed to environmental differences of the water masses, and, vice versa, the similarity of the communities from both the South-West Atlantic and the South-East Atlantic may consequently be attributed to similar water masses. This accounts for both the bottom water and the surface water as there is a strong pelagic–benthic coupling in marine environments. The organic matter flux down to the deep sea influences deep-sea biodiversity over large spatial scales (Levin et al., 2001; Smith et al., 2008). The phylogenetic clustering reported here provides further evidence for environmental filtering, as phylogenetic clustering at larger spatial scales of organisms with high dispersal potential is most likely the result of environmental filtering (Webb et al., 2002; Kraft et al., 2007).
The Angola and Guinea Basin are filled with North Atlantic Deep Water originating from the Arctic, whereas the Namibia Abyssal Plain and the abyssal plains of the South-West Atlantic are filled with Antarctic Bottom Water originating from the Antarctic. At the surface, the Angola, the Brazil and the Guinea Basin, as well as the Pernambuco Abyssal Plain, are all under the influence of the South Equatorial Current system. The Namibia Abyssal Plain, by contrast, is under the influence of the northward directed Benguela Current, and the North Argentine Basin is at the intersection of the southward directed Brazil Current, which is part of the South Equatorial Current system, and the northward directed Malvinas Current. The different water masses of the South Atlantic may, therefore, be responsible for the biogeographical pattern reported here. They could explain the similarity of the communities from the Angola and the Guinea Basin, as well as their distinction from those of the Namibia Abyssal Plain, and could equally explain the similarity of the North Argentine Basin and the Brazil Basin. The divergence of both Mediterranean communities with those from the Atlantic could also be explained by differences in the water masses, as the Mediterranean is one of the few warm deep-sea regions of the Earth with deep-water temperatures of about 13.5 °C, whereas abyssal bottom water usually has a temperature of 2–4 °C. The observed biogeographical pattern could, therefore, be the result of the environmental heterogeneity represented by water masses with distinct abiotic and biotic properties. The high similarity of the Namibia Abyssal Plain and both Mediterranean sampling areas, on the other hand, is mainly the result of few shared dominant OTUs, and not the result of similarity in species richness (Shah Salani, 2009). In fact, the Mediterranean communities are far diverse than the relatively species-poor Namibia Abyssal Plain (Supplementary Figure 1) and share only those few dominant OTUs with the Namibia Abyssal Plain. Comparisons of community membership clearly showed the differences between the Mediterranean communities and the community from the Namibia Abyssal Plain (data not shown). This is also supported by the fact that Yue and Clayton’s θ similarity coefficients within the group encompassing the Mediterranean sampling regions and the Namibia Abyssal Plain did not change significantly while raising the threshold for OTU delineation from 1 to 3% p-distance. By contrast, raising the threshold for OTU delineation lowered this similarity coefficient among all sampling regions of the South-West Atlantic (Argentine and Brazil Basin) and between the Angola and Guinea Abyssal Plain, resulting in similar sampling regions becoming more similar. At the same time, dissimilarities became more apparent. Together with the fact that raising the threshold for OTU delineation did not significantly raise the maximum number of sampling regions an OTU was present, this emphasizes both the similarities among some sampling regions as well as the dissimilarities found.
Apart from hydro-geographic features, which may pose barriers to dispersal among basins, such as the Antarctic Polar Front (Hunter and Halanych, 2008) or the Pacific Equatorial Current (Won et al., 2003), currents may influence dispersal (Young et al., 2008) and may equally explain the results from the South Atlantic, at least to some extent. The topography may be another explanation, as the Mediterranean is separated from the Atlantic by the Gibraltar Sill, which may prevent deep-sea species from the Atlantic to enter the Mediterranean (Sardà et al., 2004). Similarly, the mid-oceanic ridge may prevent dispersal among the communities from the South-East and South-West Atlantic, as shown for deep-sea bivalves, which are restricted to either the East or the West Atlantic (McClain et al., 2009). The Walvis Ridge, separating the Angola Basin from the Namibia Abyssal Plain has shown to prevent dispersal of peracarid crustaceans (Brandt et al., 2005). The different depths among some of the Atlantic and Mediterranean sampling regions may be another factor. Bathymetric patterns are known (Levin et al., 2001; Countway et al., 2007; Not et al., 2007; Brown et al., 2009), even though depth may not always be sufficient to explain community differences (Galand et al., 2010). The same holds true for differences in sediment characteristics among the sampling stations of the South-East Atlantic and the Mediterranean, as the sediment composition is known to affect community structure (Levin et al., 2001). Although the differences between communities isolated from sediment or sediment-overlaying water may be attributed to their habitat, the latter possibility is less likely as communities from sediment and sediment-overlaying water from the same sampling area (North Brazil Basin) were not significantly differentiated (P>0.05). Finally, many marine communities show a strong seasonal pattern (Levin et al., 2001; Ruhl and Smith, 2004; Fuhrman et al., 2006; Caron and Countway, 2009; Treusch et al., 2009) and can rapidly alter their community assemblage in areas with high seasonality in timescales in the order of days and weeks (Fuhrman and Hagström, 2008), rendering deep-sea microbial communities as dynamic as surface ones (Winter et al., 2009). The significant differences (P<0.05) between the communities from the South-East and the South-West Atlantic, as well as the Mediterranean, may therefore be attributed to differences in sampling time.
Lateral transport of organic material produced in coastal regions into greater depths is of major importance in maintaining diverse assemblages of microbial organisms at the seafloor (Salihoglu et al., 1991; Rabitti et al., 1994). Deep-sea abundance and biodiversity are, therefore, significantly correlated with depth and distance to the nearest coast, with the latter factor being more decisive (Boetius et al., 1996; Kröncke et al., 2003). Boetius et al. (1996) reported in their study of the eastern Mediterranean that the distance from coast is much more important than water depth for food availability at the seafloor and thus species diversity. Our results support these findings, with the community from the Ierapetra Basin being more diverse than most others.
Our data furthermore indicate that dispersal limitations by geographical distance may be another factor, as we had a very low but significant positive correlation between genetic and geographic distance (r=0.07, P<0.001). Several publications have emphasized the effect of spatial distance on microbial diversity (Papke et al., 2003; Whitaker et al., 2003; Ramette and Tiedje, 2007), and comprehensive reviews on microbial biogeography show that there is often a distance–decay relationship among microbial communities (Green and Bohannan, 2006; Martiny et al., 2006). Schauer et al. (2009) also reported a weak distance-decay relationship among the three abyssal plains of the South-East Atlantic. Likewise, communities of deep-sea microbes from depths of 3000 m of the Pacific showed a steady decline in similarity over a distance of 3500 km (Hewson et al., 2006). Distance–decay relationships may, therefore, be common given the vast distances found among abyssal plains, but may rather be of ecological nature, as geographical distance often correlates with environmental changes (Schauer et al., 2009).
A methodological problem could arise from the use of rDNA, as recent publications reported of a constant flux of inactive thermophilic bacteria into the arctic seabed, influencing the microbial community structure in this environment (Hubert et al., 2009). The high dispersal capacities and the possibility to form resting stages of many microbes make it indeed difficult to discriminate surface-derived microbes using rDNA. This would require other molecular markers, such as rRNA (Stoeck et al., 2007). Another methodological problem is most certainly the differences in sampling size, in terms of clone numbers. This may affect the result of community comparison between the different sampling areas. However, the most dominant OTUs will most probably always be detected and, as it is generally assumed that the dominant taxa contribute most the ecosystem’s function and consequently best describe that ecosystem (Scheckenbach et al., 2010), abundance-based statistics (for example, Yue and Clayton’s θ similarity coefficient) should therefore deliver comparable results. Equally, differences in sampling volume most likely affect sampling efficiency, but again, it can be assumed that the most dominant taxa will always be detected. And as OUT-based statistics produced similar results than sequence/phylogeny-based statistics, the reported dissimilarities between some sampling regions are most likely the result of true differences, even though the results may very well be biased owing to differences in sampling size and volume. Furthermore, different extraction methods and sample characteristics are known to affect sampling efficiency (Maier et al., 2009). As already stated above, the differences between communities isolated from sediment and sediment-overlaying water may therefore be attributed to the different extraction methods used. This, however, did not seem to significantly bias our results, as the community extracted from the sediment-overlaying water of the North Brazil Basin was very similar to the other communities of this region (North Argentine Basin, South Brazil Basin, North Brazil Basin) that were extracted from sediments, and was even not significantly differentiated from these (P>0.05, weighted UniFrac test).
The circumstances of our sampling and the missing abiotic and biotic parameters, among others the missing trophic levels, do not allow to elucidating the decisive deterministic and stochastic factors controlling the community composition reported here. Environmental factors, such as water masses and time, may be decisive factors, although, according to analysis of molecular variances, depth, habitat and temperature may all have a role (P<0.001), as well as geographic distance. It is, therefore, very likely that multiple processes operate at the same time to structure communities of deep-sea kinetoplastids. In this context, note that most communities were phylogenetically clustered, thus providing evidence for environmental filtering. Caron and Countway (2009) stated that even slight changes in environmental parameters can cause massive shifts in microbial community structure. This makes our findings of statistically indistinguishable communities of kinetoplastids in the range of thousands of kilometers from both the South-East Atlantic (Angola and Guinea Basin) and the South-West Atlantic (Argentine and Brazil Basin) all the more interesting, as they point to constant and homogenous environmental conditions prevailing over large spatial scales at abyssal depths.
References
Agogué H, Brink M, Dinasquet J, Herndl GJ . (2008). Major gradients in putatively nitrifying and non-nitrifying archaea in the deep North Atlantic. Nature 456: 788–791.
Arístegui J, Gasol JM, Duarte CM, Herndl GJ . (2009). Microbial oceanography of the dark ocean’s pelagic realm. Limnol Oceanogr 54: 1501–1529.
Azam F, Malfatti F . (2007). Microbial structuring of marine ecosystems. Nat Rev Microbiol 5: 782–791.
Azam F, Worden AZ . (2004). Oceanography. Microbes, molecules, and marine ecosystems. Science 303: 1622–1624.
Boetius A, Scheibe S, Tselepides A, Thiel H . (1996). Microbial biomass and activities in deep-sea sediments of the eastern Mediterranean: trenches are benthic hotspots. Deep Sea Res I 43: 1439–1460.
Brandt A, Brenkeb N, Andresa HG, Brixa S, Guerrero-Kommritza J, Mühlenhardt-Siegela U et al. (2005). Diversity of peracarid crustaceans (Malacostraca) from the abyssal plain of the Angola Basin. Organ Divers Evol 5: 105–112.
Brown MV, Philip GK, Bunge JA, Smith MC, Bissett A, Lauro FM et al. (2009). Microbial community structure in the North Pacific Ocean. ISME J 3: 1374–1386.
Caron DA, Countway PD . (2009). Hypotheses on the role of the protistan rare biosphere in a changing world. Aquat Microb Ecol 57: 227–238.
Cavender-Bares J, Ackerly DA, Baum D, Bazzaz FA . (2004). Phylogenetic overdispersion in Floridian oak communities. Am Nat 63: 823–843.
Cermeño P, Falkowski PG . (2009). Controls on diatom biogeography in the ocean. Science 325: 1539–1541.
Chao A, Lee SM . (1992). Estimating the number of classes via sample coverage. J Am Stat Assoc 87: 210–217.
Clark CG . (1992). DNA purification from polysaccharide-rich cells. In: Lee JJ, Soldo AT (eds). Protocols in Protozoology 1. Allen Press: Lawrence, pp D31–D32.
Countway PD, Gast RJ, Dennett MR, Savai P, Rose JM, Caron DA . (2007). Distinct protistan assemblages characterize the euphotic zone and deep sea (2500 m) of the western North Atlantic (Sargasso Sea and Gulf Stream). Environ Microbiol 9: 1219–1232.
Danovaro R, Gambi C, Dell'Anno A, Corinaldesi C, Fraschetti S, Vanreusel A et al. (2008a). Exponential decline of deep-sea ecosystem functioning linked to benthic biodiversity loss. Curr Biol 18: 1–8.
Danovaro R, Gambi C, Lampadariou N, Tselepides A . (2008b). Deep-sea biodiversity in the Mediterranean Basin: testing for longitudinal, bathymetric and energetic gradients. Ecography 31: 231–244.
Darling KF, Wade CM, Stewart IA, Kroon D, Dingle R, Brown AJ . (2000). Molecular evidence for genetic mixing of Arctic and Antarctic subpolar populations of planktonic foraminifers. Nature 405: 43–47.
De Vargas C, Norris R, Zaninetti L, Gibb SW, Pawlowski J . (1999). Molecular evidence of cryptic speciation in planktonic foraminifers and their relation to oceanic provinces. Proc Natl Acad Sci USA 96: 2864–2868.
Desper R, Gascuel O . (2002). Fast and accurate phylogeny reconstruction algorithms based on the minimum-evolution principle. J Comput Biol 19: 687–705.
Edel-Hermann V, Dreumont C, Pérez-Piqueres A, Steinberg C . (2004). Terminal restriction fragment length polymorphism analysis of ribosomal RNA genes to assess changes in fungal community structure in soils. FEMS Microbiol Ecol 47: 397–404.
Excoiffier L, Smouse PE, Quattro JM . (1992). Analysis of molecular variance inferred from metric distances among DNA haplotypes: application to human mitochondrial DNA restriction data. Genetics 131: 479–491.
Faith DP . (1992). Conservation evaluation and phylogenetic diversity. Biol Conserv 61: 1–10.
Fuhrman JA, Hagström A . (2008). Bacterial and archaeal community structure and its pattern. In: Kirchman, DL (ed). Microbial Ecology of the Oceans, 2nd edn Wiley: Hoboken.
Fuhrman JA, Hewson I, Schwalbach MS, Steele JA, Brown MV, Naeem S . (2006). Annually reoccurring bacterial communities are predictable from ocean conditions. Proc Natl Acad Sci USA 103: 13104–13109.
Fuhrman JA . (2009). Microbial community structure and its functional implications. Nature 459: 193–199.
Fukuda H, Sohrin R, Nagata T, Koike I . (2007). Size distribution and biomass of nanoflagellates in meso- and bathypelagic layers of the subarctic Pacific. Aquat Microb Ecol 46: 203–207.
Galand PE, Casamayor EO, Kirchman DL, Lovejoy C . (2009). Ecology of the rare microbial biosphere of the Arctic Ocean. Proc Natl Acad Sci USA 106: 22427–22432.
Galand PE, Potvin M, Casamayor EO, Lovejoy C . (2010). Hydrography shapes bacterial biogeography of the deep Arctic Ocean. ISME J 4: 564–576.
Gotelli NJ, Entsminger GI . (2003). Swap algorithms in null model analysis. Ecology 84: 532–535.
Green J, Bohannan BJ . (2006). Spatial scaling of microbial biodiversity. Trends Ecol Evol 21: 501–507.
Hausmann K, Hülsmann N, Radek R . (2003). Protistology, 3rd edn. E Schweizerbart’sche Verlagsbuchhandlung: Stuttgart.
Hewson I, Steele JA, Capone DG, Fuhrman JA . (2006). Remarkable heterogeneity in meso- to bathypelagic bacterioplankton assemblage composition. Limnol Oceanogr 51: 1274–1283.
Hubert C, Loy A, Nickel M, Arnosti C, Baranyi C, Brüchert V et al. (2009). A constant flux of diverse thermophilic bacteria into the cold Arctic seabed. Science 325: 1541–1544.
Hunter RL, Halanych KM . (2008). Evaluating connectivity in the brooding brittle star Astrotoma agassizii across the Drake Passage in the Southern Ocean. J Heredity 99: 137–148.
Katoh K, Toh H . (2010). Parallelization of the MAFFT multiple sequence alignment program. Bioinformatics 26: 1899–1900.
Kembel SW, Cowan PD, Helmus MR, Cornwell WK, Morlon H, Ackerly DD et al. (2010). Picante: R tools for integrating phylogenies and ecology. Bioinformatics 26: 1463–1464.
Kirchman DL, Cottrell MT, Lovejoy C . (2010). The structure of bacterial communities in the western Arctic Ocean as revealed by pyrosequencing of 16S rRNA genes. Environ Microbiol 12: 1132–1143.
Kouridaki I, Polymenakou PN, Tselepides A, Mandalakis M, Smith Jr KL . (2010). Phylogenetic diversity of sediment bacteria from the deep Northeastern Pacific Ocean: a comparison with the deep Eastern Mediterranean Sea. Int Microbiol 13: 143–150.
Kraft NJ, Cornwell WK, Webb CO, Ackerly DD . (2007). Trait evolution, community assembly, and the phylogenetic structure of ecological communities. Am Nat 170: 271–283.
Kröncke I, Türkay M, Fiege D . (2003). Macrofauna communities in the eastern Mediterranean deep sea. Mar Ecol 24: 193–216.
Lara E, Moreira D, Vereshchaka A, López-García P . (2008). Pan-oceanic distribution of new highly diverse clades of deep-sea diplonemids. Environ Microbiol 11: 47–55.
Lecroq B, Gooday AJ, Pawlowski J . (2009). Global genetic homogeneity in the deep-sea foraminiferan Epistominella exigua (Rotaliida: Pseudoparrellidae). Zootaxa 2096: 23–32.
Letunic I, Bork P . (2007). Interactive Tree Of Life (iTOL): an online tool for phylogenetic tree display and annotation. Bioinformatics 23: 127–128.
Levin LA, Etter RJ, Rex MA, Gooday AJ, Smith CR, Pineda J et al. (2001). Environmental influences on regional deep-sea species diversity. Annu Rev Ecol Syst 32: 51–93.
López-García P, Rodríguez-Valera F, Pedrós-Alió C, Moreira D . (2001). Unexpected diversity of small eukaryotes in deep-sea Antarctic plankton. Nature 409: 603–607.
Lozupone C, Knight R . (2005). UniFrac: a new phylogenetic method for comparing microbial communities. Appl Environ Microbiol 71: 8228–8235.
Maier RM, Pepper IL, Gerba CP . (2009). Environmental Microbiology, 2nd edn Academic Press: New York, pp 245–246.
Martiny JB, Bohannan BJ, Brown JH, Colwell RK, Fuhrman JA, Green JL et al. (2006). Microbial biogeography: putting microorganisms on the map. Nat Rev Microbiol 4: 102–112.
McClain CR, Hardy SM . (2010). The dynamics of biogeographic ranges in the deep sea. Proc Biol Sci 277: 3533–3546.
McClain CR, Rex MA, Etter RJ . (2009). Deep-sea macroecology. In: Witman JD, Roy K (eds). Marine Macroecology. University of Chicago Press: Chicago, pp 65–100.
Moreira D, López-García P, Vickerman K . (2004). An updated view of kinetoplastid phylogeny using environmental sequences and a closer outgroup: proposal for a new classification of the class Kinetoplastea. Int J Syst Evol Microbiol 54: 1861–1875.
Not F, Gausling R, Azam F, Heidelberg JF, Worden AZ . (2007). Vertical distribution of picoeukaryotic diversity in the Sargasso Sea. Environ Microbiol 9: 1233–1252.
Papke RT, Ramsing NB, Bateson MM, Ward DM . (2003). Geographical isolation in hot spring cyanobacteria. Environ Microbiol 5: 650–659.
Pawlowski J, Fahrni J, Lecroq B, Longet D, Cornelius N, Excoffier L et al. (2007). Bipolar gene flow in deep-sea benthic foraminifera. Mol Ecol 16: 4089–4096.
Pruesse E, Quast C, Knittel K, Fuchs B, Ludwig W, Peplies J et al. (2007). SILVA: a comprehensive online resource for quality checked and aligned ribosomal RNA sequence data compatible with ARB. Nucleic Acids Res 35: 7188–7196.
R Development Core Team (2010). R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. http://www.R-project.org/.
Rabitti S, Bianchi F, Boldrin A, Da Ros L, Socal G, Totti C . (1994). Particulate matter and phytoplankton in the Ionian Sea. Oceanol Acta 17: 297–307.
Ramette A, Tiedje JM . (2007). Biogeography: an emerging cornerstone for understanding prokaryotic diversity, ecology, and evolution. Microb Ecol 53: 197–207.
Rex MA, Etter RJ, Morris JS, Crouse J, McClain CR, Johnson NA et al. (2006). Global bathymetric patterns of standing stock and body size in the deep-sea benthos. Mar Ecol Prog Ser 317: 1–8.
Rex MA, Etter RJ . (2010). Deep-Sea Biodiversity Pattern and Scale. Harvard University Press: Cambridge, MA.
Rice P, Longden I, Bleasby A . (2000). EMBOSS: the European Molecular Biology Open Software Suite. Trends Genet 16: 276–277.
Ruhl HA, Smith Jr KL . (2004). Shifts in deep-sea community structure linked to climate and food supply. Science 305: 513–515.
Salihoglu I, Saydam C, Bastürk Ö, Yilmaz K, Gö D, Hatipoglu E et al. (1991). Transport of nutrients and chlorophyll-a by mesoscale eddies in the northeastern Mediterranean. Mar Chem 29: 375–390.
Sardà F, Calafat A, Flexas MM, Tselepides A, Canals M, Espino M et al. (2004). An introduction to Mediterranean deep-sea biology. Sci Marina 68: 7–38.
Schauer R, Bienhold C, Ramette A, Harder J . (2009). Bacterial diversity and biogeography in deep-sea surface sediments of the South Atlantic Ocean. ISME J 4: 159–170.
Scheckenbach F, Hausmann K, Wylezich C, Weitere M, Arndt H . (2010). Large-scale patterns in biodiversity of microbial eukaryotes from the abyssal sea floor. Proc Natl Acad Sci USA 107: 115–120.
Schloss PD, Westcott SL, Ryabin T, Hall JR, Hartmann M, Hollister EB et al. (2009). Introducing Mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol 75: 7537–7541.
Shah Salani F . (2009). Kinetoplastids from the Abyss—emerging patterns in the comparative analysis of phylogenetic community structure and diversity. Diploma thesis, University of Cologne, Cologne, Germany.
Smith CR, De Leo FC, Bernardino AF, Sweetman AK, Arbizu PM . (2008). Abyssal food limitation, ecosystem structure and climate change. Trends Ecol Evol 23: 518–528.
Stamatakis A . (2006). RAxML-VI-HPC: maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models. Bioinformatics 22: 2688–2690.
Stoeck T, Taylor GT, Epstein SS . (2003). Novel eukaryotes from the permanently anoxic Cariaco Basin (Caribbean Sea). Appl Environ Microbiol 69: 5656–5663.
Stoeck T, Zuendorf A, Breiner HW, Behnke A . (2007). A molecular approach to identify active microbes in environmental eukaryote clone libraries. Microb Ecol 53: 328–339.
Thioulouse J, Chessel D, Dolédec S, Olivier JM . (1997). ADE-4: a multivariate analysis and graphical display software. Stat Comput 7: 75–83.
Treusch AH, Vergin KL, Finlay LA, Donatz MG, Burton RM, Carlson CA et al. (2009). Seasonality and vertical structure of microbial communities in an ocean gyre. ISME J 3: 1148–1163.
Varela MM, van Aken HM, Herndl GJ . (2008). Abundance and activity of Chloroflexi-type SAR202 bacterioplankton in the meso- and bathypelagic waters of the (sub)tropical Atlantic. Environ Microbiol 10: 1903–1911.
Von der Heyden S, Cavalier-Smith T . (2005). Culturing and environmental DNA sequencing uncover hidden kinetoplastid biodiversity and a major marine clade within ancestrally freshwater Neobodo designis. Int J Syst Evol Microbiol 55: 2605–2621.
Webb C, Ackerly D, McPeek M, Donoghue M . (2002). Phylogenies and community ecology. Annu Rev Ecol Syst 33: 475–505.
Whitaker RJ, Grogan DW, Taylor JW . (2003). Geographic barriers isolate endemic populations of hyperthermophilic archaea. Science 301: 976–978.
Winter C, Kerros ME, Weinbauer MG . (2009). Seasonal changes of bacterial and archaeal communities in the dark ocean: evidence from the Mediterranean Sea. Limnol Oceanogr 54: 160–170.
Won Y, Young CR, Lutz RA, Vrijenhoek RC . (2003). Dispersal barriers and isolation among deep-sea mussel populations (Mytilidae: Bathymodiolus) from eastern Pacific hydrothermal vents. Mol Ecol 12: 169–184.
Young CR, Fujio S, Vrijenhoek RC . (2008). Directional dispersal between mid-ocean ridges: deep-ocean circulation and gene flow in Ridgeia piscesae. Mol Ecol 17: 1718–1731.
Yue JC, Clayton MK . (2005). A similarity measure based on species proportions. Commun Stat Theor M 34: 2123–2131.
Acknowledgements
We thank Michael Türkay; Pedro Martínez Arbizu; Kai George; and the captains, the crews and the scientific crews of R/V Meteor cruises 63/2 (expedition DIVA2), 71/2 and 79/1 (expedition DIVA3). This work was part of the Census of the Diversity of Abyssal Marine Life (CeDAMar) and was supported by the German Research Foundation (DFG AR 288/15-1). We thank Claudia Wylezich for a thorough review of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on The ISME Journal website
Rights and permissions
About this article
Cite this article
Salani, F., Arndt, H., Hausmann, K. et al. Analysis of the community structure of abyssal kinetoplastids revealed similar communities at larger spatial scales. ISME J 6, 713–723 (2012). https://doi.org/10.1038/ismej.2011.138
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2011.138
Keywords
This article is cited by
-
Large variability of bathypelagic microbial eukaryotic communities across the world’s oceans
The ISME Journal (2016)