Abstract
Variation in traits causes bacterial populations to respond in contrasting ways to environmental drivers. Learning about this will help us understand the ecology of individual populations in complex ecosystems. We used 454 pyrosequencing of the hypervariable region V6 of the 16S rRNA gene to study seasonal dynamics in Baltic Sea bacterioplankton communities, and link community and population changes to biological and chemical factors. Surface samples were collected from May to October 2003 and in May 2004 at the Landsort Deep in the central Baltic Sea Proper. The analysis rendered, on average, 20 200 sequence reads for each of the eight samples analyzed, providing the first detailed description of Baltic Sea bacterial communities. Community composition varied dramatically over time, supporting the idea of strong temporal shifts in bacterioplankton assemblages, and clustered according to season (including two May samples from consecutive years), suggesting repeatable seasonal succession. Overall, community change was most highly correlated with change in phosphorus concentration and temperature. Individual bacterial populations were also identified that tightly co-varied with different Cyanobacteria populations. Comparing the abundance profiles of operational taxonomic units at different phylogenetic distances revealed a weak but significant negative correlation between abundance profile similarity and genetic distance, potentially reflecting habitat filtering of evolutionarily conserved functional traits in the studied bacterioplankton.
Similar content being viewed by others
Introduction
It is becoming increasingly clear that aquatic microbial communities are highly dynamic. Learning about temporal changes in microbial populations in relation to environmental parameters is a promising avenue in microbial ecology, as it provides insights into the environmental control and functional traits of individual populations, as well as to how populations are interlinked. Long-term monitoring of aquatic bacterial communities with molecular fingerprinting methods have shown that community composition follow annually reoccurring patterns (Crump and Hobbie, 2005; Fuhrman et al., 2006; Shade et al., 2007). The recent efforts to model, formalize and explain temporal changes in microbial communities builds on an existing body of knowledge, wherein pronounced seasonal changes in certain components of bacterioplankton communities have been described in response to environmental change (for example, see Pernthaler et al., 1998; Pinhassi and Hagström, 2000; Schauer et al., 2003; Kan et al., 2006). For example, phytoplankton blooms are known to cause pronounced changes in bacterioplankton communities and populations in lakes (Riemann and Winding, 2001; Eiler and Bertilsson, 2004; Kent et al., 2007) and in marine waters (Riemann et al., 2000; Pinhassi et al., 2004; Fandino et al., 2005).
The Baltic Sea is the second largest brackish water reservoir on Earth, serving as a drainage area for ∼90 million people in 14 different countries (Rönnberg and Bonsdorff, 2004). Its surface water is characterized by a salinity gradient from ∼2 to 8 practical salinity units (PSU) from north to south, and the water column in the Baltic Sea proper is stratified because of a strong pycnocline (Stigebrandt, 2001). Typical estuarine environments, such as river outlets, are mixing zones between fresh and marine waters, in which short retention times limit the possibilities for autochthonous communities to establish (Crump et al., 2004). In contrast, the central Baltic Sea has a long retention time (>5 years (Wulff and Stigebrandt, 1989)) and is conceivably dominated by microbes that have adapted to this specific environment. In the Baltic Sea proper (∼6 PSU), Riemann et al. (2008) found that surface bacterioplankton communities seemed to be strongly influenced by typical freshwater bacterial groups within Actinobacteria, Verrucomicrobia and Betaproteobacteria, whereas many typical marine taxa appeared to be missing. Hence, although the low salinity of the Baltic Sea surface water seems to be conducive for growth of freshwater bacteria (Kisand et al., 2005; Riemann et al., 2008), typical marine bacterial populations may be constrained by requirements for higher salinity. It should be noted that estuarine bacterioplankton communities are exposed to gradients in environmental parameters other than salinity, for example, dissolved organic matter, nutrients and temperature. Hence, ultimately, a complex matrix of variables likely selects for a bacterioplankton community uniquely adapted to the estuarine environment.
It is becoming commonly accepted that the biodiversity of bacterioplankton is composed of two elements. First, a set of abundant taxa that likely perform most of the ecosystem function. These are detected using the standard repertoire of molecular tools. Second, a seedbank of presumably less active, rare taxa, which are currently difficult to detect and quantify (Pedrós-Alio, 2006). Although the presence of a rare biosphere in aquatic environments seems to be generally accepted, our current knowledge about the receptiveness of these bacteria to the environmental changes, their potential seedbank function and their importance for the adaptability and functional plasticity of aquatic bacterioplankton assemblages is largely missing.
So far, studies assessing the temporal dynamics of aquatic microbial communities have either addressed the dynamics of certain phylogenetic groups, in isolation from the rest of the microbial community, or have been based on molecular fingerprinting tools, which do not automatically give taxonomic information on the observed community components. Moreover, resolutions of fingerprinting methods are orders of magnitude from identifying low-abundance populations known to contribute greatly to the diversity of planktonic microbial communities. High-throughput next-generation sequencing, such as 454 pyrosequencing (Margulies et al., 2005), of rRNA genes offers an alternative, in which detailed community structure can be achieved in combination with fairly high taxonomic resolution (Sogin et al., 2006; Andersson et al., 2008). For instance, 454 pyrosequencing analysis of ∼118 000 16S sequences from the North Atlantic deep sea revealed a tremendous microbial diversity accounted for by thousands of low-abundance populations (Sogin et al., 2006). In addition, this type of high-throughput rRNA gene sequencing enables statistically robust assessments of community and population dynamics.
In this study, we used extensive 454 pyrosequencing of the V6 region of the 16S rRNA gene to simultaneously follow seasonal changes in rare and abundant populations in the pelagic bacterial community of the Landsort Deep in the central Baltic Sea. The dynamic properties of bacterial communities as well as individual populations at contrasting phylogenetic resolution were analyzed. Both environmental drivers and population linkages were used to explain the observed seasonality in the communities. We show that the studied bacterioplankton community feature strong temporal shifts and seasonal clustering, and link the dynamics of individual taxa and phylogenetic groups with environmental drivers.
Methods
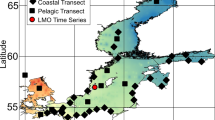
Sampling, DNA extraction and microbial community dynamics, as analyzed by denaturing gradient gel electrophoresis and 16S rRNA gene clone libraries, are detailed in Riemann et al. (2008). Briefly, samples were collected from 3 m depth at the Landsort Deep station (BY31, 58°35.90′N 18°14.21′E, depth 459 m) every 1–3 weeks from March 2003 to May 2004 using a 5 l Niskin bottle. Background environmental data were retrieved from Riemann et al. (2008) and/or the Swedish Marine Monitoring Program (http://www.smhi.se). A subset of the environmental parameters is shown in Table 1. For DNA extractions, cells from pre-filtered (<3.0 μm) water samples were collected on 0.22 μm Sterivex filters (Millipore, Solna, Sweden) using washed, sterile tubing and utensils. Filters were kept frozen at −20 °C until community DNA was extracted using an enzyme/phenol–chloroform protocol (Riemann et al., 2008). Extracted DNA was stored frozen in Tris–EDTA buffer (10 mM Tris, 1 mM EDTA, pH 8.0) until use.
Eight samples were selected for 454 pyrosequencing at the ICoMM (International Census of Marine Microbes) core facility at the Marine Biology Laboratory (Woods Hole, MA, USA). Sampling dates were chosen to cover cold spring conditions, warm summer conditions with sequential Cyanobacterial blooms and the post-bloom phase in early autumn. In addition, a spring sample from the consecutive year was analyzed to test for annually reoccurring patterns in community composition. To amplify the V6 region of bacterial 16S rRNA genes, a primer cocktail containing 5 versions of the 967 forward primer and 4 versions of the 1046 reverse primer was used (Huber et al., 2007). Pyrosequencing was performed on the 454 GS20 platform following the detailed protocol described in Sogin et al. (2006). Primer sequences were trimmed from the beginning and end of each read and sequences likely to be of low quality were removed, as described in Huse et al. (2007). All 454 sequences can be downloaded from the VAMPS database (http://vamps.mbl.edu/; project id: BSP) as well as from the NCBI Short Read Archive, accession number: SRA009836.
All non-redundant 454 sequences were extracted and cross-compared with BLASTN (Altschul et al., 1997). The similarity between two sequences was calculated as the number of matching base pairs divided by the number of base pairs available for alignment. Number of matching base pairs=alignment identity × alignment length. Number of base pairs available for alignment=alignment length+number of base pairs that flank the alignment. For example, if the alignment is preceded by one base on sequence A and three bases on sequence B, and followed by one base on each sequence, the number of base pairs available will be alignment length+two. In other words, the similarity between two sequences, A and B, can be expressed as alignment identity × alignment length/(alignment length+min (alignment startA, alignment startB)+min(lengthA−alignment endA, lengthB−alignment endB)). Sequences were clustered into operational taxonomic units (OTUs) with complete linkage clustering using an in-house developed Perl (http://www.perl.org/) script.
The 269 420 bacterial 16S rRNA gene sequences longer than 1200 bp with good Pintail scores were downloaded from Ribosomal Database Project v. 10.7 (RDP; http://rdp.cme.msu.edu/) (Cole et al., 2009) and formatted into a local BLAST database. Each 454 sequence (one per group of identical sequences) was BLASTN searched against the RDP database with default parameters, and inherited the taxonomic annotation (down to genus level) of the best scoring RDP hit, fulfilling the criteria of ⩾90% identity over an alignment of length ⩾40 bp. If no such hit was found, the sequence was classified as ‘no match’. Each OTU was classified according to the 454 sequence in the OTU with highest number of reads in the total data set.
Total bacterial abundance varied over the season and peaked in August (Table 1). To account for this when interpreting seasonal dynamics in groups and populations, OTU abundances were calculated by multiplying the relative OTU frequencies with total bacterial abundance in each of the samples.
All statistical analyses and graphs were generated in R (http://www.r-project.org/). For k-means clustering (MacQueen, 1967), the Lloyd–Forgy algorithm was used with Pearson correlation for distance calculations. BIOENV analysis was conducted with the R package Vegan, using Spearman rank correlation and the Morisita–Horn community dissimilarity index.
Results
A subset of the seasonal environmental data is shown in Table 1. Temperature increased from ∼1 °C in March to 15 °C and 20 °C in June and July, respectively. Phytoplankton biomass (proxied by chlorophyll a) was low in March (1 μg l−1), increased to 6.8 μg l−1 in April (data not shown) and then dropped again. The spring bloom caused inorganic nitrogen to decrease rapidly to ∼0.02 μM. As blooms of the Cyanobacteria Aphanizomenon sp. and Pseudanabaena sp. developed in late summer, chlorophyll a increased to a level of 2–3 μg l−1. On the last sampling date (October 8), the Cyanobacterial abundance was low (0–0.7 m l−1) and the temperature had decreased to 10 °C. Bacterial abundance changed from ∼1 × 106 ml−1 in spring and early summer to a peak level of 3–4 × 106 ml−1 in July/August.
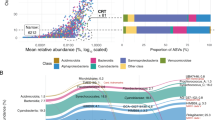
On average, 20 200 pyrosequencing reads of the V6 region of the 16S rRNA gene were obtained from each of eight surface water samples collected from May to October 2003 and in May 2004 (Table 2). Sequences were clustered into OTUs by complete linkage clustering. To determine a relevant sequence similarity cutoff for delineating OTUs, an in silico simulation was carried out using near-full-length (>1200 bp) bacterial 16S rRNA gene sequences from RDP. A 97% similarity cutoff for the V6 region sequences typically resolved near-full-length 16S rRNA gene sequences that were less than 97% similar, and this cutoff was used to delineate OTUs in the downstream analyses (Figure 1; Supplementary Figure 1).
Distribution of pair-wise sequence similarities for full-length (>1200 bp) RDP (Ribosomal Database Project) sequences having ⩾97% identity in their V6 regions. All 454 sequences that exactly matched an RDP sequence were selected and their matching RDP sequences retrieved. The sequences were cross-compared and pair-wise sequence similarities were calculated using: (a) the 454 sequences (applying the same metric as for OTU clustering (Methods)), and (b) the full-length RDP sequences using online tools at RDP. Pairs with ⩾97% V6 sequence similarity were selected and their RDP sequence similarities are displayed in the histogram.
A total of 4624 OTUs (97% cutoff) were obtained after removing single-read OTUs (4863) that could have resulted from sequencing errors. Of these 4624 OTUs, 1211 (70% of total reads) were 100% identical to sequences in RDP, whereas 808 (7% of reads) were <95% similar to the closest RDP match, and likely represent novel bacterial species (Supplementary Figure 2). Of the 4624 OTUs, 1182 were observed in a single seasonal sample, whereas only 76 were observed in all 8 samples. The most abundant OTU in each sample was also observed in all other samples; however, their representation varied dramatically. For example, the dominant OTU in May 2003 (8% of reads) was only detected with a single read on July 30 (<0.01% of reads), but became abundant again in May 2004 (10% of reads), illustrating how a component of the microbial seedbank can become important in response to environmental change (see Supplementary Table 1 and 2 for detailed information on the most abundant OTUs).
Despite the pronounced dynamics in the bacterioplankton community, pair-wise comparisons of OTU frequencies among the sampled communities revealed that the samples in general (seven out of eight) were most similar to a sample adjacent in time, and hence the samples clustered according to time point (Figure 2). This was independent of the similarity cutoff (90–99% similarity) used for OTU grouping (data not shown). In addition, the two May samples from consecutive years were highly similar, suggesting an annually reoccurring pattern in community composition.
Pair-wise bacterial community similarity matrix for the eight seasonal Baltic Sea samples. Similarities (in gray scale) were calculated by Pearson correlation of OTU (operational taxonomic unit) frequencies and exemplified by the insert scatter plot showing August 27 versus October 8 (each dot is one OTU; frequencies displayed in log scale). The dendrogram represents complete linkage clustering of the samples based on the pair-wise similarities.
Samples were dominated by Actinobacteria, Bacteroidetes, Cyanobacteria, Verrucomicrobia and Proteobacteria of the alpha, beta and gamma classes (Table 2; Supplementary Figure 3). At this taxonomic resolution, a peak of Alphaproteobacteria was observed in May, a Cyanobacterial peak in early summer (June 4), another peak of Alphaproteobacteria occurred in late summer (July 30) and a peak of Actinobacteria was apparent in the autumn samples (August–October). Resolving the data into OTUs (Figure 3a) revealed some interesting differences in ‘blooming patterns’ between phyla and classes, for example, whereas the Cyanobacterial peak in June 4 was composed largely of sequence reads from two OTUs (representing 88% of Cyanobacteria), the expansion of Alphaproteobacteria on July 30 comprised a large group of OTUs (the most abundant representing only 16% of Alphaproteobacteria). To further resolve the data, we used a simple clustering method, k-means clustering (MacQueen, 1967), to partition the OTUs into seven clusters based on similarities in abundance profiles. This clearly showed that the spring and late summer Alphaproteobacterial peaks were composed of distinct OTUs with contrasting seasonality (Figure 3b, clusters 1, 5 and 7). Among the Actinobacteria, low-abundance OTUs that decreased gradually in abundance from May to July 30 were detected, distinct from those peaking in autumn (Figure 3b, clusters 2 and 5).
Abundance profiles for operational taxonomic units (OTUs) represented by at least 100 reads in the total data set. In panel (a), each graph represents one phylum or class for Proteobacteria. In panel (b), the same OTUs (colored as in panel a) have been clustered according to seasonality into seven clusters by k-means clustering. The y axes display OTU abundances with bacterial cell counts taken into consideration.
To examine how biotic and abiotic factors were related to overall community composition, change in each environmental parameter was compared with change in community composition. Difference in total phosphorus concentration, followed by temperature, was most strongly correlated with community dissimilarity (Table 3). We also performed BIOENV analysis, which finds the set of environmental parameters that in combination correlates strongest with overall change in community composition (Clarke and Ainsworth, 1993). The highest rank correlation was obtained with a combination of temperature, chlorophyll a, silicate and total phosphorus concentration (ρ=0.66). To analyze how individual populations correlated with individual environmental parameters, the abundance profile of each of the 155 most frequent OTUs (represented by >100 reads) was correlated with each of the measured environmental parameters (Figure 4). The analysis revealed that multiple Alphaproteobacteria correlated strongly (Pearson r>0.92) with total nitrogen concentration and slightly less strongly with Pseudanabaena abundance. Moreover, a few specific OTUs co-varied with filamentous Cyanobacteria: a Bacteroidetes correlated with Aphanizomenon and Anabaena, one Gammaproteobacteria with Pseudanabaena and another one with Anabaena, and a Verrucomicrobia correlated with Aphanizomenon. These tight correlations (all Pearson r>0.96, P<0.0001) may indicate either strong functional coupling or coregulation by some unidentified environmental variable.
Correlations between environmental parameters and individual operational taxonomic units (OTUs) (represented by >100 reads). Colors indicate r-values of Pearson correlations between environmental parameters (columns) and OTUs (rows). The OTUs were ordered according to taxonomic affiliations (in alphabetic order). The dendrogram represents clustering of environmental parameters based on similarities in r-values, and was included to highlight correlations among environmental parameters.
It may be speculated that phylogenetically related bacteria have similar roles in the environment. According to the k-means clustering (Figure 3) and the correlations with environmental parameters (Figure 4), OTUs belonging to the same phylum tended to co-vary, indicating similarities in functionality. To address the potential link between phylogeny and function further, a phylogenetic tree was constructed based on the most frequent OTUs of a single phylum, the Actinobacteria, and the abundance profiles of the OTUs were compared (Figure 5). Although exceptions occurred (for example, within clade AcIB), closely related OTUs tended to display more similar abundance profiles than distantly related ones, and pair-wise similarity in abundance profile (measured by Spearman correlation) was significantly negatively correlated with pair-wise genetic distance (Pearson r=−0.12, Mantel test P=0.041). The correlation remained when the analysis was extended to include OTUs from all bacterial phyla with at least 10 reads in the data set (Pearson r=−0.13, Mantel test P<0.0001). The relationship was not caused only by very closely related OTUs being highly synchronized, as abundance profile similarities decreased gradually up to genetic distances of 20% (Figure 6).
Phylogeny and abundance profiles for the most frequent (min 200 reads) operational taxonomic units (OTUs) of Actinobacteria. The phylogenetic tree is based on best matching RDP (Ribosomal Database Project) (>1200 bp) sequences, and was constructed using the Weighbor algorithm (Bruno et al., 2000) in RDP. Only OTUs with an RDP best match of at least 97% identity over at least 50 bp were included. Square sizes represent OTU abundances (averages in gray). Classifications into actinobacterial clades were based on studies carried out by Warnecke et al. (2004); Allgaier and Grossart (2006); Holmfeldt et al. (2009) using GenBank accession numbers of RDP sequences.
Distributions of pair-wise abundance profile similarities in intervals of increasing phylogenetic distance. Phylogenetic distances were based on best matching (>1200 bp) RDP (Ribosomal Database Project) sequence and calculated in RDP using Jukes–Cantor correction. Pair-wise abundance profile similarity was measured using Spearman rank order correlation. Each box shows the distribution of abundance profile similarities (Spearman, ρ) within a genetic distance interval; 50% of data points are within the box, 75% within the whiskers and the thick black line represents the median value. Only operational taxonomic units (OTUs) with at least 10 reads in the data set and with an RDP match of at least 97% identity over at least 50 bp were included.
Discussion
In concordance with an earlier cloning-based study, including some of the samples analyzed here (Riemann et al., 2008), the Baltic Sea bacterioplankton hosted several phylogenetic groups of Bacteria known to be prominent in freshwater (Glöckner et al., 2000; Zwart et al., 2003). Freshwater and marine bacterioplankton communities typically show limited overlap in composition (Troussellier et al., 2002; Selje and Simon, 2003), whereas estuaries appear as mixing zones. However, given the location of the sampling station far from any river mouth, direct influence of advected freshwater bacteria on the indigenous bacterioplankton community is unlikely (Riemann et al., 2008).
Hence, the observation that about a third of the 454 sequence reads were related to bacterial phyla considered to be characteristic for freshwater ecosystems (Actinobacteria (25%), Betaproteobacteria (6%) and Verrucomicrobia (4%)), suggests that members of these broader phylogenetic groups also thrive at the local brackish environmental conditions in the Baltic Proper (salinity of ∼6). Actinobacteria were shown to account for about a quarter of the bacterioplankton in the Northern Baltic Sea (salinity 2–5), as determined by fluorescent in situ hybridization (Holmfeldt et al., 2009). Here, we show that they are well represented also in the central Baltic Sea.
The most common Cyanobacterial phylotypes were related to Synechococcus spp. In the Baltic Sea, these cells generally have a diameter of <1 μm (Stal et al., 2003) and therefore readily pass our 3 μm pre-filter. These Cyanobacteria are commonly found in the Baltic proper during summer (Stal et al., 2003; Labrenz et al., 2007; Haverkamp et al., 2009). Their highest abundance was observed when chlorophyll a levels were relatively low, for example, in between the phytoplankton spring bloom in April and the bloom of heterocystous Cyanobacteria in July and August, suggesting that these unicellular Cyanobacteria are successful competitors during this period of low nutrient availability, increasing predation pressure and increasing temperatures.
Repeatable seasonal succession patterns (phenology) in bacterioplankton community composition have recently been shown for lakes (Shade et al., 2007), rivers (Crump and Hobbie, 2005), coastal waters (Fuhrman et al., 2006) and the open ocean (Morris et al., 2005). Whereas earlier studies targeting this phenomenon were based on fingerprinting methods that only target the most abundant members of the community, the present study represents the first attempt to apply extensive 454 sequencing to study seasonal dynamics, capturing both abundant and rare populations and providing taxonomic information on the observed OTUs. With this powerful method, we effectively show how members of the ‘rare biosphere’ can become important in response to environmental change, rebutting the idea of the rare biosphere as being merely free DNA or dead cells (Stoeck and Epstein, 2009). The clustering of samples according to time point, and the similarity in community composition between the two May samples from consecutive years, suggest annually reoccurring patterns in bacterial composition also in the brackish Baltic Sea proper. Similar to the much more well-studied phytoplankton communities (Reynolds, 1984), phenology thus seems to be a general feature of pelagic bacterial communities that experience climate-driven seasonal environmental changes.
One plausible driver of seasonal change in community composition is temperature (Shiah and Ducklow, 1994) and this could certainly be relevant in the Baltic Sea, as growth of bacterioplankton in this system is at least seasonally limited by temperature (Hagström and Larsson, 1984). Our analysis suggested that temperature indeed was a major structuring factor. In addition, phosphorus co-varied strongly with community composition. It has previously been shown that the availability of phosphorus may limit bacterioplankton growth in the open oceans (for example, see (Cotner et al., 1997)) and in the Northern Baltic Sea (Zweifel et al., 1993), and select for specific bacteria adapted to such nutrient-deprived conditions (Pinhassi and Hagström, 2000). Hence, conceivably, the pronounced seasonal oscillations in phosphorus concentration observed at the sampling station, with potentially limiting conditions associated with the Cyanobacterial summer blooms (Table 1), exert a strong selection pressure onto the bacterial community. However, more time points would be needed to disentangle the specific effects of the individual environmental parameters on the community.
Linkages between the combined bacterioplankton community composition and phytoplankton blooms have been identified in freshwater systems (Riemann and Winding, 2001; Eiler and Bertilsson, 2004). This is not surprising, as phytoplankton blooms may represent a significant input of biologically labile organic compounds available to the bacterioplankton community. A recent study covering a wide range of aquatic systems (lakes, the Baltic Sea and treated drinking water) suggest that there may be more intricate linkages between heterotrophic bacteria and bloom-forming Cyanobacteria, as many isolated bacterial strains were capable of either enhancing, or occasionally inhibiting, the growth of bloom-forming Cyanobacteria (Berg et al., 2009). The strong positive correlations observed between a few individual OTUs and bloom-forming Cyanobacteria in this study indicate functional couplings and motivate directed studies on their potential interactions.
We observed an inverse correlation between genetic distance and similarity in abundance profile, which implies that closely related bacteria tend to occupy similar niches in the environment and respond in similar ways to environmental parameters. This in turn reflects that gene content, and thus functional traits, are strongly linked to phylogeny (Snel et al., 1999), even if this relationship to some extent is obscured by horizontal gene transfer (Ochman et al., 2000). Although communities may become phylogenetically clustered because habitat filtering selects for species with certain functional traits, interspecies competition may lead to phylogenetic dispersal by restricting all but the most successful species occupying a specific niche from surviving. For macroorganisms, both phylogenetic clustering (Webb, 2000) and dispersal (Slingsby and Verboom, 2006) have been observed, of which the latter seems to operate at a finer phylogenetic scale (Slingsby and Verboom, 2006). In addition, microbial communities have been shown to be phylogenetically clustered (for example, see Martin, 2002; Philippot et al., 2009). For example, mesocosms inoculated with the same starting community, but fed with different amounts of nitrogen and phosphorus, were more phylogenetically clustered than expected by chance, implying habitat filtering (Horner-Devine and Bohannan, 2006). Our study shows quantitative phylogenetic clustering in a community dynamics context. However, although closely related OTUs in our data set on average displayed more similar dynamics than distantly related ones, clear exceptions were observed (Figure 5), indicating adaptation to different niches among closely related OTUs. Fan-like phylogenetic structures have been observed in, for example, coastal marine microbial communities and in the human gut microbiota (Acinas et al., 2004; Ley et al., 2006). To what extent such fine-scale diversity reflects neutral diversity versus adaptation to different microniches has not yet been conclusively established (but see for example, Hunt et al., 2008). As read lengths of high-throughput sequencing methods increase, high-resolution time-series experiments have the potential to reveal to what extent different strains of phylogenetic fans co-vary, and hence shed light on to what extent the diversity is in fact neutral.
Accession codes
References
Acinas SG, Klepac-Ceraj V, Hunt DE, Pharino C, Ceraj I, Distel DL et al. (2004). Fine-scale phylogenetic architecture of a complex bacterial community. Nature 430: 551–554.
Allgaier M, Grossart HP . (2006). Diversity and seasonal dynamics of Actinobacteria populations in four lakes in northeastern Germany. Appl Environ Microbiol 72: 3489–3497.
Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z, Miller W et al. (1997). Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25: 3389–3402.
Andersson AF, Lindberg M, Jakobsson H, Backhed F, Nyren P, Engstrand L . (2008). Comparative analysis of human gut microbiota by barcoded pyrosequencing. PLoS ONE 3: e2836.
Berg KA, Lyra C, Sivonen K, Paulin L, Suomalainen S, Tuomi P et al. (2009). High diversity of cultivable heterotrophic bacteria in association with cyanobacterial water blooms. ISME J 3: 314–325.
Bruno WJ, Socci ND, Halpern AL . (2000). Weighted neighbor joining: a likelihood-based approach to distance-based phylogeny reconstruction. Mol Biol Evol 17: 189–197.
Clarke KR, Ainsworth M . (1993). A method of linking multivariate community structure to environmental variables. Mar Ecol Prog Ser 92: 205–219.
Cole JR, Wang Q, Cardenas E, Fish J, Chai B, Farris RJ et al. (2009). The Ribosomal Database Project: improved alignments and new tools for rRNA analysis. Nucleic Acids Res 37 (Database issue): D141–D145.
Cotner JB, Ammerman JW, Peele ER, Bentzen E . (1997). Phosphorus-limited bacterioplankton growth in the Sargasso Sea. Aquat Microb Ecol 13: 141–149.
Crump BC, Hobbie JE . (2005). Synchrony and seasonality in bacterioplankton communities of two temperate rivers. Limnol Oceanogr 50: 1718–1729.
Crump BC, Hopkinson CS, Sogin ML, Hobbie JE . (2004). Microbial biogeography along an estuarine salinity gradient: combined influences of bacterial growth and residence time. Appl Environ Microbiol 70: 1494–1505.
Eiler A, Bertilsson S . (2004). Composition of freshwater bacterial communities associated with cyanobacterial blooms in four Swedish lakes. Environ Microbiol 6: 1228–1243.
Fandino LB, Riemann L, Steward GF, Azam F . (2005). Population dynamics of Cytophaga-Flavobacteria during marine phytoplankton blooms analyzed by real-time quantitative PCR. Aquat Microb Ecol 40: 251–257.
Fuhrman JA, Hewson I, Schwalbach MS, Steele JA, Brown MV, Naeem S . (2006). Annually reoccurring bacterial communities are predictable from ocean conditions. Proc Natl Acad Sci USA 103: 13104–13109.
Glöckner FO, Zaichikov E, Belkova N, Denissova L, Pernthaler J, Pernthaler A et al. (2000). Comparative 16S rRNA analysis of lake bacterioplankton reveals globally distributed phylogenetic clusters including an abundant group of actinobacteria. Appl Environ Microbiol 66: 5053–5065.
Hagström Å, Larsson U . (1984). Diel and seasonal variation in growth rates of pelagic bacteria. In: Hobbie JE, Williams PJL (eds). Heterotrophic Activity in the Sea. Plenum Publishing Corp: NY, USA. pp 249–262.
Haverkamp TH, Schouten D, Doeleman M, Wollenzien U, Huisman J, Stal LJ . (2009). Colorful microdiversity of Synechococcus strains (picocyanobacteria) isolated from the Baltic Sea. ISME J 3: 397–408.
Holmfeldt K, Dziallas C, Titelman J, Pohlmann K, Grossart H-P, Riemann L . (2009). Regulation of diverse and abundant freshwater Actinobacteria along environmental gradients in the brackish northern Baltic Sea. Environ Microbiol 11: 2042–2054.
Horner-Devine MC, Bohannan BJ . (2006). Phylogenetic clustering and overdispersion in bacterial communities. Ecology 87 (7 Suppl): S100–S108.
Huber JA, Mark Welch DB, Morrison HG, Huse SM, Neal PR, Butterfield DA et al. (2007). Microbial population structures in the deep marine biosphere. Science 318: 97–100.
Hunt DE, David LA, Gevers D, Preheim SP, Alm EJ, Polz MF . (2008). Resource partitioning and sympatric differentiation among closely related bacterioplankton. Science 320: 1081–1085.
Huse SM, Huber JA, Morrison HG, Sogin ML, Welch DM . (2007). Accuracy and quality of massively parallel DNA pyrosequencing. Genome Biol 8: R143.
Kan J, Crump BC, Wang K, Chen F . (2006). Bacterioplankton community in Chesapeake Bay: predictable or random assemblages. Limnol Oceanogr 51: 2157–2169.
Kent AD, Yannarell AC, Rusak JA, Triplett EW, McMahon KD . (2007). Synchrony in aquatic microbial community dynamics. ISME J 1: 38–47.
Kisand V, Andersson N, Wikner J . (2005). Bacterial freshwater species successfully immigrate to the brackish water environment in the northern Baltic. Limnol Oceanogr 50: 945–956.
Labrenz M, Jost G, Jürgens K . (2007). Distribution of abundant prokaryotic organisms in the water column of the central Baltic Sea with an oxic-anoxic interface. Aquat Microb Ecol 46: 177–190.
Ley RE, Peterson DA, Gordon JI . (2006). Ecological and evolutionary forces shaping microbial diversity in the human intestine. Cell 124: 837–848.
MacQueen JB (ed). (1967). Some methods for classification and analysis of multivariate observations. Proceedings of the 5th Berkeley Symposium on Mathematical Statistics and Probability. University of California Press, Berkeley, CA, USA.
Margulies M, Egholm M, Altman WE, Attiya S, Bader JS, Bemben LA et al. (2005). Genome sequencing in microfabricated high-density picolitre reactors. Nature 437: 376–380.
Martin AP . (2002). Phylogenetic approaches for describing and comparing the diversity of microbial communities. Appl Environ Microbiol 68: 3673–3682.
Morris RM, Vergin KL, Cho JC, Rappe MS, Carlson CA, Giovannoni SJ . (2005). Temporal and spatial response of bacterioplankton lineages to annual convective overturn at the Bermuda Atlantic Time-series Study site. Limnol Oceanogr 50: 1687–1696.
Ochman H, Lawrence JG, Groisman EA . (2000). Lateral gene transfer and the nature of bacterial innovation. Nature 405: 299–304.
Pedrós-Alio C . (2006). Marine microbial diversity: can it be determined? Trends Microbiol 14: 257–263.
Pernthaler J, Glöckner F-O, Unterholzner S, Alfreider A, Psenner R, Amann R . (1998). Seasonal community and population dynamics of pelagic bacteria and archaea in a high mountain lake. Appl Environ Microbiol 64: 4299–4306.
Philippot L, Bru D, Saby NP, Cuhel J, Arrouays D, Simek M et al. (2009). Spatial patterns of bacterial taxa in nature reflect ecological traits of deep branches of the 16S rRNA bacterial tree. Environ Microbiol (in press; advance online publication, doi:10.1111/j.1462-2920.2009.02014.x).
Pinhassi J, Hagström Å . (2000). Seasonal succession in marine bacterioplankton. Aquat Microb Ecol 21: 245–256.
Pinhassi J, Sala MM, Havskum H, Peters F, Guadayol O, Malits A et al. (2004). Changes in bacterioplankton composition under different phytoplankton regimens. Appl Environ Microbiol 70: 6753–6766.
Reynolds CS . (1984). The Ecology of Freshwater Phytoplankton. Cambridge University Press: Cambridge, UK.
Riemann L, Leitet C, Pommier T, Simu K, Holmfeldt K, Larsson U et al. (2008). The native bacterioplankton community in the central Baltic Sea is influenced by freshwater bacterial species. Appl Environ Microbiol 74: 503–515.
Riemann L, Steward GF, Azam F . (2000). Dynamics of bacterial community composition and activity during a mesocosm diatom bloom. Appl Environ Microbiol 66: 578–587.
Riemann L, Winding A . (2001). Community dynamics of free-living and particle-associated bacterial assemblages during a freshwater phytoplankton bloom. Microb Ecol 42: 274–285.
Rönnberg C, Bonsdorff E . (2004). Baltic Sea eutrophication: area-specific ecological consequences. Hydrobiologia 514: 227–241.
Schauer M, Balagué V, Pedrós-Alió C, Massana R . (2003). Seasonal changes in the taxonomic composition of bacterioplankton in a coastal oligotrophic system. Aquat Microb Ecol 31: 163–174.
Selje N, Simon M . (2003). Composition and dynamics of particle-associated and free-living bacterial communities in the Weser estuary, Germany. Aquat Microb Ecol 30: 221–237.
Shade A, Kent AD, Jones SE, Newton RJ, Triplett EW, McMahon KD . (2007). Interannual dynamics and phenology of bacterial communities in a eutrophic lake. Limnol Oceanogr 52: 487–494.
Shiah F-K, Ducklow W . (1994). Temperature regulation of heterotrophic bacterioplankton abundance, production, and specific growth rate in Chesapeake Bay. Limnol Oceanogr 39: 1243–1258.
Slingsby JA, Verboom GA . (2006). Phylogenetic relatedness limits co-occurrence at fine spatial scales: evidence from the schoenoid sedges (Cyperaceae: Schoeneae) of the Cape Floristic Region, South Africa. Am Nat 168: 14–27.
Snel B, Bork P, Huynen MA . (1999). Genome phylogeny based on gene content. Nat Genet 21: 108–110.
Sogin ML, Morrison HG, Huber JA, Mark Welch D, Huse SM, Neal PR et al. (2006). Microbial diversity in the deep sea and the underexplored ‘rare biosphere’. Proc Natl Acad Sci USA 103: 12115–12120.
Stal LJ, Albertano P, Bergman B, Bröckel K, Gallon JR, Hayes PK et al. (2003). BASIC: Baltic Sea cyanobacteria. An investigation of the structure and dynamics of water blooms of cyanobacteria in the Baltic Sea—responses to a changing environment. Cont Shelf Res 23: 1695–1714.
Stigebrandt A . (2001). Physical oceanography of the Baltic Sea. In: Wulff F, Rahm L, Larsson P (eds). A Systems Analysis of the Baltic Sea. Springer Verlag: Heidelberg, Germany, pp 19–74.
Stoeck T, Epstein S . (2009). Crystal ball—2009: protists and the rare biosphere. Environ Microbiol Rep 1: 20–22.
Troussellier M, Schäfer H, Batailler N, Bernard L, Courties C, Lebaron P et al. (2002). Bacterial activity and genetic richness along an estuarine gradient (Rhone River plume, France). Aquat Microb Ecol 28: 13–24.
Warnecke F, Amann R, Pernthaler J . (2004). Actinobacterial 16S rRNA genes from freshwater habitats cluster in four distinct lineages. Environ Microbiol 6: 242–253.
Webb CO . (2000). Exploring the phylogenetic structure of ecological communities: an example for rain forest trees. Am Nat 156: 145–155.
Wulff F, Stigebrandt A . (1989). A time-dependent budget model for nutrients in the Baltic Sea. Glob Biogeochem Cycles 3: 63–78.
Zwart G, van Hannen EJ, Kamst-van Agterveld MP, Van der Gucht K, Lindstrom ES, Van Wichelen J et al. (2003). Rapid screening for freshwater bacterial groups by using reverse line blot hybridization. Appl Environ Microbiol 69: 5875–5883.
Zweifel UL, Norrman B, Hagström Å . (1993). Consumption of dissolved organic matter by marine bacteria and demand for inorganic nutrients. Mar Ecol Prog Ser 101: 23–32.
Acknowledgements
This study was supported by the Swedish Research Council (grants to SB and LR) and the Carl Trygger Foundation (grant to AFA). We thank Ulf Larsson, Leif Lundgren and Berndt Abrahamsson, Department of Systems Ecology, Stockholm University, for help with environmental data and sampling. Pyrosequencing was provided by the International Census of Marine Microbes (ICoMM) with financial support from a WM Keck foundation award to the Marine Biological Laboratory at Woods Hole. We thank the entire ICoMM team for their support during the completion of the study.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Supplementary Information accompanies the paper on The ISME Journal website (http://www.nature.com/ismej)
Supplementary information
Rights and permissions
About this article
Cite this article
Andersson, A., Riemann, L. & Bertilsson, S. Pyrosequencing reveals contrasting seasonal dynamics of taxa within Baltic Sea bacterioplankton communities. ISME J 4, 171–181 (2010). https://doi.org/10.1038/ismej.2009.108
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2009.108