Abstract
Microorganisms are strongly influenced by the bottom-up effects of resource supply. While many species respond to fluctuations in the concentration of resources, microbial diversity may also be affected by the heterogeneity of the resource pool, which often reflects a mixture of distinct molecules. To test this hypothesis, we examined resource–diversity relationships for bacterioplankton in a set of north temperate lakes that varied in their concentration and composition of dissolved organic matter (DOM), which is an important resource for heterotrophic bacteria. Using 16S rRNA transcript sequencing and ecosystem metabolomics, we documented strong relationships between bacterial alpha-diversity (richness and evenness) and the bulk concentration and the number of molecules in the DOM pool. Similarly, bacterial community beta-diversity was related to both DOM concentration and composition. However, in some lakes the relative abundance of resource generalists, which was inversely related to the DOM concentration, may have reduced the effect of DOM heterogeneity on community composition. Together, our results demonstrate the potential metabolic interactions between bacteria and organic matter and suggest that changes in organic matter composition may alter the structure and function of bacterial communities.
Similar content being viewed by others
Introduction
Resource supply has strong bottom-up effects on the diversity of consumer communities. Theory suggests that resource enrichment can promote diversity and food-web complexity [1,2,3,4]. Studies have shown that, in the absence of top-down control (i.e., predation), ecosystems with higher resource concentrations support more diverse and productive plant, algal, and microbial communities [5,6,7,8,9]. However, the relationship between resources and diversity often seems idiosyncratic [10,11,12]. Diversity can increase linearly with resource concentration [13], but it can also exhibit more complex, non-linear relationships where diversity peaks at intermediate concentrations [14]. Such patterns have been attributed to a range of processes including variation in competitive ability among consumers [14], shared limitations across species [13], and trophic interactions [15, 16].
Another feature that may influence patterns of biodiversity is the heterogeneity of the resource pool. Although it is convenient to model resources as homogenous pools, in reality, many resources exist as mixtures of multiple forms [17,18,19]. Often overlooked, resource heterogeneity has the potential to promote consumer diversity via niche partitioning [19, 20], including in microbial communities. For example, some aquatic bacteria are capable of specializing on different forms of carbon [21,22,23,24], and changes in the available carbon resources can restructure aquatic bacterial communities [25, 26]. Such differences in substrate preference contribute to diversity-maintaining mechanisms like niche partitioning, which influences the stability and functioning of microbial communities [20, 24, 27,28,29].
Ultimately, the effects of resource heterogeneity on biodiversity may depend on the degree to which bacterial communities are comprised of generalists or specialists. If communities are dominated by resource generalists, then the total concentration of a resource should have a stronger influence on diversity than the heterogeneity within the resource pool because resources will be essentially substitutable [13]. These generalist, or multivorous, bacteria should outperform specialists because they are able to simultaneously consume and metabolize numerous organic molecules [30]. In contrast, if communities are made up of resource specialists, then resource heterogeneity may promote consumer diversity by providing unique resource niches for consumers to partition [31, 32]. As such, resource heterogeneity and resource acquisition strategy (i.e., generalist versus specialist) may help resolve commonly observed variation in resource-diversity relationships.
For heterotrophic bacteria, organic matter is an important resource that is used for growth and physiological maintenance. However, organic matter is a heterogeneous collection of molecules that differ in chemical structure, origin, and age [33,34,35,36,37]. In aquatic ecosystems, dissolved organic matter (DOM) has traditionally been classified based on origin (autochthonous vs. allochthonous) and bioavailability (labile vs. recalcitrant). DOM can also be characterized based on its optical properties [35, 38] and functional groups (e.g., humic acids) [39]. However, other chemical features, including molecular weight, oxidation state, stoichiometry, and chemical structure, can influence the metabolism of organisms that consume DOM [40,41,42,43]. Recent technological advances, including Fourier transform ion cyclotron resonance mass spectrometry [44,45,46] and high-resolution liquid chromatography tandem mass spectrometry (MS) [47, 48], have made it possible to more thoroughly characterize DOM diversity at the molecular level and link various chemical properties to the structure of bacterial communities [37, 46,47,48]. With such tools in hand, we are now able to shed new light on the importance of fine-scale resource heterogeneity for resource–diversity relationships in bacterial communities.
In this study, we tested how resource heterogeneity contributes to resource–diversity relationships by characterizing aquatic bacterial communities and DOM. We characterized aquatic bacterial community diversity using 16S rRNA transcript sequencing, and we characterized the heterogeneity of the DOM pool using ecosystem metabolomics. We used linear models to test for relationships between bacterial alpha-diversity and DOM, and we used multivariate models to test for relationships between bacterial beta-diversity and DOM. To determine the role of resource acquisition strategy, we used species-resource (i.e., operational taxonomic unit (OTU)–DOM) co-occurrence to test whether resource–diversity relationships were influenced by the degree to which bacterial communities were dominated by specialists or generalists.
Methods
Study system and sampling
The Huron Mountains nature preserve is a 5300 ha tract of private land in the upper peninsula of Michigan, USA (Fig. 1). The area is part of the Superior Bedrock Uplands region [49]. The surrounding forests are primarily old-growth hemlock-northern hardwoods [50], and the inland water bodies are part of the Pine River Watershed, which drains into Lake Superior. Using a van Dorn sampler, we obtained surface water samples (0.5 m) from 10 lakes in the Huron Mountains during July 2012 (Fig. S1, Table 1). In addition, we measured dissolved oxygen concentrations, temperature, pH, and conductivity at the time of sampling using Quanta Hydrolab water quality sonde, and we measured chlorophyll a concentration in the lab after cold ethanol extraction of 0.7 µm-filtered (Whatman GF/F) water samples using a Turner Biosystems Fluorometer (Table 1).
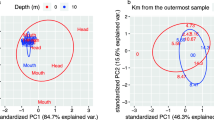
Principal coordinates analysis (PCoA) ordination of dissolved organic matter (DOM). The distances between symbols represent the dissimilarity between DOM in each lake. Using three axes, we can explain 71% of the variation in DOM composition. The third axis (not shown) captures 14% of the variation. Symbol sizes reflect variation in the concentration of dissolved organic carbon (DOC). Vectors represent the correlations between DOM composition and various physical and chemical attributes of each lake including: pH, elevation (elev), area, DOC, total nitrogen (TN), total phosphorus (TP), and chlorophyll a (Chl). Solid arrows represent significant and dotted represent nonsignificant correlations. Significance based on 10,000 permutations
Resource concentrations
With the water samples, we measured the concentrations of dissolved organic carbon (DOC), total nitrogen (TN), and total phosphorus (TP). We measured DOC concentrations by oxidation and non-dispersive infrared detection on 0.7 µm-filtered (Whatman, GF/F) samples using a Shimadzu TOC-V carbon analyzer. We measured TN on unfiltered samples using a Lachat FIA 8500 auto-analyzer (Hach, Loveland CO) after ammonium peroxydisulfate/sulfuric acid digestion [51]. We measured TP on unfiltered samples spectrophotometrically using the ammonium molybdate method and oxidation by persulfate digestion [52].
Resource heterogeneity
To estimate resource heterogeneity, we characterized the composition of extractable DOM for each lake using ecosystem metabolomics [53,54,55]. We extracted DOM from each sample using solid phase extraction (SPE), which is able to recover 40–60% of the total DOM based on chemical sorption properties [56]. Briefly, we acidified 1 L of 0.7 µm-filtered (Whatman, GF/F) water to pH 3.0 with 4 N HCl. We then passed the water sample through an SPE cartridge (Discovery-18, Supelco, Bellefonte PA) at a flow rate ≤ 5 mL min−1 using vacuum pressure. Columns were preconditioned using 6 mL 100% methanol followed by 6 mL pH 3.0 ultra-pure H2O. We filtered the sample until no sample remained or until the cartridge became clogged (recording the final volume filtered) and dried the filter with N2 gas for 5 min. We eluted the DOM from the column using 100% methanol and evaporated the methanol at 25 °C using vacuum centrifugation. A consistent amount of purified DOM was then separated on Waters Acquity ultra-performance liquid chromatography T3 column (1.8 µM, 1.0 × 100 mm) using water with a 0.1% formic acid-acetonitrile gradient and analyzed using negative electrospray ionization with quadrupole time of flight TOF MS (Waters G2 Q-TOF) and indiscriminate tandem MS (idMS/MS) at the Colorado State University Proteomics and Metabolomics Facility. Q-TOF MS provides high resolution, accurate mass quantification and idMS/MS provides high collision energy fragmentation without precursor ion selection acquired concurrently with low-collision energy MS data. For each sample, raw data files were converted to cdf format, and a matrix of molecular features as defined by retention time and ion mass (m/z) was generated for feature detection and alignment using the XCMS R package [57]. Raw peak areas were normalized to total ion signal, and the mean area of the chromatographic peak was calculated from duplicate injections. Features were grouped using the RAMClust R package, which groups features into spectra based co-elution and covariance across the full dataset, whereby spectra are used to determine the identity of observed compounds in the experiment [58]. Features were annotated using RAMClust. Briefly, molecular weight was inferred using by calling the interpretMSSpectrum R package [59]. Chemical annotations were assigned using the RAMClust annotate function, and structural annotations were imported using MSFinder [60]. Synonyms were retrieved using the RAMClust get.synonyms function, and when needed structures were retrieved using the RAMClust getSmilesInchi function. Last, compounds were assigned to chemical ontogeny using ClassyFire [61]. Further annotation details and parameters can be found in the Supplemental. We used field-prepared ultrapure water as controls and subtracted control peaks from sample peak heights. We multiplied control peaks by 1.1 to provide conservative blank subtraction. We define “DOM components” as the chemical features identified in DOM samples by ecosystem metabolomics. We used principal coordinates analysis (PCoA) to compare the composition of DOM in each lake and we used the envfit function in the vegan R package to determine which physical and chemical properties distinguished lakes based on their DOM composition [62]. The envfit function tests for correlations between the environmental variables and the PCoA axes, and it uses a permutation test to determine significance.
Composition of the active bacterial community
We used an RNA based approach to characterize the composition of the active bacterial community by sequencing the 16 S rRNA gene transcript. We extracted total nucleic acids using the MoBio Power Water RNA extraction kit (Carlsbad, CA). Nucleic acid extracts were cleaned via ethanol precipitation and RNA extracts were treated with DNase I (Invitrogen) to degrade residual DNA. We synthesized cDNA via the SuperScript III First Strand Synthesis Kit using random hexamer primers (Invitrogen). Once cDNA samples were cleaned and quantified, we amplified the 16S rRNA gene transcript (cDNA) using barcoded primers (515F and 806R) designed to work with the Illumina MiSeq platform [63]. We purified the sequence libraries using the AMPure XP purification kit, quantified using the QuantIt PicoGreen kit (Invitrogen), and pooled libraries at equal molar ratios (final concentration: 20 ng per). After pooling, we sequenced the libraries on the Illumina MiSeq platform using 250 × 250 bp paired end reads (Illumina Reagent Kit v2) at the Indiana University Center for Genomics and Bioinformatics Sequencing Facility. Paired-end raw 16S rRNA sequence reads were assembled into contigs and filtered based on quality score, length, and ambiguous base calls. After filtering, we aligned our sequences to the Silva Database (version 123). Chimeric sequences were detected and removed using the VSEARCH algorithm [64]. We then created OTUs by first splitting the sequences based on taxonomic class (using the RDP taxonomy) and then binning sequences in OTUs based on 97% sequence similarity using the OptiClust algorithm [65]. All initial sequence processing was completed using the software package mothur (version 1.40.5 [66]).
Resource heterogeneity and community diversity
First, we tested the hypothesis that resource heterogeneity affects bacterial community alpha-diversity. We used linear models to determine if DOM concentration or the number of DOM components would affect the richness and evenness of bacterial communities. We removed singleton OTUs (i.e., those found only once across all samples), and we subsampled using rarefaction to correct for differences in sample size due to sequencing depth [67, 68]. We calculated richness as the number of OTUs observed and evenness using Simpson’s evenness [69]. We transformed (Box–Cox), centered, and scaled (i.e., divided by standard deviation) DOC concentration and OTU richness to meet model assumptions regarding the distribution of residuals (see Supplemental [70]). We used the Box–Cox-transformed DOC concentration as the measure of DOM concentration, and we calculated the number of DOM components as the number of distinct DOM peaks observed in each sample.
Next, we tested the hypothesis that DOM heterogeneity affects community beta-diversity by comparing DOM concentrations and composition to bacterial community composition. We used distance-based redundancy analysis (dbRDA [71]) to test for relationships between: (1) resource concentration and community composition and (2) resource composition and community composition. dbRDA is a multivariate linear model technique that uses quantitative factors explaining differences in multivariate community composition data. We used the Box–Cox-transformed DOC concentration as the measure of DOM concentration. To use DOM composition as a predictor in our dbRDA model, we used PCoA, based on relative abundances and Bray–Curtis dissimilarity, to decompose DOM composition into orthogonal linear components [72]. To represent the DOM composition, we used the DOM PCoA axis scores for each sample. As the response in the dbRDA model, we relativized OTU abundances and used Bray–Curtis distances to compare community composition across samples. Significance tests of our dbRDA model were conducted based on 10,000 permutations. All calculations were done in the R statistical environment [73] using the vegan package [62].
Consumer–resource specialization
We tested the hypothesis that the response to DOM heterogeneity depends on whether bacterial communities were dominated by generalists or specialists by using consumer–resource (i.e., OTU–DOM) co-occurrence to define generalists and specialists. We defined resource generalists and specialists based on co-occurrence analysis, which was performed using Spearman’s rank correlations between DOM components and bacterial OTUs. We used the relative abundances of DOM components and the relative transcript abundances of bacterial OTUs. We inferred interactions based on correlations with coefficients >|0.7| [74], and we tested for significance using a permutation test based on randomizations with the independent-swap algorithm [75]. We defined resource generalists as those taxa with four or more significant negative resource correlations. While this cannot be used to directly attribute consumption or production of DOM components, we are using these correlations to infer potential metabolic associations. To understand the spatial extent of individual taxa, we defined cosmopolitan taxa as those found in ≥90% of the sampled lakes and we determined how many resource generalists were also cosmopolitan taxa. All calculations were done in the R statistical environment.
Results
Resource composition and heterogeneity
The lakes sampled in our survey captured a range of resource conditions that could affect patterns of bacterial diversity (Table 1). For example, the concentrations of DOC ranged from 4.22 to 28.99 mg C L−1 and were highly correlated with TN (rho = 0.97, p < 0.001, Fig. S2). Using ecosystem metabolomics, we characterized the DOM pool and detected 712 DOM components across the surveyed lakes. Based on the relative concentrations of DOM components, sites were on average 37% dissimilar in DOM composition. Using PCoA, the model explained 71% of the variation in DOM composition across sites using three dimensions (Fig. 1). The variation in DOM composition, based on the PCoA, was significantly related to DOC (r2 = 0.68, p = 0.01), TN (r2 = 0.70, p = 0.01), Chl a (r2 = 0.69, p = 0.02), and pH (r2 = 0.58, p = 0.03), but there were no significant relationships with TP (r2 = 0.27, p = 0.34), altitude (r2 = 0.31, p = 0.74), or surface area (r2 = 0.30, p = 0.26) (Fig. 1). In addition, we found an inverse relationship between the number of DOM components and the concentration of DOC (p < 0.01). We used DOC to represent resource concentration and the DOM PCoA scores to represent DOM composition in further analyses. We identified influential DOM components as those correlated (rho > |0.70|) with variation in the DOM PCoA axes (Fig. S3). These 172 influential components represent a broad range in molecular weight, stoichiometric ratio, and oxidation state (see Table 2 for a summary and Table S1 for a complete description).
Community composition and resource–diversity relationships
Across the sampled lakes, we identified 5085 bacterial OTUs based on 16S rRNA transcript sequencing (see Supplement for sequence counts). When rarified (60,000 sequences), lakes varied in OTU richness and evenness (Fig. 2). Using Bray–Curtis distances and relative transcript abundances, lakes were on average 62% dissimilar based on bacterial community composition.
Bacterial community diversity relationships with resource concentration and the number of DOM components. Resource (DOM) concentration and OTU richness have been Box–Cox transformed to meet model assumptions. There are significant positive relationships between OTU richness and resource concentration and between OTU evenness and the number of DOM components. There are significant negative relationships between OTU evenness and resource concentration and between OTU richness and the number of DOM components. Dashed line represents linear regression fit along with 95% confidence intervals. An untransformed version of this figure can be found in the supplemental (Fig. S4)
First, we tested for relationships between DOM chemistry and bacterial alpha-diversity. We used linear regression to test for resource–diversity relationships between bacterial community diversity (richness and evenness) and both DOM concentration and the number of DOM components in a sample. As predicted, bacterial alpha-diversity was affected by resource concentration (Fig. 2). OTU richness was positively related to DOM concentration (r2 = 0.66, p = 0.008), but inversely related to the number of DOM components (r2 = 0.50, p = 0.023). In contrast, OTU evenness was positively related to the number of DOM components (r2 = 0.67, p = 0.003) but inversely related to DOM concentration (r2 = 0.49, p = 0.022). We found a similar relationship when we characterized bacterial communities using 16S rRNA gene sequences (see Supplemental).
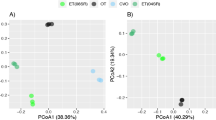
Next, we tested for relationships between DOM (concentration and composition) and bacterial beta-diversity using dbRDA. Based on the dbRDA models, DOM concentration explained 28% of the variation in bacterial community composition (p = 0.002), and DOM composition explained 45% of the variation in bacterial community composition (p = 0.03, Fig. 3). We found a similar relationship when we characterized bacterial communities using 16S rRNA gene sequences (see Supplemental). However, when we partitioned variation among the DOM PCoA axes, only DOM Axis 2 was significant (r2 = 0.70, p = 0.017). In addition, this DOM PCoA Axis 2 (DOM 2) was correlated to variation along OTU PCoA Axis 1 (rho = 0.83, p = 0.002; Fig. 3). Last, we tested for relationships between DOM concentration and DOM composition. We found a significant correlation between DOM concentration and DOM Axis 2 (rho = 0.69, p = 0.03).
Principal coordinates analysis (PCoA) ordination of bacterial communities. Vectors represent the correlation between the dissolved organic matter (DOM) heterogeneity and the bacterial community composition. The two vectors are based on correlations between community composition and the site scores from the DOM PCoA axes one and two. We used distance-based redundancy analysis to test the relationship between DOM site scores and bacterial community composition
Consumer–resource specialization
Based on consumer–resource co-occurrence analysis and spatial occurrence, we classified generalist bacteria. We found that 1.3% of taxa (68 OTUs) were resource generalists. In addition, using spatial occurrence we determined that 4.6% of the taxa (233 OTUs) were cosmopolitan. Of the resource generalists, 73.5% (50 OTUs) were also found to be cosmopolitan taxa (Table 3). Proportionally, resource generalists and cosmopolitan taxa were relatively abundant across all lakes (Fig. 4). For both groups, there was a significant inverse relationship between relative abundance and resource concentration (Fig. 4). In addition, the relative abundance of resource generalists was related to DOM composition based on DOM Axis 2 (rho = 0.81, p = 0.004), but not DOM Axis 1 (rho = 0.08, p = 0.82). Taxonomically, both resource generalists and cosmopolitan taxa were diverse. For the resource generalists, the majority belonged to the classes Alphaproteobacteria (14) and Planctomycetacia (11), but Verrucomicrobiae (8) and Actinobacteria (7) were also common. At the family level, the resource generalists represented groups, including Acetobacteraceae, Caulobacteraceae, Planctomycetaceae, Sphinomonadaceae, and Verrucomicrobiaceae. For the cosmopolitan taxa, the majority belonged to the classes Alphaproteobacteria (58) and Betaproteobacteria (50), but Gammaproteobacteria (14), Actinobacteria (21), Planctomycetacia (17), and Sphingobacteria (12) were also common. At the family level, the cosmopolitan taxa represent groups including Acetobacteraceaea, Alcaligenaceae, Bulkholderiaceae, Caulobacteraceae, Chitinophagaceae, Comomonadaceae, Flavobacteriaceae, Planctomycetaceae, Rhodobacteraceae, Spartobacteria, Sphinomonadaceae, and Verrucomicrobiaceae. For a complete list of generalists including taxonomy, see Tables S2 and S3.
The proportion of generalists and cosmopolitan taxa in aquatic bacterial communities. We defined operational taxonomic units (OTUs) as generalists using consumer–resource co-occurrence (top) and as cosmopolitan based on spatial occurrence (top). We used OTU relative abundances to calculate the proportion in each community. For both, we used a linear model to determine if there was a relationship between the proportion of generalists and the concentration of dissolved organic carbon (DOC). For both, we found a significant negative relationship. Dashed line represents linear regression fit along with 95% confidence intervals. The light gray dotted line represents 50% of the community and is used as a reference
Discussion
We found evidence that DOM concentration and heterogeneity affect aquatic bacterial resource-diversity relationships. We documented significant relationships between DOM and bacterial community alpha- and beta-diversity. Together, our results suggest that, in addition to bulk concentration, the heterogeneity of DOM resources influences aquatic bacterial diversity within and among communities. Our study supports the view that organic matter composition plays an important role in structuring aquatic bacterial communities, and that changes in organic matter composition owing to land-use modifications and changing terrestrial inputs may alter the structure and function of aquatic bacterial communities.
Resource heterogeneity affects bacterial diversity
This study revealed that DOM resources were heterogeneous across lakes—on average lakes were 37% dissimilar in their DOM composition; and therefore, resource heterogeneity may help explain the variation in resource–diversity relationships along resource concentration gradients. We found that while DOM concentration explained 28% of the variation, DOM composition explained 45% of the variation in bacterial community composition across lakes. These findings suggest that different types of bacteria use and potentially specialize on different types of DOM [26, 27, 46]. For example, it has been shown that some bacteria primarily use algal-derived resources [76, 77] while others primarily use terrestrial-derived resources [78, 79]. Therefore, lakes receiving different resource inputs may be expected to contain different bacterial communities. Thus, DOM resource heterogeneity is a potential mechanism to explain the diversity of bacteria within and among lake communities.
One of the strongest relationships we observed was the positive correlation between OTU evenness and the number of DOM components (Fig. 2). The number of DOM components is likely to influence the evenness of the bacterial community because evenness, a measure of equitability among taxa, may reflect changes in community assembly by modifying the relationship between resource availability, species interactions, and species traits [80, 81]. If DOM resources represented niches to be partitioned, the number of DOM components should promote species diversity since DOM components provide unique niches for species to partition [19, 31, 82]. Because greater resource heterogeneity was associated with increased evenness but not richness, our findings suggest that the increased evenness of the bacterial community represents changes in abundances but not the addition of new taxa. Together, our results support the hypothesis that DOM resource heterogeneity contributes to observed resource–diversity relationships, and we propose that DOM resource heterogeneity may promote more diverse heterotrophic bacterial communities by increasing species equitability.
Resource substitutability
The substitutability of resources could influence strength and direction of resource–diversity relationships. One possible explanation for why resource heterogeneity may only have weak effects in some habitats is that many resources are substitutable. Two resources are substitutable when either can each be used for growth and reproduction while the other is absent [83]. For example, some phototrophs are able to use ammonium, nitrate, or organic nitrogen as a source of nitrogen [84,85,86], while crustacean zooplankton such as Daphnia can use algae, cyanobacteria, or bacteria as different food sources [87]. Likewise, aquatic ecosystems contain numerous phosphorus resources but some have similar effects on the structure and function of aquatic bacterial communities [29].
We found numerous DOM components that appear to have similar consumer–resource co-occurrence patterns (Fig. S5). One explanation for this pattern is that many DOM components represent substitutable resources. At a chemical level, resources with the same core structure may be enzymatically substitutable. For example, vanillate and ferulate share an internal benzene structure and are used by the same metabolic pathway (beta-ketoadipate pathway: [88]). In addition, extracellular enzymes often degrade aliphatic polymers of different lengths into identical monomers [89]. Alternatively, multivorous bacteria with numerous metabolic pathways may use different molecules in similar ways if they contain the similar molecular components, are easily transported, and yield the same amount of energy [30]. As such, our data indicated that many DOM components are likely substitutable. While we were able to provide chemical annotations for most of the DOM components in our study, it is still unclear how to determine which components may be substitutable. This does not mean relationships between DOM and bacterial composition are indescribable, but methods need to be developed to classify and group DOM components into meaningful categories based on functional and metabolic forms.
Generalist communities
Our results suggest that resource generalists can be common in aquatic bacterial communities [90,91,92] and may explain, in part, why the effect of resource heterogeneity on community composition is stronger is some lakes than others. Across the sampled lakes, we documented an inverse relationship between the abundance of generalists and the concentrations of resources. This relationship was correlated with variation along the second axis of the DOM PCoA, but not the first DOM PCoA axis which explains the majority of the DOM variation. One possibility is that consumers have multiple metabolic pathways for resource acquisition. For example, evidence from comparative genomics suggests that aquatic bacteria capable of using complex organic matter also have the potential to use numerous different resources, and may thus be generalists [93,94,95].
It is often assumed that most bacteria are specialists. For example, multiple studies have identified taxa that specialize on particular resources [26, 76, 96,97,98]. The ability to use multiple resources requires the production of extra enzymes and transporters; therefore, it is costly to use numerous resources [99]. As such, specialists may be energetically favored in some environments. Likewise, numerous studies have indicated that habitat specialists (e.g., sediment and aquatic) dominate microbial communities [100,101,102,103]. However, this study suggests that generalists were relatively common in the lakes surveyed (Fig. 4). These findings are supported by another study which found that resource generalists dominated coastal bacterial communities [92]. Likewise, it has been proposed that generalist, multivorous bacteria may actually have a fitness advantages if resources are essentially substitutable [30]. It should be noted though, that we found both resource generalists and specialists (Fig. S5) and resource specialists would contribute to resource-diversity relationships. However, generalists may limit the ability of resource heterogeneity to promote diversity when generalists are more dominant than specialists.
While advances in technology have allowed for in-depth characterization of organic matter chemistry and bacterial communities, there are still limitations. First, the DOM extraction and detection may be biased toward some groups of molecules [56]. While we may have missed some important components of the DOM pool (e.g., hydrophilic compounds), we likely captured the complex terrestrial-derived organic matter that often dominates inland aquatic ecosystems such as small ponds and lakes [104]. This DOM has been shown to be important for bacterial community structure and function [42, 105, 106]. However, our methods may have missed less complex labile molecules (e.g., sugars and free amino acids) that have also been shown to affect bacterial communities [77]. Many labile molecules are consumed rapidly and may therefore escape detection. Second, our consumer-resource interaction results are based on a single time point and therefore only suggest possible bacteria-DOM metabolic interactions. We use these correlations to make inferences about the degree to which taxa are generalists. To make stronger inferences, we would need to conduct time-course experiments capturing periods of resource fluctuations often associated with hydrologic events, and we would need to perform experimental manipulations of DOM concentration and composition. In addition, we are using 16S rRNA sequencing to characterize community composition, but this method may not be able to capture cryptic diversity and strain specialization [107, 108]. For example, our results identified members of the class Actinobacteria as generalist, but most Actinobacteria have been identified as specialists [103, 109] and this discrepancy may be due to our phylogenetic resolution [110]. Last, we assume that bacterial communities are under local selection due to resource availability, but other local (e.g., predation and the physical environment) and regional (e.g., dispersal) factors can affect community composition [111,112,113,114,115,116]. For example, high-dispersal rates can overwhelm local selection due to mass effects [117], which is especially important in aquatic bacterial communities that receive organisms from the neighboring terrestrial landscape [113, 118]. Furthermore, small waterbodies, such as Lily and Pony, have higher terrestrial influence due to a high shoreline-area ratio. Terrestrial influence increases the input of bacteria to these small waterbodies and this can inflate bacterial diversity and therefore constrain the resource–diversity relationship. Regardless, our results, and other genomic studies, suggest that resource generalist are common in some aquatic bacterial communities, and this should be investigated further.
Conclusions
This study revealed that resource heterogeneity influenced resource–diversity relationships and the contribution of DOM heterogeneity to these relationships was greater than the contribution of DOM concentration. In addition, we found that generalist taxa were relatively common in many of the lakes that we sampled, but generalists were predicted to reduce effect of resource heterogeneity on the resource–diversity relationship. However, the presence of generalists does not limit the role of resource heterogeneity. First, because resource generalists do not use all resources and can still show preferences [119], resource variation can still have an effect on community composition. Second, in addition to generalists we also found evidence of resource specialists and others have found strong evidence for resource and habitat specialists in inland aquatic ecosystems [98, 100, 106]. Because the presence of generalist taxa may limit the affect resource heterogeneity has on local communities, we propose that consumer properties (i.e., generalist) and resource properties (i.e., availability) determine how strong communities respond to resource heterogeneity. Therefore, to better understand how bacterial communities will respond to environmental changes, such as temporal and spatial variation in organic matter inputs due to changes in plant community distributions or global climate change, we need to consider which resources are substitutable and which groups of resources will change in similar and predictive ways.
References
Abrams PA, Roth J. The responses of unstable food chains to enrichment. Evol Ecol. 1994;8:150–71.
Worm B, Lotze HK, Hillebrand H, Sommer U. Consumer versus resource control of species diversity and ecosystem functioning. Nature. 2002;417:848–51.
Hairston NGJ, Hairston NG. Cause-effect relationships in energy-flow, trophic structure, and interspecific interactions. Am Nat. 1993;142:379–411.
Polis GA, Strong DR. Food web complexity and community dynamics. Am Nat. 1996;147:813–46.
Waldrop MP, Zak DR, Blackwood CB, Curtis CD, Tilman D. Resource availability controls fungal diversity across a plant diversity gradient. Ecol Lett. 2006;9:1127–35.
Claire Horner-Devine M, Leibold Ma, Smith VH, Bohannan BJM. Bacterial diversity patterns along a gradient of primary productivity. Ecol Lett. 2003;6:613–22.
Hulot FD, Lacroix G, Lescher-Moutoué F, Loreau M. Functional diversity governs ecosystem response to nutrient enrichment. Nature. 2000;405:340–4.
Leibold MA, Chase JM, Shurin Jonathan B, Downing AL. Species turnover and the regulation of trophic structure. Annu Rev Ecol Syst. 1997;28:467–94.
Smith VH. Microbial diversity-productivity relationships in aquatic ecosystems. FEMS Microbiol Ecol. 2007;62:181–6.
Tilman D, Kilham SS, Kilham P. Phytoplankton community ecology: the role of limiting nutrients. Annu Rev Ecol Syst. 1982;13:349–72.
Mittelbach GG, Steiner CF, Scheiner SM, Gross KL, Reynolds HL, Waide RB, et al. What is the observed relationship between species richness and productivity? Ecology. 2001;82:2381–96.
Rosenzweig ML. Paradox of enrichment: destabilization of exploitation ecosystems in ecological time. Science. 1971;171:385–7.
Stevens MH, Carson WP. Resource quantity, not resource heterogeneity, maintains plant diversity. Ecol Lett. 2002;5:420–6.
Leibold MA. Biodiversity and nutrient enrichment in pond plankton communities. Evol Ecol Res. 1999;1:73–95.
Carpenter SR, Cole JJ, Hodgson JR, Kitchell JF, Pace ML, Bade DL. et al. Trophic cascades nutrients, and lake productivity: whole-lake experiments. Ecol Monogrpahs. 2001;71:163–86.
Holt RD, Grover JP, Tilman D. Simple rules for interspecific dominance in systems with exploitative and apparent competition. Am Nat. 1994;144:741–71.
Turner BL. Resource partitioning for soil phosphorus: a hypothesis. J Ecol. 2008;96:698–702.
Ashton IW, Miller AE, Bowman WD, Suding KN. Niche complementarity due to plasticity in resource use: plant partitioning of chemical N forms. Ecology. 2010;91:3252–60.
Schoener TW. Resource partitioning in ecological communities. Science. 1974;185:27–39.
Finke DL, Snyder WE. Niche partitioning increases resource exploitation by diverse communities. Science. 2008;321:1488–90.
Töpper B, Thingstad TF, Sandaa R-A. Effects of differences in organic supply on bacterial diversity subject to viral lysis. FEMS Microbiol Ecol. 2013;83:202–13.
Alonso-Saez L, Aristegui J, Pinhassi J, Gomez-Consarnau L, Gonzalez JM, Vaque D, et al. Bacterial assemblage structure and carbon metabolism along a productivity gradient in the NE Atlantic Ocean. Aquat Microb Ecol. 2007;46:43–53.
Horňák K, Kasalický V, Šimek K, Grossart HP. Strain-specific consumption and transformation of alga-derived dissolved organic matter by members of the limnohabitans-C and polynucleobacter-B clusters of betaproteobacteria. Environ Microbiol. 2017;19:4519–35.
Salcher MM, Posch T, Pernthaler J. In situ substrate preferences of abundant bacterioplankton populations in a prealpine freshwater lake. ISME J. 2013;7:896–907.
Alonso-Sáez L, Gasol JM. Seasonal variations in the contributions of different bacterial groups to the uptake of low-molecular-weight compounds in northwestern Mediterranean coastal waters. Appl Environ Microbiol. 2007;73:3528–35.
Gómez-Consarnau L, Lindh MV, Gasol JM, Pinhassi J. Structuring of bacterioplankton communities by specific dissolved organic carbon compounds. Environ Microbiol. 2012;14:2361–78.
Jones SE, Newton RJ, McMahon KD. Evidence for structuring of bacterial community composition by organic carbon source in temperate lakes. Environ Microbiol. 2009;11:2463–72.
Loreau M, Hector A. Partitioning selection and complementarity in biodiversity experiments. Nature. 2001;412:72–6.
Muscarella ME, Bird K, Larsen ML, Placella SA, Lennon JT. Phosphorus resource heterogeneity in microbial food webs. Aquat Microb Ecol. 2014;73:259–72.
Egli T. How to live at very low substrate concentration. Water Res. 2010;44:4826–37.
Glasser JWJ. Evolution of efficiencies and strategies of resource exploitation. Ecology. 1984;65:1570–8.
Levine JM, HilleRisLambers J. The importance of niches for the maintenance of species diversity. Nature. 2009;461:254–7.
Stevenson FJ. Humus chemistry: genesis, composition, reactions. 2nd ed. New York, NY: John Wiley & Sons, Inc.; 1984.
Dittmar T, Stubbins A (2014). Dissolved organic matter in aquatic systems. In: Holland HD, Turekian KK (editors) Vol. 1. Treatise on geochemistry. Elsevier, p. 125–56. ISBN: 978-0-08-098300-4, ebook, Oxford UK; San Diego, CA, USA.
Fellman JB, Miller MP, Cory RM, D’Amore DV, White D. Characterizing dissolved organic matter using PARAFAC modeling of fluorescence spectroscopy: a comparison of two models. Environ Sci Technol. 2009;43:6228–34.
Jaffé R, McKnight D, Maie N, Cory R, McDowell WH, Campbell JL. Spatial and temporal variations in DOM composition in ecosystems: the importance of long-term monitoring of optical properties. J Geophys Res. 2008;113:G04032.
Moran MA, Kujawinski EB, Stubbins A, Fatland R, Aluwihare LI, Buchan A, et al. Deciphering ocean carbon in a changing world. Proc Natl Acad Sci. 2016;113:3143–51.
Weishaar JL, Aiken GR, Bergamaschi B A, Fram MS, Fujii R, Mopper K. Evaluation of specific ultraviolet absorbance as an indicator of the chemical composition and reactivity of dissolved organic carbon. Environ Sci Technol. 2003;37:4702–8.
Croué J-P. Isolation of humic and non-humic NOM fractions: structural characterization. Environ Monit Assess. 2004;92:193–207.
Cherif M, Loreau M. Stoichiometric constraints on resource use, competitive interactions, and elemental cycling in microbial decomposers. Am Nat. 2007;169:709–24.
Cory RM, McKnight DM. Fluorescence spectroscopy reveals ubiquitous presence of oxidized and reduced quinones in dissolved organic matter. Environ Sci Technol. 2005;39:8142–9.
Lennon JT, Pfaff LE. Source and supply of terrestrial organic matter affects aquatic microbial metabolism. Aquat Microb Ecol. 2005;39:107–19.
Berggren M, Ström L, Laudon H, Karlsson J, Jonsson A, Giesler R, et al. Lake secondary production fueled by rapid transfer of low molecular weight organic carbon from terrestrial sources to aquatic consumers. Ecol Lett. 2010;13:870–80.
Martínez-Pérez AM, Osterholz H, Nieto-Cid M, Álvarez M, Dittmar T, Álvarez-Salgado XA. Molecular composition of dissolved organic matter in the Mediterranean Sea. Limnol Oceanogr. 2017;62:2699–712.
Osterholz H, Singer G, Wemheuer B, Daniel R, Simon M, Niggemann J, et al. Deciphering associations between dissolved organic molecules and bacterial communities in a pelagic marine system. ISME J. 2016;10:1717–30.
Osterholz H, Kirchman DL, Niggemann J, Dittmar T. Diversity of bacterial communities and dissolved organic matter in a temperate estuary. FEMS Microbiol Ecol. 2018;94:1–11.
Petras D, Koester I, Da Silva R, Stephens BM, Haas AF, Nelson CE, et al. High-resolution liquid chromatography tandem mass spectrometry enables large scale molecular characterization of dissolved organic matter. Front Mar Sci. 2017;4:405.
Broeckling CD, Broz AK, Bergelson J, Manter DK, Vivanco JM. Root exudates regulate soil fungal community composition and diversity. Appl Environ Microbiol. 2008;74:738–44.
Schaetzl RJ, Enander H, Luehmann MD, Lusch DP, Fish C, Bigsby M, et al. Mapping the physiography of Michigan with GIS. Phys Geogr. 2013;34:2–39.
Woods KD. Dynamics in late-successional hemlock-hardwood forests over three decades. Ecology. 2000;81:110–26.
Lachat. (2005). Standard operating procedure for total and total dissolved phosphorous (Lachat Method 10-115-01-1-F for QuikChem FIA + 8000).
Wetzel RG, Likens GE. Inorganic nutrients: nitrogen, phosphorus, and other nutrients. In: Limnological analyses. 3rd ed, (eds. Robert G. Wetzel and Gene E. Likens) pp. 85–111, (Springer-Verlag, New York, 2000).
Dias DA, Svendsen C, Jones OAH, Griffin JL, Maguire ML, Spurgeon DJ. Metabolomics and its use in ecology. Austral Ecol. 2013;38:713–20.
Fegel T, Boot CM, Baron JS, Hall EK. Inking the chemistry and reactivity of dissolved organic matter from glaciers and rock glaciers. bioRxiv. e-pub ahead of print, https://doi.org/10.1101/115808; 2017.
Raes J, Bork P. Molecular eco-systems biology: towards an understanding of community function. Nat Rev Microbiol. 2008;6:693–9.
Dittmar T, Koch B, Hertkorn N, Kattner G. A simple and efficient method for the solid-phase extraction of dissolved organic matter (SPE-DOM) from seawater. Limnol Oceanogr Methods. 2008;6:230–5.
Smith CA, Tautenhahn R, Neumann S, Benton P, Conley C, Rainer J (2016). xcms: LC/MS and GC/MS data analysis package.
Broeckling CD, Afsar FA, Neumann S, Prenni JE. RAMClust: a Novel feature clustering method enables spectral-matching-based annotation for metabolomics data. Anal Chem. 2014;86:6812–7.
Jaeger C, Méret M, Schmitt CA, Lisec J. Compound annotation in liquid chromatography/high-resolution mass spectrometry based metabolomics: robust adduct ion determination as a prerequisite to structure prediction in electrospray ionization mass spectra. Rapid Commun Mass Spectrom. 2017;31:1261–6.
Tsugawa H, Kind T, Nakabayashi R, Yukihira D, Tanaka W, Cajka T, et al. Hydrogen rearrangement rules: computational MS/MS fragmentation and structure elucidation using MS-FINDER software. Anal Chem. 2016;88:7946–58.
Djoumbou Feunang Y, Eisner R, Knox C, Chepelev L, Hastings J, Owen G, et al. ClassyFire: automated chemical classification with a comprehensive, computable taxonomy. J Cheminform. 2016;8:61.
Oksanen J, Blanchet FG, Kindt R, Legendre P, Minchin PR, O’Hara RB, et al. Vegan: community ecology package. http://cran.r-project.org/package=vegan; 2013.
Caporaso JG, Lauber CL, Walters WA, Berg-Lyons D, Huntley J, Fierer N, et al. Ultra-high-throughput microbial community analysis on the Illumina HiSeq and MiSeq platforms. ISME J. 2012;6:1621–4.
Rognes T, Flouri T, Nichols B, Quince C, Mahé F. VSEARCH: a versatile open source tool for metagenomics. PeerJ. 2016;4:e2584.
Westcott SL, Schloss PD. OptiClust, an improved method for assigning amplicon-based sequence data to operational taxonomic units. mSphere. 2017;2:e00073–17.
Schloss PD, Westcott SL, Ryabin T, Hall JR, Hartmann M, Hollister EB, et al. Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol. 2009;75:7537–41.
Hughes JB, Hellmann JJ. The application of rarefaction techniques to molecular inventories of microbial diversity. Methods Enzymol. 2005;397:292–308.
James FC, Rathbun S. Rarefaction, relative abundance, and diversity of avian communities. Auk. 1981;98:785–800.
Smith B, Wilson JB. A consumer’s guide to evenness indices. Oikos. 1996;76:70–82.
Neter J, Kutner MH, Nachtsheim CJ, Wasserman W. Applied linear statistical models. Chicago, Illinois, USA: Irwin; 1996.
Legendre P, Anderson MJ. Distance-based redundancy analysis: testing multispecies responses in multifactorial ecological experiments. Ecol Monogr. 1999;69:1–24.
Legendre P, Legendre LFJ. Numerical ecology. 3rd ed. (Elsevier, Oxford, UK, 2012).
R Core Team. R: A language and environment for statistical computing. http://www.r-project.org/; 2012.
Williams RJ, Howe A, Hofmockel KS. Demonstrating microbial co-occurrence pattern analyses within and between ecosystems. Front Microbiol. 2014;5:1–10.
Gotelli NJ. Null model analysis of species co-occurrence patterns. Ecology. 2000;81:2606–21.
Jaspers E, Overmann J. Ecological significance of microdiversity: identical 16S rRNA gene sequences can be found in bacteria with highly divergent genomes and ecophysiologies. Appl Environmantal Microbiol. 2004;70:4831–9.
Sarmento H, Gasol JM. Use of phytoplankton-derived dissolved organic carbon by different types of bacterioplankton. Environ Microbiol. 2012;14:2348–60.
Guillemette F, McCallister SL, del Giorgio PA. Selective consumption and metabolic allocation of terrestrial and algal carbon determine allochthony in lakes. ISME J. 2015;10:1373–82.
Roehm CL, Giesler R, Karlsson J. Bioavailability of terrestrial organic carbon to lake bacteria: the case of a degrading subarctic permafrost mire complex. J Geophys Res. 2009;114:G03006.
Hill MO. Diversity and evenness: a unifying notation and its consequences. Ecology. 1973;54:427–32.
Hillebrand H, Bennett DM, Cadotte MW. Consequences of dominance: a review of evenness effects on local and regional ecosystem processes. Ecology. 2008;89:2712–24.
Werner EE. Species packing and niche complementarity in three sunfishes. Am Nat. 1977;111:553–78.
Tilman D. Resources: a graphical-mechanistic approach to competition and predation. Am Nat. 1980;116:362–93.
Haynes RJ, Goh KM. Ammonium and nitrate nutrition of plants. Biol Rev. 1978;53:465–510.
McKane RB, Johnson LC, Shaver GR, Nadelhoffer KJ, Rastetter EB, Fry B, et al. Resource-based niches provide a basis for plant species diversity and dominance in arctic tundra. Nature. 2002;415:68–71.
Schimel JP, Bennett J. Nitrogen mineralization: challenges of a changing paradigm. Ecology. 2004;85:591–602.
Demott WR. Utilization of a cyanobacterium and a phosphorus-deficient green alga as complementary resources by daphnids. Ecology. 1998;79:2463–81.
Buchan A, Collier LS, Neidle EL. Key aromatic-ring-cleaving enzyme, protocatechuate 3,4-dioxygenase, in the ecologically important marine Roseobacter lineage. Appl Environ Microbiol. 2000;66:4662–72.
Rojo F. Degradation of alkanes by bacteria: minireview. Environ Microbiol. 2009;11:2477–90.
Sriswasdi S, Yang CC, Iwasaki W. Generalist species drive microbial dispersion and evolution. Nat Commun 2017;8:1162.
Barberán A, Ramirez KS, Leff JW, Bradford MA, Wall DH, Fierer N (2014). Why are some microbes more ubiquitous than others? Predicting the habitat breadth of soil bacteria. Ecol Lett. 17: 794–802.
Mou X, Sun S, Edwards RA, Hodson RE, Moran MA. Bacterial carbon processing by generalist species in the coastal ocean. Nature. 2008;451:708–11.
Lauro FM, McDougald D, Thomas T, Williams TJ, Egan S, Rice S, et al. The genomic basis of trophic strategy in marine bacteria. Proc Natl Acad Sci. 2009;106:15527–33.
Livermore JA, Emrich SJ, Tan J, Jones SE. Freshwater bacterial lifestyles inferred from comparative genomics. Environ Microbiol. 2014;16:746–58.
Newton RJ, Griffin LE, Bowles KM, Meile C, Gifford S, Givens CE, et al. Genome characteristics of a generalist marine bacterial lineage. ISME J. 2010;4:784–98.
Hunt DE, David LA, Gevers D, Preheim SP, Alm EJ, Polz MF. Resource partitioning and sympatric differentiation among closely related bacterioplankton. Science. 2008;320:1081–5.
Mccarren J, Becker JW, Repeta DJ, Shi Y, Young CR, Malmstrom RR, et al. Microbial community transcriptomes reveal microbes and metabolic pathways associated with dissolved organic matter turnover in the sea. Proc Natl Acad Sci. 2010;107:16420–7.
Bird K. Generalist and specialist strategies of phopsohrus acquisition by aquatic bacteria. East Lansing, MI: Michigan State University; 2012. Masters thesis.
Johnson DR, Goldschmidt F, Lilja EE, Ackermann M. Metabolic specialization and the assembly of microbial communities. ISME J. 2012;6:1985–91.
Székely AJ, Langenheder S. The importance of species sorting differs between habitat generalists and specialists in bacterial communities. FEMS Microbiol Ecol. 2014;87:102–12.
Langenheder S, Ragnarsson H. The role of environmental and spatial factors for the composition of aquatic bacterial communities. Ecology. 2007;88:2154–61.
Mariadassou M, Pichon S, Ebert D. Microbial ecosystems are dominated by specialist taxa. Ecol Lett. 2015;18:974–82.
Monard C, Gantner S, Bertilsson S, Hallin S, Stenlid J. Habitat generalists and specialists in microbial communities across a terrestrial-freshwater gradient. Sci Rep. 2016;6:37719.
Wilkinson GM, Pace ML, Cole JJ. Terrestrial dominance of organic matter in north temperate lakes. Global Biogeochem Cycles. 2013;27:43–51.
Lapierre J-F, Guillemette F, Berggren M, Del Giorgio PA. Increases in terrestrially derived carbon stimulate organic carbon processing and CO2 emissions in boreal aquatic ecosystems. Nat Commun. 2013;4:2972.
Muscarella ME, Jones SE, Lennon JT (2016) Species sorting along a subsidy gradient alters bacterial community stability. Ecology. 2016;97:2034–43.
Hahn MW, Jezberová J, Koll U, Saueressig-Beck T, Schmidt J. Complete ecological isolation and cryptic diversity in Polynucleobacter bacteria not resolved by 16S rRNA gene sequences. ISME J. 2016;10:1642–55.
Rodriguez-R LM, Castro JC, Kyrpides NC, Cole JR, Tiedje JM, Konstantinidis KT. How much do rRNA gene surveys underestimate extant bacterial diversity? Appl Environ Microbiol. 2018;84:00014-18.
Newton RJ, Jones SE, Eiler A, McMahon KD, Bertilsson S. A guide to the natural history of freshwater lake bacteria. Microbiol Mol Biol Rev. 2011;75:14–49.
Martiny AC, Treseder K, Pusch G. Phylogenetic conservatism of functional traits in microorganisms. ISME J. 2013;7:830–8.
Winter C, Bouvier T, Weinbauer MG, Thingstad TF. Trade-offs between competition and defense specialists among unicellular planktonic organisms: the ‘killing the winner’ hypothesis revisited. Microbiol Mol Biol Rev. 2010;74:42–57.
Zeglin LH. Stream microbial diversity in response to environmental changes: review and synthesis of existing research. Front Microbiol. 2015;6:454.
Crump BC, Amaral-Zettler LA, Kling GW. Microbial diversity in arctic freshwaters is structured by inoculation of microbes from soils. ISME J. 2012;6:1629–39.
Gasol JM, Pedrós-Alió C, Vaqué D. Regulation of bacterial assemblages in oligotrophic plankton systems: Results from experimental and emperical approaches. Antonie van Leeuwenhoek. 2002;81:435–52.
Langenheder S, Lindström ES. Local and regional factors influencing bacterial community assembly. Environ Microbiol Rep. 2012;4:1–9.
Rousk J, Bååth E, Brookes PC, Lauber CL, Lozupone C, Caporaso JG, et al. Soil bacterial and fungal communities across a pH gradient in an arable soil. ISME J. 2010;4:1340–51.
Leibold MA, Holyoak M, Mouquet N, Amarasekare P, Chase JM, Hoopes MF, et al. The metacommunity concept: a framework for multi-scale community ecology. Ecol Lett. 2004;7:601–13.
Ruiz-Gonzalez C, Nino-Garcia JP, del Giorgio PA. Terrestrial origin of bacterial communities in complex boreal freshwater networks. Ecol Lett. 2015;18:1198–206.
Behmer ST, Joern A. Coexisting generalist herbivores occupy unique nutritional feeding niches. Proc Natl Acad Sci. 2008;105:1977–82.
Acknowledgements
We acknowledge constructive feedback from AL Peralta, NI Wisnoski, and two anonymous reviewers. We thank BK Lehmkuhl for technical support and W Thorpe for logistical support. This work was supported by the Huron Mountain Wildlife Foundation (M.E.M. and J.T.L.), the National Science Foundation DEB-1442246 (J.T.L.) and DEB-1501164 (M.E.M. and J.T.L.), and US Army Research Office Grant W911NF-14-1-0411 (J.T.L.). All code and data used in this study can be found in a public GitHub repository (https://www.github.com/LennonLab/ResourceHeterogeneity; https://doi.org/10.5281/zenodo.2656098) and NCBI (BioProject PRJNA540276).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Muscarella, M.E., Boot, C.M., Broeckling, C.D. et al. Resource heterogeneity structures aquatic bacterial communities. ISME J 13, 2183–2195 (2019). https://doi.org/10.1038/s41396-019-0427-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-019-0427-7
This article is cited by
-
Contrasting Effects of Leaf Litter Quality and Diversity on Oviposition of Mosquitoes
Neotropical Entomology (2023)
-
Reduced chemodiversity suppresses rhizosphere microbiome functioning in the mono-cropped agroecosystems
Microbiome (2022)
-
Ecological networks of dissolved organic matter and microorganisms under global change
Nature Communications (2022)
-
Blood meal source and mixed blood-feeding influence gut bacterial community composition in Aedes aegypti
Parasites & Vectors (2021)
-
Non-additive microbial community responses to environmental complexity
Nature Communications (2021)