Abstract
Malassezia furfur was the first species described within the cosmopolitan yeast genus Malassezia, which now comprises 13 species. Reported isolation rates of these species from healthy and diseased human skin show geographic variations. PCR-fingerprinting with the wild-type phage M13 primer (5′-GAGGGTGGCGGTTCT-3′) was applied to investigate phylogeographic associations of M. furfur strains isolated from Scandinavians residing permanently in Greece, in comparison to clinical isolates from Greek, Bulgarian and Chinese native residents. Seven M. furfur strains from Scandinavians were compared with the Neotype strain (CBS1878), CBS global collection strains (n=10) and clinical isolates from Greece (n=4), Bulgaria (n=15) and China (n=6). Scandinavian, Greek and Bulgarian M. furfur strains mostly formed distinct group clusters, providing initial evidence for an association with the host's geographical origin and with the underlying skin condition. These initial data address the hypothesis that M. furfur could be a eukaryotic candidate eligible for phylogeographic studies.
Similar content being viewed by others
Main
The genus Malassezia includes 10 anthropophilic and obligatory lipophilic species (M. globosa, M. restricta, M. slooffiae, M. obtusa, M. furfur, M. sympodialis, M. japonica, M. yamatoensis, M. dermatis, M. nana) and 3 zoophilic species (M. pachydermatis, M. caprae, M. equina) (Cabañes et al., 2007; Ashbee, 2007). It is considered a member of the class Exobasidiomycetes that mainly includes plant pathogens (Begerow et al., 2006). Malassezia yeasts show global distribution on human skin and display adaptable, occasionally species-specific biological properties related to their human host. (Xu et al., 2007). Moreover, worldwide epidemiological studies sampling healthy volunteers and Malassezia associated dermatoses, such as pityriasis versicolor, atopic dermatitis and seborrheic dermatitis, report isolation of mixed species from healthy and diseased skin, as well as clear geographic variations in Malassezia species isolation rates (Ashbee, 2007).
In this study PCR-fingerprinting, a widely available method offering direct multi-centre assessment of primary epidemiological data, was used to investigate potential phylogeographic associations of M. furfur strains isolated from Scandinavians permanently residing in Greece, as well as clinical isolates from Greece, Bulgaria and China. M. furfur strains formed distinct group clusters, associated with the host's geographical origin and with the underlying skin condition.
Seven M. furfur strains isolated from the forehead (A) or midscapula area (B) of Scandinavian volunteers residing permanently in Greece for 0.5–47 years (median 16) were compared with the Neotype (CBS 1878) (Centraalbureau voor Schimmelcultures, Utrecht, Netherlands), global strains (n=10) and 25 clinical isolates from Greece (n=4), Bulgaria (n=15) and China (n=6; Table 1). For isolation and identification to species level, Malassezia strains were grown on modified Dixon's agar at 32 °C (Gaitanis et al., 2006a, 2006b). M. furfur strains were identified by their inability to grow on Sabouraud-dextrose agar without lipid supplementation, the positive assimilation of Tweens 20, 40, 60, 80 and Cremophor El (Guého et al, 1996; Mayser et al, 1997). Conventional identification was confirmed by PCR–RFLP analysis of the internal transcribed spacer 2 region (Gaitanis et al 2006a, 2006b). DNA was extracted by a modified hexadecyltrimethylammonium bromide method (Velegraki et al, 1999). Additional automated DNA extractions (Maxwell 16, Promega, Madison, WI, USA) were carried out from single, 2–3 mm diameter M. furfur colonies from Dixon's agar, using a modification (not shown) of the Maxwell 16 Cell DNA Purification protocol (Promega), yielding 20 ng DNA (A260/A280: 1.03). Automated M. furfur extraction was engaged to evaluate its standardization aptitude and throughput in upcoming medium-scale intercontinental population studies, its basic downstream applications, such as direct use in PCR, restriction enzyme digestion and agarose gel electrophoresis, and downstream real-time PCR research applications.
For PCR fingerprinting, the minisatellite-specific core sequence of the wild-type phage M13 (5′-GAGGGTGGCGGTTCT-3′) (Meyer and Mitchell, 1995) was used. The 25 μl of the PCR master mix contained 2 μl MgCl2 (25 mM), 2.5 μl MgCl2-free storage buffer B (Promega), 1.5U Taq Polymerase (Promega), 0.015 mM of each dNTP (HT Biotechnology Ltd, Cambridge, UK), 50 pmol of the M13 primer and 200–250 ng of template DNA. The PCR reaction was carried out in a Techgene Techne thermal cycler (Stone Staffordshire, UK) and consisted of 1 initial denaturation cycle (5 min; 94 °C) and 45 amplification cycles (1 min; 94 °C, 1 min; 42 °C and 2 min; 72 °C). PCR products were electrophoresed in 4–6% 29:1 acrylimide/bis-acrylimide stacking gels for 3.5 h at 180 V, followed by ethidium bromide staining and UV visualization (Herolab, E.A.S.Y., Weisloch, Germany). The M13 PCR-fingerprinting has been exhaustively validated previously in the basidiomycetous yeast Cryptococcus neoformans (Meyer and Mitchell, 1995). In this study, the stability of M13 fingerprints for all M. furfur strains was confirmed in five independent experiments within a 3-year period.
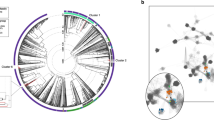
The dendrogram was constructed by the Bionumerics software Version 4 (Bio-Maths, Bilthoven, Belgium; Figure 1) using the Dice Coefficient of similarity and cluster analysis with the unweighted pair-group method with arithmetic averages (UPGMA) using 1.00% position tolerance and no optimization, to obtain the greatest variation in similarity. A cutoff point of 24% similarity was used discriminating six major groups.
Rooted dendogram constructed using unweighted pair-group method with arithmetic averages (UPMGA) analysis of M. furfur microsatellite PCR fingerprints as generated by Bionumerics software. B: Bulgarian M. furfur strains; CH: Chinese; GS: Scandinavians residing in Greece; SD: seborrheic dermatitis; PV: pityriasis versicolor; HS: healthy subjects; CBS: Centraalbureau voor Schimmelcultures, Utrecht, the Netherlands. In case there was a simultaneous isolation of two M. furfur strains from the forehead and midscapula area they were differentiated as A and B, respectively.
Overall, M. furfur fingerprints showed a trend to associate with the host's geographic origin and a correlation with the underlying skin condition (Figure 1). Six out of seven Scandinavian M. furfur isolates formed Group IV with 35% homology forming a distinct cluster from Groups V and VI (20% homology) that included all Greek and the global selection of strains (Figure 1). The single M. furfur Scandinavian (GS1A) isolate that was grouped together with the Bulgarian isolates in Group I (37% similarity; Figure 1) could represent a transient member (Mayser et al., 2001) of the Malassezia skin flora.
Generally, data on skin Malassezia population dynamics are scant (Paulino et al., 2006) and not directly comparable. Yet, indicative of the same phylogeographic trend is that a M. furfur seborrheic dermatitis isolate from a Dutch male residing for 5 years in China showed identical complete internal transcribed spacer sequences with M. furfur CBS7982, from a European healthy individual (Ran et al., 2008).
Bulgarian strains clustered in Groups I and II with clear disease associations. Specifically, seborrheic dermatitis isolates (n=10) clustered in Group I with 51% similarity and pityriasis versicolor (n=3) and dandruff (DF) (n=2) isolates in Group II (similarity 77%). Although the number of isolates analyzed is small, the observed disease-specific M. furfur clustering concurs with previously recorded disease-specific Malassezia species subtypes (Paulino et al, 2006; Tajima et al, 2008) based on IGS, internal transcribed spacer and 18S sequencing. Interestingly, the two systemic M. furfur isolates from France and Greece (Table 1) clustered together showing their common European and disease origin (Gupta et al, 2004). Of the Chinese isolates, four out of six formed a single Group III with 22% homology to Group I, whereas two out of six grouped separately. More strains from this large geographic region should be analyzed to reveal consequential human phylogeographic segregation. Variable M. furfur subtypes (Figure 1: strains GS4A/GS4B and GS9A/GS9B) from different anatomical regions showed different M13 fingerprints. As multiple Malassezia species are isolated from individuals (Gaitanis et al., 2006b; Tajima et al, 2008), an analogous intra-species polymorphism is expected.
The undemanding sampling procedure and the satisfactory M. furfur isolation rate of 5–23.3% (Sweden, Sandström-Falk et al., 2005; Iran, Tarazooie et al., 2004) may appoint M. furfur a candidate eukaryotic organism for studying human phylogeography. Moreover, compared with other Malassezia species, M. furfur survives longer in storage, thus supporting retrospective studies, strain exchange and inter-laboratory investigations. Also, M. furfur is designated as a Biosafety Level 2 agent, this not impending international handling and distribution (Devlin, 2006).
Generally, PCR-fingerprinting holds an inborn methodological disadvantage, because the randomly amplified DNA fragments are used in a deterministic way to identify similarity or dissimilarity of isolates. By contrast, the minisatellite-derived oligonucleotide M13 is capable of achieving specific amplification of inter-repeat sequences by target minisatellite sequences dispersed throughout the fungal genome. Despite the somewhat inscrutable role of minisatellites in genomic plasticity on an evolutionary time scale, M13 PCR-fingerprinting used as a single-typing method in yeasts has been proven, upon stringent PCR reaction parameter standardization, highly reproducible. Since it is long shown to successfully detect inter- and intra-species variation (Meyer and Mitchell, 1995), it represents a cost-effective initial screening method engaging M.furfur in human phylogeography studies.
To our knowledge, this is the first time that a eukaryote serves as a potential marker for human phylogeography. Human viruses (HIV, dengue virus, rabbies virus) (Holmes, 2004) and bacteria (Helicobacter pylori) (Falush et al., 2003) have been used previously for this purpose. By contrast, other human fungal pathogens of global distribution, such as Trichophyton rubrum, though shown clonally reproduced in two geographically distinct populations, (Gräser et al., 2007), maintain genetic homogeneity (Summerbell et al., 1999) rendering it unsuitable for this purpose.
The data presented herein have to be confirmed with inclusion of additional M. furfur isolates from more geographically defined population cohorts using robust typing methods, such as multilocus sequencing, to expand knowledge on Malassezia species ecology and pathobiology and further assess its candidacy for phylogeograpic studies.
References
Ashbee HR . (2007). Update on the genus Malassezia. Med Mycol 45: 287–303.
Begerow D, Stoll M, Baurer R . (2006). A phylogenetic hypothesis of Ustilagomycotina based on multiple gene analyses and morphological data. Mycologia 98: 906–916.
Cabañes FJ, Theelen B, Castellá, Boekhout T . (2007). Two new lipid-dependent Malassezia species from domestic animals. FEMS Yeast Res 7: 1064–1076.
Devlin RK . (2006). Invasive fungal infections caused by Candida and Malassezia species in the neonatal intensive care unit. Adv Neonatal Care 6: 68–77.
Falush D, Wirth T, Linz B, Pritchard J, Stephens M, Kidd M et al. (2003). Traces of human migrations in Helicobacter pylori populations. Science 299: 1582–1585.
Gaitanis G, Robert V, Velegraki A . (2006a). Verifiable single nucleotide polymorphisms of the internal transcribed spacer 2 region for the identification of 11 Malassezia species. J Derm Sc 43: 214–217.
Gaitanis G, Velegraki A, Alexopoulos EC, Chasapi V, Tsigonia A, Katsambas A . (2006b). Distribution of Malassezia species in pityriasis versicolor and seborrhoeic dermatitis in Greece. Typing of the major pityriasis versicolor isolate M. globosa. Br J Dermatol 154: 854–859.
Gräser Y, Fröhlich J, Presber W, de Hoog S. (2007). Microsatellite markers reveal geographic population differentiation in Trichophyton rubrum. J Med Microbiol 56: 1058–1065.
Guého E, Midgley G, Guillot J . (1996). The genus Malassezia with description of four new species. Antonie Van Leeuwenhoek 69: 337–355.
Gupta AK, Boekhout T, Theelen B, Summerbell R, Batra R . (2004). Identification and typing of Malassezia species by amplified fragment length polymorphism and sequence analyses of the internal transcribed spacer and large-subunit regions of ribosomal DNA. J Clin Microbiol 42: 4253–4260.
Holmes EC . (2004). The phylogeography of human viruses. Mol Ecol 13: 745–756.
Mayser P, Schütz M, Schuppe HC, Jung A, Schill WB . (2001). Frequency and spectrum of Malassezia yeasts in the area of the prepuce and glans penis. BJU Int 88: 554–558.
Mayser P, Haze P, Papavassilis C, Pickel M, Gruender K, Guého E . (1997). Differentiation of Malassezia species: selectivity of cremophor EL, castor oil and ricinoleic acid for M furfur. Br J Dermatol 137: 208–213.
Meyer W, Mitchell T . (1995). Polymerase chain reaction fingerprinting in fungi using single primers specific to minisatellites and simple repetitive DNA sequences: strains variation in Cryptococcus neoformans. Electrophoresis 16: 1648–1656.
Paulino LC, Tseng C-H, Strober BE, Blaser M . (2006). Molecular analysis of fungal microbiota in samples from healthy human skin and psoriatic lesions. J Clin Microbiol 44: 2933–2941.
Ran Y, He X, Zhang H, Dai Y, Li L, Bulmer GS . (2008). Seborrheic dermatitis flare in a Dutch male due to commensal Malassezia furfur overgrowth. Med Mycol 46: 611–614.
Sandström Falk MH, Tengvall Linder M, Johansson C M-H, Bartosik J, Bäck O, Tarrnhult T et al. (2005). The prevalence of Malassezia yeasts in patients with atopic dermatis, seborrheic dermatitis and healthy controls. Acta Derm-Venereol 85: 17–23.
Summerbell RC, Haugland RA, Li A, Gupta AK . (1999). rRNA gene internal transcribed spacer 1 and 2 gene sequences of asexual, anthropophilic dermatophytes related to Trichophyton rubrum. J Clin Microbiol 37: 4005–4011.
Tajima M, Sugita T, Nishikawa A, Tsuboi R . (2008). Molecular analysis of Malassezia microflora in seborrheic dermatitis patients: comparison with other diseases and healthy subjects. J Invest Dermatol 128: 345–351.
Tarazooie B, Kordbacheh P, Zaini F, Zomoridian K, Saadat F, Zeraati H et al. (2004). Study of the distribution of Malassezia species in patients with pityriasis versicolor and healthy individuals in Tehran, Iran. BMC Dermatol 4: 5.
Velegraki A, Kambouris M, Kostourou A, Chalevelakis G, Legakis NJ . (1999). Rapid extraction of fungal DNA from clinical samples for PCR amplification Med. Mycol 37: 69–73.
Xu J, Saunders CW, Hu P, Grant RA, Boekhout T, Kuramae EE et al. (2007). Dandruff-associated Malassezia genomes reveal convergent and divergent virulence traits shared with plant and human fungal pathogens. Proc Natl Acad Sci USA 104: 18730–18735.
Acknowledgements
This study was encouraged by the International Society for Human and Animal Mycology (ISHAM) Malassezia Working Group and funded by the Greek State Scholarships Foundation (IKY) and the University of Athens Grant ‘Kapodistrias’ K.A. 70/4/5905 and K.A 70/3/6915. We thank the Scandinavian community of Athens, the NOKIA Hellas personnel and Helena Boundouvi for organizing the sampling sessions, G Tzanakaki and K Kessanopoulos, National Reference Laboratory for Meningococcal Disease, School of Public Health, Athens, Greece, for use of the Bionumerics Software and S Kritikou for technical assistance.
Author information
Authors and Affiliations
Corresponding author
Additional information
Conflict of interest
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Gaitanis, G., Velegraki, A., Alexopoulos, E. et al. Malassezia furfur fingerprints as possible markers for human phylogeography. ISME J 3, 498–502 (2009). https://doi.org/10.1038/ismej.2008.112
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2008.112
Keywords
This article is cited by
-
Delimiting species in Basidiomycota: a review
Fungal Diversity (2021)
-
Secreted Hydrolytic and Haemolytic Activities of Malassezia Clinical Strains
Mycopathologia (2019)