Abstract
To elucidate fungal diversity in methane hydrate-bearing deep-sea marine sediments in the South China Sea, internal transcribed spacer (ITS) regions of rRNA genes from five different sediment DNA samples were amplified and phylogenetically analyzed. Total five ITS libraries were constructed and 413 clones selected randomly were grouped into 24 restriction patterns by Amplified Ribosomal DNA Restriction Analysis (ARDRA). ITS sequences of 44 representative clones were determined and compared with the GenBank database using gapped-BLAST. The phylogenetic analysis showed that the ITS sequences (71–97% similarity) were similar to those of Phoma, Lodderomyces, Malassezia, Cryptococcus, Cylindrocarpon, Hortaea, Pichia, Aspergillus and Candida. The remaining sequences were not associated to any known fungi or fungal sequences in the public database. The results suggested that methane hydrate-bearing deep-sea marine sediments harbor diverse fungi. This is the first report on fungal communities from methane hydrate-bearing deep-sea marine sediments in South China Sea.
Similar content being viewed by others
Introduction
Marine fungi are ubiquitous in the ocean and have been isolated from a variety of detritus environments (Rappé et al., 1998; Buchan et al., 2002). Many marine fungi could tolerate and adapt to extreme conditions like prokaryotes and some fungi have been isolated from the deep-sea mud (Takami et al., 1997; Nagahama et al., 2003). Deep-sea sedimentary environments have occurred continuously throughout the Earth's history and have the potential to harbor diverse novel sequences that may be relevant to the early evolution of eukaryotes. However, current models of deep-sea eukaryotic evolution have relied on comparisons among a few microbes, reconstructing the phylogenetic relations from such ecological environments is necessary for us to understand better the origin and evolution of early eukaryotes on this planet.
Methane hydrates are typically found in marine sediments on continental margins or hundreds of meters below the ground in circumpolar permafrost regions where low temperature and high pressure favor the formation of hydrates (MacDonald et al., 1994; Sassen et al., 1999). Methane is predominantly of biological origin in marine sediments that are frequently rich in organic matter. Some novel archaeal and bacterial clones have been observed in hydrate-bearing sediments (Reed et al., 2002). Small eukaryotes in deep sea seems to be involved in decomposition of organic matter, which may play important roles in flows of energy in deep-sea biosphere. However, there are few investigations of the fungal communities associated with methane hydrate-bearing sediments.
Although diverse eukaryotic microbes were found in deep-sea sediments by cultivation and microscopy, the abundance and diversity of naturally occurring fungi have not been well characterized. Most microbes resisted cultivation, and microscopic descriptions do not discriminate similar organisms that are physiologically different. Culture-independent methods based on the rRNA genes of microbes were usually used to study the microbial communities in natural habitats. The internal transcribed spacer (ITS) regions of fungal rRNA genes have been identified as discriminative targets for molecular analysis of fungal communities and their high sequence variability relative to the flanking sequences makes them valuable for genus- and species-level identification (Buchan et al., 2002). To describe more fully fungal diversity in methane hydrate-bearing deep-sea marine sediments and find new sequences useful for phylogenetic studies, fungal ITS regions of rRNA genes from five different sediment DNA samples were amplified and phylogenetically analyzed.
Materials and methods
Sediment samples
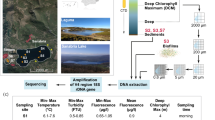
Sediment samples for DNA extraction were collected from the subsurface sediments at water depths of 350 m (A), 884 m (C), 1123 m (D), 2965 m (F) and 3011 m (G) in South China Sea by the Department of Guangzhou Marine Geological Survey. These samples were considered to be associated with gas hydrate based on the signatures of bottom simulating reflector (BSR), amplitude blanking zone with high velocity above BSR, drilling results and hydrate data (Zhang et al., 2002; Liu et al., 2005). The samples were frozen at −20°C before being processed. To minimize contamination, the superficial parts exposed to the air were discarded when processed. DNA extractions were conducted under sterile conditions and the manipulation of each sample was performed separately to avoid cross contamination.
DNA extraction
DNA extraction was performed using combined physical and chemical treatments (Miller, 2001; Gabor et al., 2003; Robe et al., 2003). Two grams (wet weight) of sediments was added to 10 ml polypropylene tubes. Then 3.6 ml of the extraction buffer (0.25 M NaH2PO4, 0.1 M Tris-HCl (pH 8.0), 0.1 M EDTA (pH 8.0), 1% hexadecylmethylammonium bromide (CTAB)), 0.4 ml of 20% SDS solution and equal amounts of glass beads (250–400 mesh) were added. The mixture was homogenized for 1 min using a Vortex, frozen in liquid nitrogen for 1 min and incubated at 68°C for 1 h by inverting the tube at intervals of 15 min. The tube was centrifuged at 13 000 g for 15 min, and then the supernatant was transferred into a new 10-ml tube. Potassium acetate (5 M, 300 μl) and 40% polyethylene glycol 8000 (1 ml) solutions were added, precipitated at –20°C for 15 min and centrifuged at 13 000 g for 15 min. The pellets were dissolved in 1 ml 2 × CTAB (2% CTAB, 1.4 M NaCl, 0.1 M EDTA, pH 8.0) solution and incubated at 68°C for 15 min. One milliliter of chloroform was added and the solution was gently mixed, then centrifuged at 13 000 g for 10 min. Aqueous DNA was precipitated by 1 ml isopropanol at –20°C for 15 min. After centrifugation at 13 000 g for 15 min, supernatant was removed carefully and the pellet was washed with 1 ml of 70% ethanol. The DNA was resolved in 100 μl of TE buffer for further studies. The DNA of each sample was extracted in duplicate.
ITS amplification
The ITS region was amplified using fungal-specific ITS 1F (5′-CTTGGTCATTTAGAGGAAGTAA-3′) and universal ITS 4 (5′-TCCTCCGCTTATTGATATGC-3′) primers (White et al., 1990; Gardes and Bruns, 1993). The PCR products are about 600 bp long, including ITS 1, 5.8 S and ITS 2 regions of the rRNA operon. Approximately 10–100 ng of template DNA, 2.5 μl 10 × PCR buffer (500 mM KCl, 100 mM Tris-HCl (pH 9.0), 15 mM 1%, (w/v) MgCl2, Triton X-100), 0.2 U Taq DNA polymerase (Takara Biotechnology (Dalian) Co., Ltd., Dalian, China), 1 μ M ITS 1F primer, 1 μ M ITS 4 primer and 200 μ M dNTPs were combined in a total volume of 25 μl. The reaction mixtures lacking template DNA in parallel were performed as negative controls. PCR was carried out with PTC-100 cyclers (MJ Research, Inc., Waltham, MA, USA). The initial 5 min at 94°C was followed by 30 cycles of 50 s at 94°C, 50 s at 55°C and 1.5 min at 72°C. A final step of 10 min at 72°C was included to complete any partial polymerizations. All the PCR procedures were repeated once. PCR products were purified using EZNA Cycle-Pure Kits (Omega Bio-tek Inc., Doraville, GA, USA).
Construction of ITS libraries and ARDRA analyses
Purified PCR products were cloned into the pGEM-T easy vector (Promega Corporation, Madison, WI, USA), and then transformed into Escherichia coli DH5α according to the manufacturer's instructions. Clones were screened according to α-complementation on Luria–Bertani agar medium containing 100 μg ml−1 ampicillin, 5-bromo-4-chloro-3-indolyl-β-D-galactopyranoside (X-gal) (800 μg ml−1) and isopropyl β-D-thiogalactopyranoside (IPTG) (800 μg ml−1). To identify unique ITS, the inserted DNA sequence was amplified in reaction mixtures described above using T7 and SP6 primers flanking the multiclone sites of the vector. The PCR conditions were presented as follows: the initial 5 min at 94°C was followed by 30 cycles of 30 s at 94°C, 30 s at 54°C and 90 s at 72°C, final extension at 72°C for 10 min. Amplified products were selected according their size in 1% (w/v) agarose gels. The resulting products were further used for restriction analysis by the restriction enzymes HaeIII and TaqI, respectively. Digested products were size-fractioned on 2% agarose gels for 1.5 h at 5 V cm−1 and grouped according to their restriction patterns. Clones with identical patterns were considered as the same operational taxonomic unit (OTU).
Accumulation curves and diversity analysis
Library coverage was calculated using the relative distribution of OTUs and the equation described previously (Good, 1953). Rarefaction curves were drawn using the freeware program EcoSim 7.0 (Gotelli and Entsminger, 2005). The Shannon–Weaver index of general diversity (H′) was calculated for each library using the equation H′=−∑Pi ln Pi. Pi was calculated as follows: Pi=ni/N, where ni is the number of clones belonging to the same OTU pattern and N is the total number of library clones (Shannon and Weaver, 1949).
Sequencing and phylogenetic analysis
One to four clones of each OTU were selected randomly for sequencing, using the Applied Biosystems 377 sequencer in Invitrogen Biotechnology Co. (Guangzhou, China). All sequences were analyzed using the rRNA Database Project CHECK_CHIMERA program to remove chimeric sequences (Maidak et al., 2001). The retained sequences were compared with the sequences of GenBank database using gapped-BLAST (Altschul et al., 1997). Multiple alignments were obtained using the CLUSTALX 1.80 program. Phylogenetic relationships were inferred by the neighbor-joining method. Evolutionary distances were calculated with the Kimura two-parameter model (transition/transversion ratio=2.0). To avoid potential biases, ordering of taxa was randomized.
Nucleotide sequence accession numbers
The GenBank accession numbers of the nucleotide sequences determined in this study are presented in Table 2.
Results
ITS libraries and ARDRA analysis
Total 413 positive clones of the five ITS libraries (designated as A, C, D, G and E) were analyzed by ARDRA analysis (Table 1). Sequence divergence was not found among randomly selected 10 pairs of clones that were considered as one ARDRA pattern. The results indicated that the clones belonged to one single pattern. Therefore, one OTU pattern was considered as a single and unique OTU. Library coverage ranged from 76 to 95%. Sample A showed the highest coverage but the lowest number of OTU patterns. Rarefaction curve of Sample A showed a plateau, which indicated that library A covered almost phylotypes (Figure 1). Sample A also contained the least diversity of fungi. Sample D represented the highest OTU patterns but the lowest coverage value. Rarefaction curve of Sample D did not reach a plateau. The fungal diversity of Sample D was not assessed. The other three samples showed high coverage values (⩾0.83) and decreasing slopes of accumulation curves. No samples showed high diversity of ITS sequences. Calculation of H′ indicated different fungal diversities among the five samples. The OTU patterns also revealed genetic variability of fungi in methane hydrate-bearing sediments (Table 2). For example, each sample contained different OTU patterns, and only the OTU1 and OTU3 were found from all the five samples (Table 1). The numbers of clones belonging to OTU1 and OTU3 accounted for over 75% of total clones from Samples A, C, F and G, and 55% from Sample D.
Phylogenic analysis
Forty-four ITS clones representing total 24 OTU patterns were sequenced and compared with the GenBank database (Figure 2). The composition indicated that most of the ITS sequences that can be included in the order of Ascomycota and Basidomycota originated from fungi. ITS sequences from seven OTU patterns (OTU1, OTU3, OTU6, OTU13, OTU18, OTU19 and OTU23) were most closely related to the cultivable fungi Phoma glomerata. Sequences from OTU1, OTU3, OTU13, OTU18 and OTU19 showed about 97% identity with Phoma glomerata. They represented 331 fungal clones and accounted for 80% of the total 413 clones. However, OTU6 and OTU23 showed low identity to the most related sequences in the GenBank (<83%). The two sequences formed a separate clade with 95% bootstrap percentages, so they were not associated to any currently documented sequences.
Neighbor-joining tree obtained from analysis of the rDNA ITS showing the relationships of the 24 representative sequences in five libraries. The tree was rooted by using two stramenopiles sequences (Ectocarpus siliculosus and Kuckuckia sp.). Bootstrap values (n=1000 replicates) of ⩾50% are reported as percentages. The scale bar represents the number of changes per nucleotide position. ITS, internal transcribed spacer.
Clones of OTU5 were phylogenetically associated with Cylindrocarpon species and fungal sp. R55 (AY699699). Clones of OTU14 were clustered with Hortaea werneckii and clones of OTU17 clustered with Cladosporium species. These three OTU patterns belonged to Pezizomycotina, which is a subphylum of the ascomycota. Clones of OTU10 and OTU16 were associated with genus Emericella and Aspergillus. Clones of OTU7 were distantly related to Aspergillus versicolor (76.83%), and apparently represented another type of heterotrophic divergence (98% of bootstrap value).
There are diverse yeasts in methane hydrate-bearing deep-sea marine sediments. A small portion (7.5%) of total clones belongs to yeast (9 OTUs representing 31 clones). Clones of OTU8, OTU9, OTU12 and OTU22 were assigned to the order of Basidomycota. Phylogenetic analyses strongly supported the affinity between clones of OTU8 and Malassezia restricta, although their sequence identity reached only 85.33%. Clones of OTU9, OTU12 and OTU22 were grouped with Cryptococus congus. Sequence identity of the three OTUs with Cryptococus congus was low (<77%). It indicated novel ITS sequences of Cryptococus genera from the marine sediment. The most closely related sequence of OTU20 was Cladosporium sp. MA4762 with low sequence identity (71.08%). OTU20 formed a separate clade in basidomycota clade; its exact phylogenetic position was not clear (low bootstrap values). Five OTU patterns (OTU4, OTU11, OTU15, OTU21 and OTU24) were included in yeasts belonging to ascomycetes. Clones of OTU4 and OTU21 were well supported with genera Lodderomyces and Candida, respectively. A second clade of yeasts in ascomycetes was formed by clones of OTU11, OTU15 and OTU24, together with Pichia ohmeri.
Discussion
Communities of bacteria, archaea, protists and fungi account for most of the oceanic biomass. These microscopic factories are responsible for 98% of primary production and mediate all biogeochemical cycles in the oceans (Sogin et al., 2006). Marine fungi have been isolated from a variety of detritus materials: decaying woods, leaves, seaweeds, mangroves, sea grasses, sediments, calcareous and chitinous substrates (Rappé et al., 1998; Buchan et al., 2002). Some marine fungi tolerating and adapting to the deep-sea extreme conditions were also isolated (Takami et al., 1997; Nagahama et al., 2003). Culture-independent analysis has produced a very different view of fungal community structure than culture-based methods (Viaud et al., 2000). Therefore, the molecular techniques were adopted to investigate the community structure of marine fungi in the study.
Previous studies showed that 18S rRNA approach is a valuable tool in assessing the global diversity of aquatic eukaryotes (López-García et al., 2001; Moon-van der Staay et al., 2001; Stocek and Epstein, 2003; Stocek et al., 2003). However, the 18S rRNA genes are problematic because identification is commonly limited to genus or family level. This is primarily due to the relative lack of variation within 18S rRNA genes between closely related fungal species (Anderson and Cairney, 2004). The ITS regions of fungal rDNA are stretches of DNA between 18S, 5.8S and 28S rRNA genes and exhibit a high degree of polymorphism between species but are thought to be highly conserved within species. These characteristics make them valuable for genus- and species-level identification (Buchan et al., 2002). Therefore, ITS sequences can provide better taxonomic resolution than 18S rRNA genes (Anderson and Cairney, 2004). Surveys based on 18S rRNA genes indicated that most marine picoplanktons can be assigned to microalgae, flagellates and ciliates (López-García et al., 2001; Moon-van der Staay et al., 2001). However, the ITS 1F primer conjunction with ITS4 primer could specifically amplify fungal templates from mixed community DNA samples and proved to be extremely useful for unraveling the systematic and phylogeny of the dominant salt marsh fungi (Buchan et al., 2002; Chen and Cairney, 2002; Anderson et al., 2003a, 2003b; Anderson and Cairney, 2004).
Fungal communities in methane hydrate-bearing marine sediments assessed by ITS sequences appear to be composed mostly of Phoma glomerata and yeasts, even though PCR does not allow us to quantify accurately cell numbers because of differences in the number of gene copy per cell and PCR biases. Phoma is found as parasites of seaweeds, sea grasses, mollusks and sponges, so it is often isolated from marine and estuarine environment (Kohlmeyer and Volkmann-Kohlmeyer, 1991). There is some evidence that terrestrial microorganisms accumulate in deep-sea sediments (Baross et al., 1975; Pivkin, 2000). Moreover, the isolation of terrestrial fungus Cladosporium sp. from sediments below 4,400 m indicates that sedimentation is probably the most important factor responsible for the accumulation of facultative marine fungi in deep-sea sediments (Takishita et al., 2006; Shao and Sun, 2007). We therefore reconsider the general notion that the role of facultative marine fungi in ecology is of little importance. It is assumed that the marine bacteria present in deep sea have drastically reduced growth and metabolic rates due to the combination of low temperature and high pressure (Baross et al., 1975). Therefore, it is quite likely that some facultative fungi would remain viable but in an inactive state for an indeterminable period.
In general, basidiomycetous and ascomycetous yeasts often account for the majority of the total yeast population in oligotrophic oceanic waters, and among the basidiomycetous yeasts, some species of Cryptococcus and its teleomorphs have been found to be widespread across various oceanic regions including the deep-sea environment (Takishita et al., 2006). In deep-sea sediments, there is less organic debris available to be utilized by yeasts than in sediments in shallow regions. Marine sediments associated with gas hydrates or hydrocarbon seeps are characterized by a rapid increase in consumption of oxygen with depth (Yan et al., 2006). The sulfate-reducing bacteria and methanotrophic archaea are the metabolically active fraction of the microbial community in hydrate-bearing sediments (Mills et al., 2005). These consortia are principally responsible for anaerobic methanotrophy and may contribute significantly to the carbon flux in anaerobic microbial communities fueled by methane. The anaerobic methanotrophy plays a crucial role in the establishment and success of methane-seep communities by converting methane into more readily accessible carbon and energy substrates (Orphan et al., 2002). Most known methanol-assimilating yeast belong to the genera Pichia and Candida and methylotrophic yeasts have attracted interest since they were first isolated for both physiological study and industrial applications (Gellissen, 2000). Methylotrophic yeasts from extreme environments have certain advantages over mesophiles in industrial processes. Consumption of methane hydrate hydrocarbons by associated microbial consortia could play a significant role in global methane and carbon cycles and in diagnostic processes.
Molecular characterization of fungal community from hydrate-bearing sediments has provided evidence for new taxonomic group having no known closely related cultivated isolates. This clade lacks a cultivated isolates and, therefore, the specific physiology is unknown. Attempts will be made to cultivate these fungi identified in the clone libraries to determine their specific physiology and metabolic traits.
Our data suggest that previously unidentified fungal communities occur widely in organic-rich deep-sea marine sediments associated with methane hydrate in South China Sea. Some fungi that we detected represent uncultivated and physiologically uncharacterized assemblages. Nevertheless, the recognition of fungal communities that consistently occur in the presence of methane hydrates serves as a starting point for defining their ecological and biogeochemical significances. Future studies will determine how these fungi affect the cycling of carbon and establish the physiological properties that permit their survival at depth.
References
Anderson IC, Cairney JWG . (2004). Diversity and ecology of soil fungal communities: increased understanding through the application of molecular techniques. Environ Microbiol 6: 769–779.
Anderson IC, Campbell CD, Prosser JI . (2003a). Diversity of fungi in organic soils under a moorland-Scots pine (Pinus sylvestris L.) gradient. Environ Microbiol 5: 1121–1132.
Anderson IC, Campbell CD, Prosser JI . (2003b). Potential bias of fungal 18S rDNA and internal transcribed spacer polymerase chain reaction primers for estimating fungal biodiversity in soil. Environ Microbiol 5: 36–47.
Altschul SF, Madden TL, Schaffer AA, Zhang JH, Miller W, Lipman D . (1997). Gapped-BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25: 3389–3402.
Baross JA, Hanus FJ, Morita RY . (1975). Survival of human enteric and other sewage microorganisms under simulated deep-sea conditions. Appl Microbiol 30: 309–318.
Buchan A, Newell SY, Moretam JIL, Moranm MA . (2002). Analysis of internal transcribed spacer (ITS) regions of rRNA genes in fungal communities in a southeastern US salt marsh. Microb Ecol 43: 329–340.
Chen DM, Cairney JWG . (2002). Investigation of the influence of prescribed burning on ITS profiles of ectomycorrhizal and other soil fungi at three Australian sclerophyll forest sites. Mycol Res 106: 532–540.
Gabor EM, Vries EJ, Janssen DB . (2003). Efficient recovery of environmental DNA for expression cloning by indirect extraction methods. FEMS Microbiol Ecol 44: 153–163.
Gardes M, Bruns TD . (1993). ITS primers with enhanced specificity for basidiomycetes: application to the identification of mycorrhiza and rusts. Mol Ecol 2: 113–118.
Gellissen G . (2000). Heterologous protein production in methylotrophic yeasts. Appl Microbiol Biotechnol 54: 741–750.
Good IJ . (1953). The population frequencies of species and the estimation of the population parameters. Biometrika 40: 237–264.
Gotelli NJ, Entsminger GL . (2005). EcoSim: null models software for ecology. Version 7. Acquired Intelligence Inc. & Kesey-Bear. Jericho, VT 05465.http://garyentsminger.com/ecosim.htm.
Kohlmeyer J, Volkmann-Kohlmeyer B . (1991). Illustrated key to the filamentous higher marine fungi. Bot Mar 34: 1–61.
Liu XW, Li MF, Zhang YW, Zhang GX, Wu NY, Huang YY et al. (2005). Studies of seismic characteristics about gas hydrate: A case study of line HD152 in the South China Sea. Geoscience 19: 33–38 (in Chinese).
López-García P, Rodríguez-Valera F, Pedrós-Alió C, Moreira D . (2001). Unexpected diversity of small eukaryotes in deep-sea Antarctic plankton. Nature 409: 603–606.
MacDonald IR, Guinasso NL, Sassen R, Brooks JM, Lee L, Scott KT . (1994). Gas hydrate that breaches the sea-floor on the continental-slope of the Gulf of Mexico. Geology 22: 699–702.
Maidak BL, Cole JR, Lilburn TG, Parker CY, Saxman PR, Farris RJ et al. (2001). The RDP-α (ribosomal database project). Nucleic Acids Res 29: 173–174.
Miller DN . (2001). Evaluation of gel filtration resins for the removal of PCR-inhibitory substances from soils and sediments. J Microbiol Meth 44: 48–58.
Mills HJ, Martinez RJ, Story S, Sobecky PA . (2005). Characterization of microbial community structure in Gulf of Mexico gas hydrates: comparative analysis of DNA- and RNA-derived clone libraries. Appl Environ Microbiol 71: 3235–3247.
Moon-van der Staay SY, De Wachter R, Vaulot D . (2001). Oceanic 18SrDNA sequences from picoplankton reveal unsuspected eukaryotic diversity. Nature 409: 607–610.
Nagahama T, Hamamoto M, Nakase T, Takaki Y, Horikoshi K . (2003). Cryptococus surugaensis sp. nov., a novel yeast species from sediment collected on the deep-sea floor of Suruga Bay. Int J Syst Evol Microbiol 53: 2095–2098.
Orphan VJ, House CH, Hinrichs KU, Mckeegan KD, Delong EF . (2002). Mutiple archaeal groups mediate methane oxidation in anoxic cold seep sediments. Proc Natl Acad Sci USA 99: 7663–7668.
Pivkin MV . (2000). Filamentous fungi associated with holothurians from the sea of Japan, off the primorye coast of Russia. Biol Bull 198: 101–109.
Rappé MS, Suzuki MT, Vergin KL, Giovannoni SJ . (1998). Phylogenetic diversity of ultraplankton plastid small-subunit rRNA genes recovered in environmental nucleic acid samples from the Pacific and Atlantic coasts of the United States. Appl Environ Microbiol 64: 294–303.
Reed DW, Fujita Y, Delwiche ME, Blackwelder DB, Sheridan PP, Uchida T et al. (2002). Microbial communities from methane hydrate-bearing deep marine sediments in a forearc basin. Appl Environ Microbiol 68: 3759–3770.
Robe P, Nalin R, Capellano C, Vogel TM, Simonet P . (2003). Extraction of DNA from soil. Eur J Soil Biol 39: 183–190.
Sassen R, Joye S, Sweet ST, DeFreitas DA, Milkov AV, MacDonald IR . (1999). Thermogenic gas hydrates and hydrocarbon gases in comples chemosynthetic communities, Gulf of Mexcio continental slope. Org Geochem 30: 485–497.
Shannon CE, Weaver W . (1949). The Mathematical Theory of Communication. University of Illinois Press: Urbana, USA.
Shao Z, Sun F . (2007). Intracellular sequestration of manganese and phosphorous in a metal-resistant fungus Cladosporium cladosporioides from deep-sea sediment. Extremophiles 11: 435–443.
Sogin ML, Morrison HG, Huber JA, Welch DM, Huse SM, Neal PR et al. (2006). Microbial diversity in the deep sea and the underexplored ‘rare biosphere’. Proc Natl Acad Sci USA 103: 12115–12120.
Stocek T, Epstein S . (2003). Novel eukaryotic lineages inferred from small-subunit rRNA analyses of oxygen-depleted marine environments. Appl Environ Microbiol 69: 2657–2663.
Stocek T, Taylor GT, Epstein S . (2003). Novel eukaryotes from the permanently anoxic Cariaco Basin (Caribbean Sea). Appl Environ Microbiol 69: 5656–5663.
Takami H, Inoue A, Horikoshi K . (1997). Microbial flora in the deepest sea mud of the Mariana Trench. FEMS Microbiol Lett 152: 279–285.
Takishita K, Tsuchiya M, Reimer JD, Maruyama T . (2006). Molecular evidence demonstrating the basidiomycetous fungus Cryptococcus curvatus is the dominant microbial eukaryote in sediment at the Kuroshima Knoll methane seep. Extremophiles 10: 165–169.
Viaud M, Pasquier A, Brygoo Y . (2000). Diversity of soil fungi studied by PCR-RFLP of ITS. Mycol Res 104: 1027–1032.
White TJ, Bruns TD, Lee S, Taylor J . (1990). Analysis of phylogenetic relationships by amplification and direct sequencing of ribosomal RNA genes. In: Innis MA, Gelfand DH, Sninsky JJ, White J (eds). PCR Protocols: a Guide to Methods and Applications. Academic Press: New York, pp 315–322.
Yan T, Ye Q, Zhou J, Zhang CL . (2006). Diversity of functional genes for methanotrophs in sediments associated with gas hydrates and hydrocarbon seeps in the Gulf of Mexico. FEMS Microbiol Ecol 57: 251–259.
Zhang GX, Huang YY, Zhu YH, Wu BH . (2002). Prospect of gas hydrate resources in the South China Sea. Mar Geol Tern Geol 22: 75–81 (in Chinese).
Acknowledgements
This work was supported by grant from the National High Technology Research and Development Plan of China (2003AA620401). We thank Yongyang Huang and Jian Liu, Guangzhou Marine Geological Survey, for their assistance during this study.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Lai, X., Cao, L., Tan, H. et al. Fungal communities from methane hydrate-bearing deep-sea marine sediments in South China Sea. ISME J 1, 756–762 (2007). https://doi.org/10.1038/ismej.2007.51
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2007.51
Keywords
This article is cited by
-
Sedimentary DNA for tracking the long-term changes in biodiversity
Environmental Science and Pollution Research (2023)
-
Adaptation mechanisms of the soil microbial community under stoichiometric imbalances and nutrient-limiting conditions in a subtropical nitrogen-saturated forest
Plant and Soil (2023)
-
Assessment of yeasts in tropical peat swamp forests in Thailand
Mycological Progress (2020)
-
Cultivable fungi present in deep-sea sediments of Antarctica: taxonomy, diversity, and bioprospecting of bioactive compounds
Extremophiles (2020)
-
Characterization of the total and viable bacterial and fungal communities associated with the International Space Station surfaces
Microbiome (2019)