Abstract
Acoustic signals often have a significant role in pair formation and in species recognition. Determining the genetic basis of signal divergence will help to understand signal evolution by sexual selection and its role in the speciation process. An earlier study investigated quantitative trait locus for male courtship song carrier frequency (FRE) in Drosophila montana using microsatellite markers. We refined this study by adding to the linkage map markers for 10 candidate genes known to affect song production in Drosophila melanogaster. We also extended the analyses to additional song characters (pulse train length (PTL), pulse number (PN), interpulse interval, pulse length (PL) and cycle number (CN)). Our results indicate that loci in two different regions of the genome control distinct features of the courtship song. Pulse train traits (PTL and PN) mapped to the X chromosome, showing significant linkage with the period gene. In contrast, characters related to song pulse properties (PL, CN and carrier FRE) mapped to the region of chromosome 2 near the candidate gene fruitless, identifying these genes as suitable loci for further investigations. In previous studies, the pulse train traits have been found to vary substantially between Drosophila species, and so are potential species recognition signals, while the pulse traits may be more important in intra-specific mate choice.
Similar content being viewed by others
Introduction
Both sexual selection and speciation processes are important research areas in evolutionary biology (Jones and Ratterman, 2009). Speciation depends on the evolution of reproductive isolation between populations. Reproductive isolation often relies on divergence of courtship signals and preferences, which may occur because of sexual selection (Ritchie, 2007). Among courtship signals, acoustic communication is one of the most common and well-studied channels (Coleman, 2009) and it has been found to have a crucial role in the reproductive behaviours of many birds, mammals, frogs and insects.
Males of almost all fruit fly (genera Sophophora and Drosophila) species studied so far produce complex courtship songs by vibrating their wings (Ewing and Miyan, 1986; Talyn and Dowse, 2004; Hoikkala and Mazzi, 2009). These songs are species specific and often serve as a component of the critical set of signals females use to recognize conspecific males (Kyriacou and Hall, 1980; Ritchie et al., 1998). Intra-specific variation in male acoustic signals also has a role in sexual selection influencing mating decisions of females, for example, in distantly related Drosophila melanogaster and Drosophila montana (Aspi and Hoikkala, 1995; Talyn and Dowse, 2004). However, the roles of chemical, visual, tactile and acoustic signals in sexual selection and species-recognition vary between species (Cobb et al., 1986). In some species, like Drosophila subobscura (Ewing and Bennet-Clark, 1968) and several Hawaiian picture-winged Drosophila species (Hoikkala et al., 1989), the males do not produce any song during courtship, while in other species, for example, D. montana and Drosophila ezoana, the song is an obligatory step leading to insemination (Hoikkala, 1988). Research on the genetic control of the song characters in D. montana is particularly relevant to understanding the evolution of reproductive barriers based on acoustic cues. It is a model for addressing important and unresolved questions in the evolution of reproductive isolation, such as whether genes under sexual selection are located on autosomal or sex-linked chromosomes (Qvarnström and Bailey, 2009), or whether the same genes control variation within and divergence between species (Gleason and Ritchie, 2004).
Quantitative trait locus (QTL) analysis allows one to assess the number, nature and genomic locations of loci affecting traits of interests. Genome scans employing anonymous DNA markers, like microsatellites or single-nucleotide polymorphisms (SNPs), can be used to identify the genetic architecture of QTL underlying complex quantitative traits. If candidate genes (genes previously identified as having an effect on the trait) are implicated in QTL peaks, the natural next step is to examine these further, to dismiss or confirm their potential involvement. By using molecular markers located within candidate genes, one can better evaluate their potential contribution to the traits. This type of approach has been especially successful in explaining intra-specific trait variation (reviewed in Haag and True, 2001; Fitzpatrick et al., 2005).
There are many QTL studies examining patterns of inheritance of song characters within and between species, but only a few studies so far have supplemented QTL techniques with candidate gene approaches (reviewed in Hoikkala and Mazzi, 2009). Targeted mutation of known genes in D. melanogaster (subgenus Sophophora) has provided candidate genes for the evaluation of mapped QTL in fly species other than D. melanogaster. Many mutations are known to alter the pattern of male courtship song in D. melanogaster (Gleason, 2005).
Our study species, D. montana, belongs to the Drosophila virilis species group, and has courtship song consisting of polycyclic sound pulses arranged in pulse trains (Hoikkala and Lumme, 1987). This species inhabits temperate forests of the northern hemisphere, forming several distinct genetic clusters between and within the continents (Mirol et al., 2007). Earlier studies revealed significant differences in several song characters, wing shape and genital morphology, between populations from distinct geographic regions (Suvanto et al., 2000; Klappert et al., 2007; Routtu et al., 2007). The identification of polymorphic microsatellite markers in D. montana has led to the construction of a linkage map and to the mapping of QTL for song carrier frequency (FRE) differences between populations (Schäfer et al., 2010). The largest QTL mapped to an area of low recombination, containing many loci. With the D. virilis genomic sequence being recently published (Clark et al., 2007), development of molecular markers for candidate genes in non-model species from this species group has become much easier and we have exploited this opportunity to include candidate loci in our analysis of song variation in D. montana.
Specific objectives of this study were the following: (1) to develop genetic markers within candidate genes for song characters in D. montana; and to add them to the microsatellite linkage map described in Schäfer et al. (2010); (2) to include data on five additional song characters; and (3) to identify and describe QTLs responsible for the divergence of courtship song between two strains originating from natural populations in order to elucidate the genetic architecture of a complex quantitative trait. More generally, we aim to compare the results obtained with published results on QTL in the other species from the D. virilis group. In this way, we will provide insights into whether the loci accounting for song variation within a species are the same as the ones causing divergence between species and also into whether the number and locations of these loci are similar to those already known. At the same time, the candidate gene approach allows us to test the relationship between the loci for which song aberrations have been found in the model species and those underlying naturally occurring variation in song production in a non-model species.
Materials and methods
Insect material and song characters
We worked on a cross between two strains described in Schäfer et al. (2010) known to be divergent for song characters. The two genetically variable isofemale strains representing natural populations from Finland (Oulanka, O3F66) and from the USA (Colorado, C3F13) were kept under laboratory conditions for approximately 1 year before crossing. The males of the Finnish population have earlier been shown to have higher song carrier FRE than the ones from the North American population (Klappert et al., 2007; Routtu et al., 2007). Females and males from the parental strains were reciprocally mated en masse to produce a large number of offspring. The resulting F1 generation consisted of two types of offspring: F1A (from C3F13 females and O3F66 males) and F1B (from O3F66 females and C3F13 males). Individuals from the F1 generation were mated within and between the groups in four F2 combinations: F1AxF1A (F2AA), F1AxF1B (F2AB), F1BxF1B (F2BB) and F1BxF1A (F2BA) (parental females are always listed first in the cross code). Courtship song was recorded at 20±1o and analysed as described in Schäfer et al. (2010). Song data for 64 males from the P generation, 48 males from the F1 generation and 723 males from the F2 generation representing the upper and lower 30% of the song FRE distribution were included in QTL analysis. In addition to the song carrier FRE analysed in Schäfer et al. (2010), we also analysed the following song characters: cycle number (CN), pulse length (PL), pulse number (PN), pulse train length (PTL) and interpulse interval (IPI). A schematic representation of these song traits is presented in Figure 1. Mean values from three measurements of each song character for each individual were used for statistical analyses.
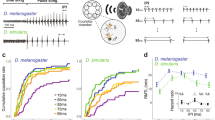
Schematic representation of a male D. montana courtship song oscillogram showing how measurements of six song traits were obtained. FRE, song carrier FRE (Hz); CN, number of cycles in a pulse; PL, pulse length (ms); PN, number of pulses per pulse train; PTL, length of a pulse train (ms); IPI, interpulse interval (ms).
Candidate gene markers and PCR
We developed new markers for polymorphisms segregating between the two D. montana populations in 10 candidate genes for song characters, from the list provided by Gleason (2005): black (bla), cacophony (cac), cystein string protein (csp), ebony (ebo), fruitless (fru), maleless (mle), paralytic (par), period (per), slowpoke (slo) and temperature-induced paralytic E (tip). These candidate genes were selected because they have been found to affect male courtship song in D. melanogaster and we were able to use genome sequence data from D. melanogaster and D. virilis to design primers that amplified part of the target locus from D. montana. To develop the markers, we based initial sequencing primers on the published D. virilis genome and used them to sequence fragments of candidate genes from three random individuals from each of the two parental D. montana strains (O3F66 and C3F13). The sequences were screened for polymorphism and the sites that differed most in allele FRE between the parental strains were used for marker development. The list of candidate genes included in our study, with their genomic location, type and sequences of respective PCR primers are presented in Supplementary Table S1. The alignments of PCR-amplified DNA sequences that form the basis of the developed markers are included in Supplementary Figure S1. Representative sequences have been submitted to GenBank with accession numbers HQ889727 to HQ889746.
Seven of the candidate gene markers were based on the presence–absence of small indels (1–15 bp insertions–deletions) within the PCR-amplified sequence. The amplified fragments of different length were fluorescently labelled and visualized on an Applied Biosystems 373048-capillary DNA analyser (Applied Biosystems, Lincoln, NE, USA). Each 10 μl PCR reaction mix contained 1 μl of template DNA (1:20 dilution in ultrapure water), 1 μl of 10 × Mg-free reaction buffer IV (Bioline, London, UK), 1 mM MgCl2, 2 mM of each dNTP, 5 μM forward and 0.5 μM reverse primer and 0.5 μM of universal FAM or HEX fluorescently labelled oligo used for labelling PCR products in the same reaction. In all 0.05 U of Taq DNA polymerase (Bioline) was used in each PCR reaction. The PCR thermocycling program started with 94 °C for 3 min for initial DNA denaturation, then 94 °C for 30 s, 58 °C for 30 s, 72 °C for 30 s, for 10 cycles, then 94 °C for 30 s, 54 °C for 30 s, 72 °C for 30 s, for 10 cycles, followed by 20 cycles of 94 °C for 30 s, 50 °C for 30 s, 72 °C for 3 min. The final extension step after the last cycle was at 72 °C for 10 min.
For the remaining three genes (para, black and ebo), the markers were based on SNPs. The initial PCR step resulted in amplification of sequence fragments of the same length, but differing in SNP composition. The PCRs were carried out in 20 μl reactions made of 2–3 μl of template DNA (1:20 dilution in ultrapure water), 2 μl of 10 × Mg-free reaction buffer IV (Bioline), 1.5 mM MgCl2, 2 mM of each dNTP, 7.5 μM of forward and reverse primers, and 0.1 U of Taq DNA polymerase (Bioline) per reaction. Thermal cycling program consisted of 95 °C for 3 min, then 95 °C for 1 min, 61 °C for 1 min, 72 °C for 1 min, for 30 cycles, followed 72 °C for 5 min. PCR-amplified DNA fragments containing SNPs were digested with restriction enzymes, which selectively cut the DNA strands at one of the SNP variants. We used restriction enzyme FokI for the paralytic gene marker, NotI for black and SexA for ebo. Digestions were carried out following the manufacturer's instructions (New England BioLabs, Ipswich, MA, USA). The resulting DNA fragments were then resolved on high-resolution agarose (RESponse Research PCR agarose 335010, RESoure Line Agaroses BIOzym) containing Syber-Safe (Invitrogen, Carlsbad, CA, USA) and visualized under ultraviolet-light.
Analyses
We compared song characters between strains and generations and calculated correlations among different song traits in the F2 cross generation using R software version 2.11.8 (R Development Core Team, 2008), as for most of the subsequent analyses we first tested for associations between alleles of candidate gene markers and song characters using a linear regression approach. In the next step, existing data on microsatellite genotypes and song carrier FRE were supplemented with genotypes for the 10 candidate gene markers and five additional song characters. New markers were added to the microsatellite linkage map using the program MAPMAKER v.3.0 (Lander et al., 1987). The updated map was subsequently used in QTL analyses with the R/qtl package version 1.15-15 (Broman et al., 2003) to map QTL and test for epistasis by the Multiple Imputation Mapping method. We chose this method because it copes well with non-normally distributed phenotypic data (for example, due to selective genotyping) and with missing genotype information (Broman and Sen, 2009). Logarithm of odds (LOD) significance thresholds for QTL detection were established using 3000 or 1000 permutations for single-QTL and two-QTL scans, respectively. We calculated 95% approximate Bayesian Credible Intervals for the detected QTL. As many song characters appeared to be inter-correlated, principal component analysis (PCA) was carried out to reduce data dimensionality and further analyse the relationship between song variation and genetic variation using QTL techniques, as outlined above. For performing PCA, we used a Bayesian method, which can deal with incomplete data via missing value imputation, implemented in the bpca function in the pcaMethods R package (Stacklies et al., 2007).
Results
Male courtship song characters
We measured six characters of the courtship song of D. montana males (Figure 1) to capture the complexity of this signal. We found significant variation between the two parental strains C3F13 and O3F66 in all measured song traits (t-test, P<0.001, Supplementary Table S2) except PTL. Males from C3F13 had more cycles per pulse (CN) and longer pulses (PL) than O3F66 males. Also, IPIs were longer in the former than the latter. In contrast, we observed fewer pulses (PN) per pulse train in C3F13 individuals, which cancelled out the effect of having longer pulses and resulted in a lack of difference in PTL between the two strains. Individuals from the F1- and F2-cross-generations showed intermediate and highly variable values of the courtship song characters, in comparison with the two parental strains (Figure 2). The high variance of the song traits in the F2 suggests that few loci of large effect might be responsible for song divergence.
Medians (±25 and 75% quantiles, min and max values) of the male courtship song characters in the parental lines C3F13 (Colorado, USA), O3F66 (Oulanka, Finland) and the reciprocal crosses of the first (F1) and second (F2) generations. F1A, cross between C3F13 females and O3F66 males; F1B, O3F66 females and C3F13 males. In the F2 generation strain labels, parental females are listed first and parental males second. For description of song characters refer to Figure 1.
We calculated pairwise correlations among song characters in the F2 and found that most characters are significantly correlated (Table 1). Song carrier FRE appeared to be strongly related to CN, but not to PL. There was also no significant association between carrier FRE and PTL, or number of pulses in a train (PN) and IPI. On the other hand, strong correlations (r>0.5) were observed between PL and CN, PTL and PN, and PL and IPI.
PCA on the song characters resulted in two reduced axes, PC1 and PC2, which accounted for 36% and 28% of variation, respectively. The distribution of scores and loadings of the song characters for the first two principal components are shown in Supplementary Figure S2. Song traits CN, PL, IPI and FRE had negative loadings for PC1, while PTL and PN had positive loadings for PC2. As a result, PC1 represented mainly variation in song pulse structure and PC2 represented variation in song pulse trains.
Candidate genes—single marker regressions
Single marker regression analysis revealed significant associations between song carrier FRE and three candidate gene markers: strong for ebo and fru, and weak for slo (Supplementary Table S3). CN in a pulse, a character closely correlated with FRE, was also associated with markers ebo and fru, while PL was linked to fru only. IPI values were also associated with markers ebo and fru, and additionally with per. Two strongly correlated traits related to pulse train properties, PTL and PN, were associated mainly with another candidate gene marker slo.
Linkage map
Genotype information from 10 candidate gene markers was added to the existing genotype data for 42 microsatellites from a previous study (Schäfer et al., 2010) to construct a refined linkage map. The order and marker distances inferred from the extended data set were in agreement with those published earlier, with candidate gene markers per, par and cac mapping on the X chromosome, markers ebo, fru and slo on chromosome 2, markers tip and csp on chromosome 3, marker bla on chromosome 4, and marker mle on chromosome 5 (Supplementary Figure S3).
Single-QTL mapping
Logarithm of odds (LOD) profiles for six song characters from single-QTL scans along the five chromosomes are shown in Figure 3. A summary of the identified QTL is given in Table 2 along with the names of markers falling into the confidence intervals harbouring each QTL. The analyses revealed two main intervals containing significant QTL for courtship song characters. The first interval was located on the X chromosome at positions 96–148 cM (most conservative 95% Bayesian Credible Interval) and contained overlapping QTL peaks for two correlated characters: PTL and PN per train. This interval also contained a marker for the candidate gene period (per). Allele effect plots (Supplementary Figure S4) showed that the per allele most common in the parental strain O3F66 (Oulanka, Finland) is associated with high values of PTL and PN. The second detected interval with significant QTL was located on chromosome 2 between 36 and 88 cM and contained LOD peaks for four inter-correlated song characters: carrier FRE, CN per pulse, PL and IPI length. Markers for the candidate genes fru and ebo fall into this interval. Allele effect plots (Supplementary Figure S4) for these two genes showed that the alleles prevalent in the parental strain O3F66 are associated with lower values of these four song characters and they are largely dominant over the alternative alleles. Hence, all allele effects are in the expected direction given the parental differences.
Multiple imputation mapping of six male courtship song characters against the marker positions on X chromosome, four autosomes and a hypothetical Y-chromosome marker. Horizontal lines indicate 95% significance thresholds for detecting a QTL for the single-QTL scan for each song character. For description of song characters refer to Figure 1.
The results of single-QTL mapping using the first two principal components of song trait variation revealed a very similar pattern (Table 2). PC1, representing mainly pulse-associated traits, had one major-effect QTL at the same location as FRE, CN, PL and IPI. PC2, representing pulse train-associated traits, had one QTL at the same location as PTL and PN, but also a second QTL on chromosome 2 close to microsatellite marker mon40 (Supplementary Figure S5).
Two-QTL mapping
The possibility of multiple QTL, suggested by the multiple peaks in the LOD curves was assessed by a two-QTL analysis. At the same time, the possibility of epistasis (interactions between two QTL) was tested. The results provided some evidence for the existence of more than one QTL for song characters, but no evidence for epistasis (Supplementary Table S4). For song characters PN and PTL, both with major-effect QTL on chromosome X, two additional QTL were indicated on chromosome 2. For FRE and IPI, both with major-effect QTL on chromosome 2, a second QTL was suggested on chromosome X; while for CN an additional QTL was suggested on chromosome 3. When the analyses were carried out for PC1 and PC2, similar conclusions were obtained (Supplementary Table S4). PC1, representing mainly traits PL, CN, IPI and FRE, had one major-effect QTL on chromosome 2, position 64 cM. For PC2, representing mainly traits PN and PTL, at least two weak-effect QTL were indicated on chromosomes X and 2. The results of two-QTL analyses have to be treated with caution, however, because of the unresolved statistical problems linked to the use of X chromosome data (Broman and Sen, 2009).
Stepwise QTL mapping
We performed stepwise QTL analysis to further evaluate the possibility of more than one QTL for song characters. A forward–backward model selection procedure was used to identify the best model, with model choice made via a penalized LOD score (Manichaikul et al., 2009). The procedure yielded a set of genetic models that were largely consistent with the results of single- and two-QTL scans. First, it showed that single-QTL models are most likely for the song characters FRE, CN, PL and IPI, with one major-effect QTL located on chromosome 2 near the candidate genes fru and ebo. These models explained between 4 and 16% of the variation in these song characters and had an associated P-value <0.00001 (except IPI, P=0.0002). Second, in line with the two-QTL scans, PN in a pulse train appeared to be associated with three QTL: one on the X chromosome (near candidate gene per) and two others, one near each end of chromosome 2 (one near candidate gene slo). The model explained 11.5% of the variation and had an associated P-value <0.00001. Finally, the analysis showed that a no-QTL model is most likely for the PTL (it is likely that this represents the duration of courtship bouts, which is determined by an interaction between males and females). The results of analyses using principal components to summarize variation in the song traits suggested that single-QTL models were most likely for PC1 and PC2: PC1 having a QTL on chromosome 2 at the location around 62 cM (near candidate gene marker fru), and PC2 also having a QTL on chromosome 2 at the location around 6 cM (near microsatellite marker mon40). These results are summarized in Supplementary Table S5 and Supplementary Figure S6. It has to be noted that the penalized LOD score criterion is currently defined only for autosomal loci (Broman and Sen, 2009), and thus analyses performed using the stepwise model selection algorithm on the data with the X chromosome should be considered with caution (QTL on X chromosome being less likely to be detected).
Discussion
In this study, we have identified several QTL associated with the divergence of male courtship song between the two natural strains of D. montana, C3F13 and O3F66 from the USA and Finland, respectively. The six song characters we investigated fell into two types: traits related to pulse properties (FRE, CN, PL and IPI, the latter strongly correlated with PL) and traits related to pulse train properties (PTL and PN). This division was reflected in principal components PC1 and PC2, and was associated with different QTL. Single, two and stepwise QTL mapping analyses revealed one major-effect QTL on the second chromosome influencing all pulse traits. A similar QTL was earlier identified as the main QTL for song carrier FRE by Schäfer et al. (2010), where this QTL was found to coincide with a region of low recombination, possibly harbouring the polymorphic inversion 2Y (Morales-Hojas et al., 2007). In contrast to song pulse traits, the song characters related to PTL (PTL and PN) were found to have a minor-effect QTL on the X chromosome, with PN having at least one additional QTL on an autosome. QTL for the pulse train characters were relatively weak and explained a small proportion of variation (4–6%), suggesting polygenic determination of these characters.
The auditory signals produced by males have a role in mate choice and species recognition. Different song characters are responsible for these two functions in D. montana (reviewed by Hoikkala et al., 2005) and our study contributes to understanding whether these traits are determined by the same or different QTL. Among the six traits we used to quantify song variability, song carrier FRE has been shown to be an important factor in mate choice (Aspi and Hoikkala, 1995; Ritchie et al., 1998; Klappert et al., 2007). Females from the Finnish population prefer males with high-FRE song (Ritchie et al., 1998, 2001; Klappert et al., 2007) and in this population song FRE correlates positively with male condition and offspring survival (Hoikkala et al., 1998, 2008). PL and CN also have a role in sexual selection in the Finnish flies, with females preferring shorter pulses with more cycles per pulse (Aspi and Hoikkala, 1995; Ritchie et al., 1998, 2001), which, consequently, results in preference for high song FRE. However, female preferences can differ between populations—flies from the Colorado population were shown to respond more to songs with lower carrier FRE and, thus, different song traits are likely to be selected for by Colorado females (Klappert et al., 2007). Despite this observation, we can reasonably assume that FRE and other pulse traits evolve under sexual selection in most populations, and therefore, might be expected to be controlled by X chromosomal genes (for example, Reinhold, 1998). We found no evidence for such sex-linkage, as only a single major-effect QTL was revealed on the second chromosome for the pulse-related song traits. This result is in agreement with work on inbred and outbred D. montana strain hybrids (Aspi, 2000; Suvanto et al., 2000) and with studies on species from the D. melanogaster group (Colegrave et al., 2000; Gleason and Ritchie, 2004), which indicated that most song traits were influenced by factors on the autosomes. Although the evidence favouring sex-linkage of genes influencing sexually selected traits is quite strong, it may depend on the mechanism of sex determination (Qvarnström and Bailey, 2009).
To date, within-species genetic variation in male song traits has only been studied in two other species from the D. virilis group: D. virilis and Drosophila littoralis. In the former species, the genetic basis of PN and PTL was initially shown to be mainly autosomal and polygenic (Huttunen and Aspi, 2003). Subsequently, significant QTL affecting PN were located on the X and on chromosomes 2, 3 and 4, and QTL located on the third chromosome were linked to PTL variation (Huttunen et al., 2004). In D. littoralis, FRE and CN were found to be mainly determined by loci on chromosome 2 (Hoikkala and Lumme, 1990), as in our results. Contrary to our findings in D. montana, the IPI of D. littoralis was linked to factors on the X chromosome. However, the IPI of D. littoralis song (∼300 ms) is much longer than that of D. montana (<40 ms) and the genes affecting IPI in the former species could well be the same ones that give rise to species differences in this trait.
We addressed an additional important question on the relationship between the loci for which song aberrations have been found in the model species and those underlying naturally occurring variation in song production in non-model species by adding candidate gene markers to the D. montana microsatellite linkage map created for QTL mapping. Candidate genes were used as markers in QTL analyses to determine whether they potentially contribute to QTL peaks. We found that, out of the 10 candidate genes studied, 2 genes consistently fell into confidence intervals of QTL. The QTL for pulse train traits (PTL, PN) located on the X chromosome harboured markers for the candidate gene period. This regulatory gene has long been known for its effects on circadian rhythm and song rhythm in D. melanogaster and other related species (Kyriacou, 2002). The mutations in period investigated by Kyriacou and Hall (1980) affected periodic oscillations of IPI, but not mean IPI or the pulse duration. Although D. montana is not known to have the long IPI cycles coded by the period gene in species of the D. melanogaster group, our result implies that this gene may influence other pulse train traits in D. montana. This suggestion comes from a study on courtship song of Bactrocera cucurbitae flies (Miyatake and Kanmiya, 2004). The investigators investigated the relationship between circadian locomotor rhythm (controlled by period) and courtship song traits. They found a positive correlation between the length of the locomotor circadian rhythm and length of the interval between pulse trains, which was not investigated in our study. In contrast, pulse train duration (PTL) in Bactrocera was not linked to circadian rhythm.
The QTL for pulse traits (FRE, CN, PL and IPI) located on the second chromosome harboured markers for the candidate gene fru. Fru belongs to the group of sex-determining genes that alter the courtship behaviour of the males, including song production (reviewed by Baker et al., 2001). It is known to have a major role in masculinising the male components of the central nervous system and influencing the production of the courtship song. Mutations in the fru gene in D. melanogaster caused complete absence of song or a pulse song of low quality, expressed as fewer pulse trains per minute, lower mean number of pulses per train and shorter IPIs (Villella et al., 1997; Rideout et al., 2007). Although the latter is not strong evidence for the function of fru in intra-specific song variation in D. montana, at least it shows that relevant traits are affected in D. melanogaster. The nonA/dissonance gene is also known to influence pulse cycle shape in flies of the D. virilis group (Campesan et al., 2001): we attempted to develop a polymorphic marker for this gene, with no success. However, nonA is located next to cacophony in the D. virilis genome and therefore most likely outside any QTL detected by us. Similarly, we did not find any QTL near the slowpoke gene, although some research results indicate that mutations in genes encoding or affecting ion-channel function (also including cac, para and nap) might be a source of intra-specific variation in courtship song of D. melanogaster (Peixoto and Hall, 1998).
QTL studies involving candidate genes can eliminate genes from further study as potential causal loci. Finding co-occurrence of QTL and candidate genes is not sufficient to determine that a particular gene is involved in determining a trait because each QTL peak covers many loci. Nevertheless, it presents a compelling case for more detailed study of the gene and the surrounding genomic region. The knock-down and knock-out approaches seem a risky way to proceed, because our candidate genes are involved in sex determination and multiple behavioural traits. One of the techniques used to investigate the role of genes in determining species-specific phenotypes is gene transfer. Unfortunately, there are only a few applications of this technique we are aware of related to Drosophila courtship song genes. A transfer of the period gene from Drosophila simulans to D. melanogaster resulted in changes in the species-specific characteristics of circadian locomotor activity and courtship song cycles (Wheeler et al., 1991). A transfer of nonA between D. virilis and D. melanogaster showed that nonA, like the period gene, might encode species-specific song characters (Campesan et al., 2001). Unfortunately, transformation studies are still technically complicated, especially for long and complex genes like, for example, fru.
A promising direction for further research lies in molecular studies on the level of DNA sequence. Therefore, the investigation of candidate genes for song in D. montana will go in two directions. First, looking into inter-specific DNA sequence variation within the virilis group of species and, then, relating sequence divergence and evolutionary rates to species-specific song characters using comparative methods. This work is currently in progress, and sufficient sequence coverage was obtained for four candidate genes: cac, fru, mle and tipE. Second, as genome sequencing becomes more feasible, we plan to sequence the transcriptome of D. montana and then use capture sequencing of candidate genes from multiple individuals and populations to test for associations between variation at the molecular and phenotype levels. Such an investigation could ultimately lead to the identification of quantitative trait genes and differences between their alleles, pinpointing the functional changes in DNA sequence that influence song variation involved in sexual selection and speciation.
Data archiving
Sequence data have been submitted to GenBank: accession numbers HQ889727–HQ889746. Genotype data have been deposited at Dryad: doi:10.5061/dryad.8q3p913p.
Accession codes
References
Aspi J (2000). Inbreeding and outbreeding depression in male courtship song characters in Drosophila montana. Heredity 84: 273–282.
Aspi J, Hoikkala A (1995). Male mating success and survival in the field with respect to size and courtship song characters in Drosophila littoralis and D. montana (Diptera: Drosophilidae). J Insect Behav 8: 67–87.
Baker BS, Taylor BJ, Hall JC (2001). Are complex behaviors specified by dedicated regulatory genes? Reasoning from Drosophila. Cell 105: 13–24.
Broman KW, Sen S (2009). A Guide to QTL Mapping with R/qtl. Springer: New York.
Broman KW, Wu H, Sen S, Churchill GA (2003). R/qtl: QTL mapping in experimental crosses. Bioinformatics 19: 889–890.
Campesan S, Dubrova Y, Hall JC, Kyriacou CP (2001). The nonA gene in Drosophila conveys species-specific behavioural characteristic. Genetics 158: 1535–1543.
Clark AG, Eisen MB, Smith DR, Bergman CM, Oliver B, Markow TA et al. (2007). Evolution of genes and genomes on the Drosophila phylogeny. Nature 450: 203–218.
Cobb M, Burnet B, Connolly K (1986). The structure of courtship in the Drosophila melanogaster species sub-group. Behaviour 97: 182–212.
Colegrave N, Hollocher H, Hinton K, Ritchie MG (2000). The courtship song of African Drosophila melanogaster. J Evol Biol 13: 143–150.
Coleman SW (2009). Taxonomic and sensory biases in the mate-choice literature: there are far too few studies of chemical and multimodal communication. Acta Ethol 12: 45–48.
Ewing AW, Bennet-Clark HC (1968). The courtship songs of Drosophila. Behaviour 31: 288–301.
Ewing AW, Miyan JA (1986). Sexual selection, sexual isolation and the evolution of song in the Drosophila repleta group of species. Anim Behav 34: 421–429.
Fitzpatrick MJ, Ben-Shahar Y, Smid HM, Vet LEM, Robinson GE, Sokolowski MB (2005). Candidate genes for behavioural ecology. Trends Ecol Evol 20: 96–104.
Gleason JM (2005). Mutations and natural genetic variation in the courtship song of Drosophila. Behav Genet 35: 265–277.
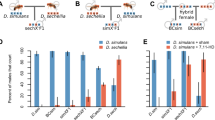
Gleason JM, Ritchie MG (2004). Do quantitative trait loci (QTL) for a courtship song difference between Drosophila simulans and D. sechellia coincide with candidate genes and intraspecific QTL? Genetics 166: 1303–1311.
Haag ES, True JR (2001). Perspective: From mutants to mechanisms? Assessing the candidate gene paradigm in evolutionary biology. Evolution 55: 1077–1084.
Hoikkala A (1988). The importance of different courtship stimuli in the mating behaviour of European species of the Drosophila virilis group. Ann Zool Fennici 25: 257–263.
Hoikkala A, Lumme J (1987). The genetic basis of evolution of the male courtship sounds in the Drosophila virilis group. Evolution 41: 827–845.
Hoikkala A, Lumme J (1990). Inheritance of male courtship sound characteristics in Drosophila littoralis. Behav Genet 20: 423–435.
Hoikkala A, Aspi J, Suvanto L (1998). Male courtship song frequency as an indicator of male genetic quality in an insect species, Drosophila montana. Proc R Soc Lond B 265: 503–508.
Hoikkala A, Mazzi D (2009). Evolution of complex acoustic signals in Drosophila species. In: Kim Y-K (ed). Handbook in Behavior Genetics. Springer Science+Business media: New York. pp 187–196.
Hoikkala A, Hoy R, Kaneshiro K (1989). High-frequency clicks of Hawaiian picture-winged Drosophila planitibia subgroup species. Anim Behav 47: 1363–1374.
Hoikkala A, Klappert K, Mazzi D (2005). Factors affecting male song evolution in Drosophila montana. Curr Top Dev Biol 67: 225–250.
Hoikkala A, Saarikettu M, Kotiaho JS, Liimatainen JO (2008). Age-related decrease in male reproductive success and song quality in Drosophila montana. Behav Ecol 19: 94–99.
Huttunen S, Aspi J (2003). Complex inheritance of male courtship song characters in Drosophila virilis. Behav Genet 33: 17–24.
Huttunen S, Aspi J, Hoikkala A, Schlötterer C (2004). QTL analysis of variation in male courtship song characters in Drosophila virilis. Heredity 92: 263–269.
Jones AG, Ratterman NL (2009). In the light of evolution III: two centuries of Darwin Sackler Colloquium: mate choice and sexual selection: what have we learned since Darwin? Proc Natl Acad Sci USA 106: 10001–10008.
Klappert K, Mazzi D, Hoikkala A, Ritchie MG (2007). Male courtship song and female preference variation between phylogeographically distinct populations of Drosophila montana. Evolution 61: 1481–1488.
Kyriacou CP (2002). Single gene mutations in Drosophila: what can they tell us about the evolution of sexual behaviour. Genetica 116: 197–202.
Kyriacou CP, Hall JC (1980). Circadian rhythm mutations in Drosophila melanogaster affect short-term fluctuations in the male′s courtship song. Proc Natl Acad Sci USA 77: 6729–6733.
Lander S, Green P, Abrahamson J, Barlow A, Daly M, Lincoln S, Newburg L (1987). MAPMAKER: an interactive computer package for constructing primary genetic linkage maps of experimental and natural populations. Genomics 1: 174–181.
Manichaikul A, Moon JY, Sen S, Yandell BS, Broman KW (2009). A model selection approach for the identification of quantitative trait loci in experimental crosses, allowing epistasis. Genetics 181: 1077–1086.
Mirol PM, Schäfer MA, Orsini L, Routtu J, Schlötterer C, Hoikkala A et al. (2007). Phylogeographic patterns in Drosophila montana. Mol Ecol 16: 1085–1097.
Miyatake T, Kanmiya K (2004). Male courtship song in circadian rhythm mutants of Bactrocera cucurbitae (Tephritidae: Diptera). J Insect Physiol 50: 85–91.
Morales-Hojas R, Päällysaho S, Vieira CP, Hoikkala A, Vieira J (2007). Comparative polytene chromosome maps of D. montana and D. virilis. Chromosoma 116: 21–27.
Nagelkerke NJD (1991). A note on a general definition of the coefficient of determination. Biometrika 78: 691–692.
Peixoto AA, Hall JC (1998). Analysis of temperature-sensitive mutants reveals new genes involved in the courtship song of Drosophila. Genetics 148: 827–838.
Qvarnström A, Bailey RI (2009). Speciation through evolution of sex-linked genes. Heredity 102: 2–15.
R Development Core Team (2008). R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing: Vienna, Austria.
Reinhold K (1998). Sex linkage among genes controlling sexually selected traits. Behav Ecol Sociobiol 44: 1–7.
Rideout EJ, Billeter JC, Goodwin SF (2007). The sex-determination genes fruitless and doublesex specify a neural substrate required for courtship song. Curr Biol 17: 1473–1478.
Ritchie MG (2007). Sexual selection and speciation. Annu Rev Ecol Evol Syst 38: 79–102.
Ritchie MG, Saarikettu M, Livingstone S, Hoikkala A (2001). Characterization of female preference functions for Drosophila montana courtship song and a test of the temperature coupling hypothesis. Evolution 55: 721–727.
Ritchie MG, Townhill RM, Hoikkala A (1998). Female preference for fly song: playback experiments confirm the targets of sexual selection. Anim Behav 56: 713–717.
Routtu J, Mazzi D, Van der Linde K, Mirol P, Butlin RK, Hoikkala A (2007). The extent of variation in male song, wing and genital characters among allopatric Drosophila montana populations. J Evol Biol 20: 1591–1601.
Schäfer MA, Mazzi D, Klappert K, Kauranen H, Vieira J, Hoikkala A, Ritchie MG, Schlötterer C (2010). A microsatellite linkage map for Drosophila montana shows large variation in recombination rates, and a courtship song trait maps to an area of low recombination. J Evol Biol 23: 518–527.
Stacklies W, Redestig H, Scholz M, Walther D, Selbig J (2007). pcaMethods: a bioconductor package providing PCA methods for incomplete data. Bioinformatics 23: 1164–1167.
Suvanto L, Liimatainen J, Tregenza T, Hoikkala A (2000). Courtship signals and mate choice of the flies of inbred Drosophila montana strains. J Evol Biol 13: 583–592.
Talyn BC, Dowse HB (2004). The role of courtship song in sexual selection and species recognition by female Drosophila melanogaster. Anim Behav 68: 1165–1180.
Villella A, Gailey DA, Berwald B, Oshima S, Barnes PT, Hall JC (1997). Extended reproductive roles of the fruitless gene in Drosophila melanogaster revealed by behavioral analysis of new fru mutants. Genetics 147: 1107–1130.
Wheeler DA, Kyriacou CP, Greenacre ML, You Q, Rutila JE, Rosbash M et al. (1991). Molecular transfer of a species-specific behavior from Drosophila simulans to Drosophila melanogaster. Science 251: 1082–1085.
Acknowledgements
This work was supported by the European Union through a Marie Curie Research Training Network (HPRN-CT-2002-00266). SY Wen acknowledges a Royal Society KC Wong International Incoming Fellowship Award. We thank S Nakagawa and two anonymous referees for their comments on the paper.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on Heredity website
Supplementary information
Rights and permissions
About this article
Cite this article
Lagisz, M., Wen, SY., Routtu, J. et al. Two distinct genomic regions, harbouring the period and fruitless genes, affect male courtship song in Drosophila montana. Heredity 108, 602–608 (2012). https://doi.org/10.1038/hdy.2011.129
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/hdy.2011.129