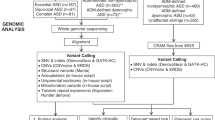
Abstract
Purpose
Developmental delay phenotypes have been associated with FMR1 premutation (PM: 55–200 CGG repeats) and “gray zone” (GZ: 45–54 CGG repeats) alleles. However, these associations have not been confirmed by larger studies to be useful in pediatric diagnostic or screening settings.
Methods
This study determined the prevalence of PM and GZ alleles in two independent cohorts of 19,076 pediatric referrals to developmental delay diagnostic testing through Victorian Clinical Genetics Service (cohort 1: N = 10,235; cohort 2: N = 8841), compared with two independent general population cohorts (newborn screening N = 1997; carrier screening by the Victorian Clinical Genetics Service prepair program N = 14,249).
Results
PM and GZ prevalence rates were not significantly increased (p > 0.05) in either developmental delay cohort (male PM: 0.12–0.22%; female PM: 0.26–0.33%; male GZ: 0.68–0.69%; female GZ: 1.59–2.13-%) compared with general population cohorts (male PM: 0.20%; female PM: 0.27–0.82%; male GZ: 0.79%; female GZ: 1.43–2.51%). Furthermore, CGG size distributions were comparable across datasets, with each having a modal value of 29 or 30 and ~1/3 females and ~1/5 males having at least one allele with ≤26 CGG repeats.
Conclusion
These data do not support the causative link between PM and GZ expansions and developmental-delay phenotypes in pediatric settings.
Similar content being viewed by others
Introduction
Neurological and neurodevelopmental disorders are associated with a trinucleotide CGG expansion of the FMR1 gene located on the long arm of the X chromosome.1 Full mutation (FM) CGG expansions of >200 repeats cause fragile X syndrome (FXS), whereby gene silencing leads to little or no production of the FMR1 protein (FMRP) that is essential for typical neurodevelopment.2 FXS is a common single-gene cause of inherited intellectual disability and autism spectrum disorder (ASD). CGG expansions of between 55 and 200 repeats, termed premutation (PM: ~1/200 females; ~1/800 males), are associated with an increased risk of having a child with FXS. This risk increases with the size of the PM CGG expansion in females, with some evidence suggesting that AGG interruptions may modify the chance of expansion.3
PM alleles are associated with fragile X–associated primary ovarian insufficiency (FXPOI), resulting in premature menopause in ~20% of women, and, among individuals over the age of 50, fragile X–associated tremor/ataxia syndrome (FXTAS) in up to 45% of PM males and 17% of PM females.4 Depressive and anxiety disorders are also associated with PM alleles.5,6
Additionally, smaller and more common CGG expansions between 45 and 54 CGG repeats, called “gray zone” or “intermediate” alleles (hereafter GZ: ~1/66 females, ~1/112 males) have been proposed to increase the risk of developing both FXTAS-like neurodegenerative disorders and FXPOI.7,8 However, this is controversial, as evidence of GZ-specific phenotypes is based on a few small studies that were impacted by selection bias.
The primary mechanisms underlying PM (and possibly GZ) presentations are postulated to be distinct from those involved in FXS. In PMs, the most important are aggregation of specific proteins mediated by overexpressed FMR1 mRNA, mitochondrial dysfunction, overexpression of long noncoding RNA of ASFMR1/FMR4, FMR5, and FMR6, and repeat associated non-ATG translation.9,10,11,12 However, decreased production of FMRP, previously considered unique to FXS, has also been described in individuals with PM alleles.13
Several smaller studies have provided some evidence for a FXS-like phenotype of ASD, developmental delay (DD), attention deficit hyperactivity disorder, and learning difficulties in proband children with PM and GZ alleles.14,15,16 However, these may also be explained by the presence of mosaicism for FM alleles on the background of PM or GZ alleles, which can be missed by standard diagnostic testing protocols, as well as possibly non-FMR1-related learning or behavioral disorders.17 Moreover, while a few prevalence studies have found enrichment of PM and GZ cases in cohorts with ASD and/or special needs,18,19 many others failed to replicate these findings, showing no association with neurodevelopmental disorders20,21,22,23 (as summarized in Supplementary Table S1 online).
Finally, a number of smaller studies associated alleles of ≤26 CGG repeats with an increased risk of developing fertility problems or behavioral problems and/or having children with developmental disability or a psychiatric illness, resembling to a degree the FMR1-related phenotypes, FXPOI and FXS.24,25 However, these finding are somewhat controversial owing to lack of replication by larger studies.
This study investigated pediatric patients with DD diagnostic test referrals (totaling ~19,000 cases) to determine the frequency of males and females with PM and GZ alleles, with statistical comparison to prevalence data from two population cohorts (newborn screening26 and population carrier screening27). The study hypotheses were that (i) PM and GZ frequencies are significantly enriched in children referred for DD testing as probands and (ii) CGG size distribution is different in a cohort of children referred for DD diagnostic testing compared with the general population.
Materials and methods
Study population
Between the 1980s and the time that this study was conducted, all clinician referrals for DD diagnostic testing at Victorian Clinical Genetics Services (VCGS) included FMR1 genetic testing as part of the standard protocol. The most common reason for referral was DD, followed by suspected intellectual disability, ASD, language delay, learning disorder, and/or FXS. The first DD cohort was sourced through the Medipath system at VCGS and comprised 10,235 pediatric DD referrals (≤18 years old) made between January 2003 and December 2009 (hereafter, DD #1 cohort). These samples are a proportion of a previously described larger cohort of individuals aged from <1 week to 89.9 years.28 Because the DD #1 cohort was sourced through an archived database, CGG size was not available. The second DD cohort included 8,841 pediatric DD referrals (≤18 years old) to VCGS between September 2013 and April 2017 (hereafter, DD #2 cohort). Data for this unpublished cohort were available through the Laboratory Information Management System (LabWare, Wilmington, DE) and included polymerase chain reaction (PCR)-based CGG sizes.29 Although the protocol of using PCR and Southern blot in FXS diagnostic testing has not changed significantly since 1992 at VCGS, chromosomal testing changed from conventional karyotyping to use of chromosomal microarray testing in 2012. This change to the protocol for DD testing at VCGS prevented pooling of the two DD cohorts. All cases identified as having an FM allele and those with pathogenic chromosome abnormalities identified elsewhere in the genome were part of the exclusion criteria for this study.
The first population cohort comprised 1997 newborns born between November 2009 and December 2010 at the John Hunter Hospital for whom consent for FMR1 CGG testing was given by a parent or guardian; the consent rate was 94%.26 The second population cohort comprised 14,249 females (17–57 years old) from the general population screened for FMR1 expansions as part of the VCGS prepair genetic-carrier screen (hereafter called the adult carrier-screening cohort).27 This program offers reproductive carrier screening to women for cystic fibrosis, FXS-associated disorders, and spinal muscular atrophy. Specific details of all four cohorts are provided in Supplementary Note S1 online.
Molecular testing protocol
First-line FMR1 testing at VCGS was performed on blood or saliva DNA. This was conducted using a fully validated PCR amplification assay with precision of ±1 repeat and limit of detection at 170 CGG repeats in males and 130 CGG repeats in females.29 Second-line confirmatory testing involved Southern blot analysis for amplified repeat sequences in the PM range, and inconclusive PCR results including “one peak” females and “no peak” males.30
All infants for whom samples were included in the newborn-screening cohort were born at John Hunter Hospital in Newcastle, Australia. Extra discs were punched from each child’s newborn-screening sample cards as part of a fragile X feasibility study by the NSW Newborn Screening Programme and Department of Molecular Genetics at the Children’s Hospital, Westmead, Australia. Two PCR methodologies were used to determine CGG size: (i) a modified PCR assay using a chimeric CGG-targeted primer31 and (ii) a standard PCR-based fragile X assay32 that was run in parallel to correlate with the chimeric primer assay. Alleles with <40 repeats were sized by nondenaturing capillary electrophoresis and alleles with ≥40 CGG repeats were sized using denaturing capillary electrophoresis.
The adult carrier-screening molecular testing was performed by triplet-primed PCR of the FMR1 CGG repeat region using the FMR1 TP-PCR commercial kit (Abbott Molecular, Lake Bluff, IL) or AmplideX FMR1 PCR kit (Asuragen, Austin, TX).33 Briefly, PCR products were denatured at 95 °C for two minutes after being mixed with a ROX 1000 size standard (Asuragen) and Hi-Di formamide (Thermo Fisher, Waltham, MA). These were then run on ABIPRISM 3730 capillary electrophoresis (Life Technologies, Foster City, CA) using POP-7 polymer (Life Technologies) with a 50-cm capillary, according to the manufacturer’s instructions. Samples that had expanded alleles showed the triplet repeat “stutter” pattern, with CGG sizing determined using Gene Mapper software version 5.0 (Life Technologies). All females identified as being in the PM range were reflexed for confirmatory testing at VCGS using PCR CGG sizing and Southern blot analysis.
Statistical analyses
The equality of the proportions of positive PM and GZ results was computed using Fisher’s exact test. All comparisons that were significant at p < 0.05 were then analyzed using pairwise comparisons, also using Fisher’s exact test, where results are presented before and after adjustment for multiple comparison using the false discovery rate (FDR). Binomial probability test was used to compare the proportions with inclusive and exclusive FMRI family history. Because of the small sample size, intergroup comparisons of CGG size in the PM range between the DD #2, adult carrier-screening, and newborn-screening cohorts were performed using a nonparametric Kruskal-Wallis test. All analyses were conducted using Stata version 13 (StataCorp, College Station, TX). PM was defined as CGG size 55–199; GZ was classified as 45–54 CGG repeats.
Results
The two DD cohorts were composed mainly of proband referrals who did not list knowledge of an FMR1 expansion in a blood relation (Table 1). Each DD cohort also included a proportion for whom no clinical notes were provided (DD #1: n = 494; DD #2: n = 440). These were included in the main analyses of proband data. In the adult carrier-screening cohort, knowledge of an FMR1 expansion in a blood relative was indicated on the test request form by a small number of females.
Frequency of males and females with positive FMR1 PM and GZ results
The PM and GZ frequencies were first determined after exclusion of males and females with a positive FMR1 family history, indicated either via clinical notes or on the adult carrier-screening test request form (Table 2). For males, there was no significant difference in PM and GZ frequency between the DD #1, DD #2, and newborn-screening cohorts (Table 2 and Figure 1).
Results correspond to Table 2. DD #1: pediatric DD referrals to VCGS between January 2003 and December 2009; DD #2: pediatric DD referrals to VCGS between September 2013 and April 2017. Population cohorts: newborn screening; adult carrier screening.
Pairwise comparison analyses of the female data showed a higher prevalence of PM females in the newborn-screening cohort versus both DD #1 (p = 0.035) and adult carrier-screening cohorts (p = 0.008) (Table 2 and Supplementary Table S2 online). However, after FDR adjustment for multiple comparisons, the difference between the newborn-screening and DD #1 cohorts was no longer significant. There was also a significant increase in the frequency of GZ females in the adult carrier cohort compared with both the newborn-screening (p = 0.032) and DD #2 cohorts (p = 0.008) (Table 2; Supplementary Table S2 online), with the latter remaining significant after FDR adjustment. Thus in the adult carrier-screening cohort, the number of PM females was depleted, yet that of GZ females was increased.
Importantly, there was no significant increase in the frequency of PM or GZ results in either DD cohort, for males or females (Table 2 and Figure 1). Repeat of all analyses excluding children referred with no clinical notes did not change the results.
Impact of family history of FMR1 expansion on prevalence estimates
To assess the impact of ascertainment bias on prevalence estimates, the frequency of males and females with PM and GZ results was next determined with inclusion of individuals who had indicated knowledge of an FMR1 expansion in a blood relation. With this change to the cohorts, male PM and GZ results remained comparable across the DD and newborn-screening cohorts (Supplementary Table S3 online).
However, for females, the addition of positive FMR1 family-history data eliminated the difference in female PM frequency between the DD #1 and newborn-screening cohorts, changing the p value from p = 0.035 to p = 0.324. It also reduced the difference in PM frequency found between the adult- and newborn-screening cohorts from p = 0.008 to p = 0.017, with the new p value not significant after FDR adjustment (Supplementary Tables S3 and S4 online). This analysis shows that the underrepresentation of females with PM alleles in the adult carrier-screening cohort compared with the newborn-screening cohort (reported in Table 2) could be artificially related to ascertainment bias.
By contrast, positive FMR1 family-history data did not influence the differences in female GZ prevalence found between adult- and newborn-screening cohorts (p = 0.032). Similarly, the increase of GZ females in the adult carrier screening versus the DD #2 cohort remained statistically significant after FDR adjustment (p = 0.008) (Supplementary Tables S3 and S4 online). Therefore it is unlikely that the greater prevalence of GZ females in the adult carrier-screening cohort (shown in Table 2) is related to an impact of ascertainment bias.
Finally, binomial probability tests showed no effect from including the positive FMR1 family-history data in the analyses, on male and GZ prevalence results. However, in the DD #1 cohort female PMs were more common when the family-history data were included (0.5%, or 1 in 199), compared with when they were removed (0.3%; or 1 in 392) (p = 0.023), although this was not significant after FDR adjustment (Supplementary Table S5 online). A comparison of estimated prevalence rates with and without a positive family history is presented in Supplementary Table S6 online.
FMR1 PM CGG size distribution
The median PM CGG size in the DD #2 cohort was CGG 57 repeats (males and females). This was not significantly different from the median CGG size in the newborn screening (CGG: 59.5 repeats) and adult carrier screening (CGG: 61 repeats) cohorts. Analyses of males and females separately did not change these results (Table 3).
CGG distribution plots and low normal allele prevalence
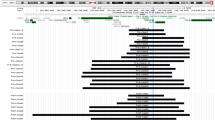
Male and female CGG-distribution plots were created for cohorts where CGG size was available (i.e., all except DD #1) (Figure 2). Because the focus of these analyses is to understand the lower end of the CGG size distribution and any differences in the “low normal” allele prevalence across the cohorts, the smaller of the two alleles in females was used for the distribution plots in Figure 2. The other allele is presented in Supplementary Figure S1 online. For each plot the modal value was 29 or 30. In both the DD #2 and adult carrier-screening cohorts, minor peaks at CGG 20 and 23 repeats were observed. These peaks were smaller in males than in females, with the minor peak at CGG 20 reaching ~5% in the male DD #2 cohort, versus ~10% in the female DD #2 and 15% in the adult carrier-screening cohorts.
Figures show male data from (a) DD #2 (N = 6670) and (b) newborn screening (n = 1016). Panels c–e show female data (smaller of two alleles): (c): DD #2 (N = 2168); (d): newborn screening (N = 981); (e): adult carrier screening (N = 14,239). Sample size is slightly different from what is reported in Table 1, as these data do not include results with >70 CGG repeats. The larger female allele is shown in Supplementary Figure S1 online. DD, developmental delay.
An analysis of the proportion of individuals with at least one “low normal” CGG repeat size (i.e., ≤26 repeats) was performed (Supplementary Tables S7 and S8 online). For males, the “low normal” CGG size was found in 19.0% of the DD #2 cohort and 21.3% of the newborn-screening cohort, but this difference was not significantly different (p = 0.09). In females, similar proportions were found in the three cohorts (p = 0.61): 35.0% of the DD #2, 33.6% of the newborn-screening, and 35.0% of the adult carrier-screening cohort. Thus there were very similar proportions across the cohorts of males and females with at least one X chromosome with the “low normal” CGG size allele. Proportions for females with two copies of a “low normal” allele (i.e., both alleles have ≤26 repeats) were: 3.0% (DD #2); 5.4% (newborn screening); and 4.1% (adult carrier screening). After FDR adjustment, significant differences remained between the DD #2 prevalence and both the newborn-screening (p = 0.002) and adult carrier-screening (p = 0.011) cohorts (Supplementary Table S8 online).
Discussion
The present investigation did not find an enrichment of children with PM and GZ alleles in pediatric DD diagnostic referrals to VCGS, compared with two population cohorts.26,27 Based on the main analyses, which excluded individuals indicating knowledge of an FMR1 expansion in a blood relation, the prevalence of PM males in the DD cohorts in Australia was estimated at between 1/461 and 1/830. This is similar to prevalence estimates from the newborn-screening cohort (1/507 males) and those from other groups that have analyzed Caucasian populations in other countries.34,35 The proportion of PM females is also similar in DD and population cohorts (~1/306–392), although slightly reduced when compared with prevalence estimates in US populations.34,36 There was also no significant difference in GZ frequency in DD versus population cohorts for males and females, excluding an increase of GZ females in the adult carrier-screening cohort versus the DD #2 cohort.
Given the lack of enrichment of PM and GZ males and females in the DD cohorts, there is unlikely to be a clinical phenotype associated with these alleles in children <18 years old that is significant enough to warrant clinician referral for further DD diagnostic testing. It is therefore unlikely that PM and GZ alleles are a cause of the conditions that are common reasons for clinician referral, such as DD, but also intellectual disability, ASD, language delay, and learning disorder.
The present findings support the results from smaller prevalence studies that also did not find any elevation in PM or GZ frequencies in cohorts with indicated DD and/or ASD.20,21,22,23 Together, this information challenges the findings from other smaller cohort prevalence studies that have reported a significant excess of developmental problems in children with PM or GZ results.18,19 In light of this, generalizability of results from a questionnaire-based study that found an excess of developmental problems in children with PM and GZ alleles but did not perform confirmatory genetic testing is also questionable.37 Furthermore, a hospital linkage analyses study38 and case studies that reported FXS-like features in male PM and GZ probands may have been impacted by selection bias.14,15,16 Differences between studies that have investigated the association between the PM and GZ alleles and developmental problems, which may have contributed to inconsistency in the literature, include (i) sample size and cohort characteristics; (ii) change over time in awareness of FMR1-related disorders by clinicians and availability of diagnostic methodologies; (iii) different definitions of PM/GZ repeat size and different ethnicities; and (iv) use of maternal allele(s) versus an independent typically developing group for control comparisons. Moreover, given that recent studies suggest that mosaicism is more common than previously thought,17 and difficult to detect using standard testing protocols because of test-sensitivity issues or deletion of PCR primer binding sites associated with somatic mosaicism,39,40 it is also plausible that some studies may have unknowingly included individuals who were mosaic for PM and FM, or GZ and FM, which could have skewed the results and interpretation.
This study investigated whether clinical impact in PM children could be driven statistically by a subgroup of individuals who have a very large CGG repeat size that is within the PM range (most individuals have <70 CGG repeats).34 Given that FMR1 alleles with >80 CGG repeats have been associated with increased risk of developing FXTAS and FXPOI,6 they may also be related to neurodevelopmental impact in the pediatric setting. However, the median PM CGG size was very comparable among all cohorts analyzed in this study, indicating no link between the larger CGG size and clinician concerns over DD or other common reasons for DD diagnostic testing (e.g., ASD).
The proportion of individuals with the “low normal” allele (≤26 CGG repeats) was similar in the DD #2 cohort and both adult carrier and newborn-screening cohorts, consistent with no association between these alleles and neurodevelopmental disorders. Specifically, ~1/3 females and 1/5 males in each cohort had at least one “low normal” allele (~1/ 18–33 females had two copies of these “low normal” alleles). Given the large sample size and replication of proportions across clinical and nonclinical cohorts, the proportions reported in this study are likely to be robust and thus may be useful as a benchmark for future comparisons in investigations of the “low normal” CGG size.
The findings should be interpreted in light of the following limitations. This study cannot comment on the severity of the phenotype in PM and GZ children identified in the DD cohorts, nor can it comment on the involvement of PM and GZ alleles in subtle phenotypes that are unlikely to prompt clinician requests for DD diagnostic testing, such as executive dysfunction, which can impact planning and organizational skills. Indeed, there is emerging evidence that adults with PM alleles are at greater risk of developing subtle visuospatial problems that resemble dorsal stream-processing vulnerability,6 plus strong evidence that adult PM females can have elevated, but often sub-threshold, symptoms of social anxiety disorder and major depressive disorder5 that may or may not be linked to FXPOI and/or FXTAS.
There is also a possibility of bias in the adult carrier-screening cohort due to (i) higher socioeconomic status, as the carrier-screening test is predominantly offered by private obstetricians on a test-pay basis, and (ii) PM females who do not have social anxiety or menopausal/infertility problems being more likely to use the adult carrier-screening service.27 Other study limitations include not having the resolution on the nondenaturing capillary electrophoresis to determine whether minor peaks at 20 and 23 were present in the newborn-screening cohort and the possibility that accurate clinical and family-history information was not always provided on the clinician test request form.
In conclusion, there was no statistically significant increase in the frequency of males and females with expanded alleles in the DD clinician referral cohorts compared with newborn and population carrier-screening cohorts. This questions the impact of PM and GZ expansions on FXS-like phenotypes in children, such as DD and ASD. This study also queries the clinical relevance of the recently described “low normal” allele in neurodevelopmental disorders. This is likely to be of interest to families that may have or plan to have children with a PM and GZ allele, such as those identified in population carrier screening or through cascade testing. The present findings also favor not testing for PM and GZ alleles in newborns, owing to lack of sufficient evidence of clinically significant neurodevelopmental impact on these children in the pediatric clinical setting and the potential to identify adult-onset conditions of incomplete penetrance associated with PM alleles.
References
Verkerk AJ, Pieretti M, Sutcliffe JS et al. Identification of a gene (FMR-1) containing a CGG repeat coincident with a breakpoint cluster region exhibiting length variation in fragile X syndrome. Cell 1991;65:905–914.
Hagerman R, Lauterborn J, Au J & Berry-Kravis E. Fragile X syndrome and targeted treatment trials. Results Probl Cell Differ 2012;54:297–335.
Volle CB & Delaney S. AGG/CCT interruptions affect nucleosome formation and positioning of healthy-length CGG/CCG triplet repeats. BMC Biochem 2013;14:14–33.
Rodriguez-Revenga L, Madrigal I, Pagonabarraga J et al. Penetrance of FMR1 premutation associated pathologies in fragile X syndrome families. Eur J Hum Genet 2009;17:1359–1362.
Roberts JE, Tonnsen BL, McCary LM, Ford AL, Golden RN & Bailey DB Jr.. Trajectory and predictors of depression and anxiety disorders in mothers with the FMR1 premutation. Biol Psychiatry 2015;79:850–857.
Kraan CM, Hocking DR, Bradshaw JL et al. Neurobehavioural evidence for the involvement of the FMR1 gene in female carriers of fragile X syndrome. Neurosci Biobehav Rev 2013;37:522–547.
Bodega B, Bione S, Dalpra L et al. Influence of intermediate and uninterrupted FMR1 CGG expansions in premature ovarian failure manifestation. Hum Reprod 2006;21:952–957.
Loesch DZ, Khaniani MS, Slater HR et al. Small CGG repeat expansion alleles of FMR1 gene are associated with parkinsonism. Clin Genet 2009;76:471–476.
Todd Peter K, Oh Seok Y, Krans A et al. CGG repeat-associated translation mediates neurodegeneration in fragile X tremor ataxia syndrome. Neuron 2013;78:440–455.
Arocena DG, Iwahashi CK, Won N et al. Induction of inclusion formation and disruption of lamin A/C structure by premutation CGG-repeat RNA in human cultured neural cells. Hum Mol Genet 2005;14:3661–3671.
Loesch DZ, Godler DE, Evans A et al. Evidence for the toxicity of bidirectional transcripts and mitochondrial dysfunction in blood associated with small CGG expansions in the FMR1 gene in patients with parkinsonism. Genet Med 2011;13:392–399.
Pastori C, Peschansky VJ, Barbouth D, Mehta A, Silva JP & Wahlestedt C. Comprehensive analysis of the transcriptional landscape of the human FMR1 gene reveals two new long noncoding RNAs differentially expressed in Fragile X syndrome and Fragile X-associated tremor/ataxia syndrome. Hum Genet 2014;133:59–67.
Ludwig AL, Espinal GM, Pretto D et al. CNS expression of murine fragile X protein (FMRP) as a function of CGG-repeat size. Hum Mol Genet 2014;23:3228–3238.
Farzin F, Perry H, Hessl D et al. Autism spectrum disorders and attention-deficit/hyperactivity disorder in boys with the fragile X premutation. J Dev Behav Pediatr 2006;27(2 Suppl):S137–S144.
Clifford S, Dissanayake C, Bui QM, Huggins R, Taylor AK & Loesch DZ. Autism spectrum phenotype in males and females with fragile X full mutation and premutation. J Autism Dev Disord 2007;37:738–747.
Goodlin-Jones BL, Tassone F, Gane LW & Hagerman RJ. Autistic spectrum disorder and the fragile X premutation. J Dev Behav Pediatr 2004;25:392–398.
Aliaga SM, Slater HR, Francis D et al. Identification of males with cryptic fragile X alleles by methylation-specific quantitative melt analysis. Clin Chem 2016;62:343–352.
Youings SA, Murray A, Dennis N et al. FRAXA and FRAXE: the results of a five year survey. J Med Genet 2000;37:415–421.
Murray A, Youings S, Dennis N et al. Population screening at the FRAXA and FRAXE loci: molecular analyses of boys with learning difficulties and their mothers. Hum Mol Genet 1996;5:727–735.
Tassone F, Choudhary NS, Durbin-Johnson B, Hansen R, Hertz-Picciotto I & Pessah I. Identification of expanded alleles of the FMR1 gene in the CHildhood Autism Risks from Genes and Environment (CHARGE) study. J Autism Dev Disord 2013;43:530–539.
Essop FB & Krause A. Diagnostic, carrier and prenatal genetic testing for fragile X syndrome and other FMR-1-related disorders in Johannesburg, South Africa: A 20-year review. S Afr Med J 2013;103:994–998.
Madrigal I, Xuncla M, Tejada MI et al. Intermediate FMR1 alleles and cognitive and/or behavioural phenotypes. Eur J Hum Genet 2011;19:921–923.
Otsuka S, Sakamoto Y, Siomi H et al. Fragile X carrier screening and FMR1 allele distribution in the Japanese population. Brain Dev 2010;32:110–114.
Mailick MR, Hong J, Rathouz P et al. Low-normal FMR1 CGG repeat length: phenotypic associations. Front Genet 2014;5:309.
Weghofer A, Kim A, Barad DH & Gleicher N. The impact of androgen metabolism and FMR1 genotypes on pregnancy potential in women with dehydroepiandrosterone (DHEA) supplementation. Hum Reprod 2012;27:3287–3293.
Christie L, Wotton T, Bennetts B et al. Maternal attitudes to newborn screening for fragile X syndrome. Am J Med Genet A 2013;161a:301–311.
Archibald AD, Smith MJ, Burgess T et al. Reproductive genetic carrier screening for cystic fibrosis, spinal muscular atrophy and fragile X syndrome in Australia: outcomes of 12,000 tests. Genet Med; e-pub ahead of print 26 October 2017.
Cotter M, Archibald AD, McClaren BJ et al. Clinical audit of genetic testing and referral patterns for fragile X and associated conditions. Am J Med Genet A 2016;170:1439–1449.
Khaniani MS, Kalitsis P, Burgess T & Slater HR. An improved diagnostic PCR assay for identification of cryptic heterozygosity for CGG triplet repeat alleles in the fragile X gene (FMR1). Mol Cytogenet 2008;1:5.
Francis D, Burgess T, Mitchell J & Slater H. Identification of small FRAXA premutations. Mol Diagn 2000;5:221–225.
Tassone F, Pan R, Amiri K, Taylor AK & Hagerman PJ. A rapid polymerase chain reaction-based screening method for identification of all expanded alleles of the fragile X (FMR1) gene in newborn and high-risk populations. J Mol Diagn 2008;10:43–49.
Fu YH, Kuhl DP, Pizzuti A et al. Variation of the CGG repeat at the fragile X site results in genetic instability: resolution of the Sherman paradox. Cell 1991;67:1047–1058.
Hantash FM, Goos DG, Tsao D et al. Qualitative assessment of FMR1 (CGG)n triplet repeat status in normal, intermediate, premutation, full mutation, and mosaic carriers in both sexes: implications for fragile X syndrome carrier and newborn screening. Genet Med 2010;12:162–173.
Tassone F, Iong KP, Tong TH et al. FMR1 CGG allele size and prevalence ascertained through newborn screening in the United States. Genome Med 2012;4:100.
Dombrowski C, Levesque S, Morel ML, Rouillard P, Morgan K & Rousseau F. Premutation and intermediate-size FMR1 alleles in 10572 males from the general population: loss of an AGG interruption is a late event in the generation of fragile X syndrome alleles. Hum Mol Genet 2002;11:371–378.
Cronister A, Teicher J, Rohlfs EM, Donnenfeld A & Hallam S. Prevalence and instability of fragile X alleles: implications for offering fragile X prenatal diagnosis. Obstet Gynecol 2008;111:596–601.
Bailey DB Jr., Raspa M, Olmsted M & Holiday DB. Co-occurring conditions associated with FMR1 gene variations: findings from a national parent survey. Am J Med Genet A 2008;146A:2060–2069.
Renda MM, Voigt RG, Babovic-Vuksanovic D et al. Neurodevelopmental disabilities in children with intermediate and premutation range fragile X cytosine-guanine-guanine expansions. J Child Neurol 2012;29:326–330.
Hwang YT, Aliaga SM, Arpone M et al. Partially methylated alleles, microdeletion, and tissue mosaicism in a fragile X male with tremor and ataxia at 30 years of age: a case report. Am J Med Genet A 2016;170:3327–3332.
Jiraanont P, Kumar M, Tang HT et al. Size and methylation mosaicism in males with fragile X syndrome. Expert Rev Mol Diagn 2017;17:1023–1032.
Acknowledgments
This study was supported by an NHMRC Early Career fellowship to C.M.K. (no. 1112934); the Victorian Government’s Operational Infrastructure Support Program, with the salaries supported by NHMRC project grants (nos. 104299 and1103389 to D.E.G.); Murdoch Children’s Research Institute, Royal Children’s Hospital Foundation (D.E.G.); the Financial Markets Foundation for Children (Australia) (nos. 2017–361 to D.E.G., C.M.K., and D.J.A.); the Genetics of Learning Disability (GOLD) Service (T.L.W., M.F., and L.M.C.); and John Kelleher from the Fragile X Association of Australia (T.L.W.). The authors are grateful to the mothers and the staff at the John Hunter Hospital, Newcastle, Australia, for their participation in and support of the study. We also thank Richard Wesseling for designing the Access database for the collection of data and for assisting with the extraction of data for evaluation. Finally, we thank the members of the prepair carrier-screening laboratory team (Alisha Monaco, Karina Sandoval, Lisa Ward, and Melissa Chow).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Disclosure
The authors declare no conflict of interest.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Kraan, C.M., Bui, Q.M., Field, M. et al. FMR1 allele size distribution in 35,000 males and females: a comparison of developmental delay and general population cohorts. Genet Med 20, 1627–1634 (2018). https://doi.org/10.1038/gim.2018.52
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/gim.2018.52
Key Words
This article is cited by
-
Maternal FMR1 alleles expansion in newborns during transmission: a prospective cohort study
Pediatric Research (2023)
-
Family history of FXTAS is associated with age-related cognitive-linguistic decline among mothers with the FMR1 premutation
Journal of Neurodevelopmental Disorders (2022)
-
Cortical gyrification and its relationships with molecular measures and cognition in children with the FMR1 premutation
Scientific Reports (2020)
-
Abnormally Methylated FMR1 in Absence of a Detectable Full Mutation in a U.S.A Patient Cohort Referred for Fragile X Testing
Scientific Reports (2019)
-
Fragile X syndrome carrier screening in pregnant women in Chinese Han population
Scientific Reports (2019)