Abstract
Purpose
The implementation of newborn screening for lysosomal disorders has uncovered overall poor specificity, psychosocial harm experienced by caregivers, and costly follow-up testing of false-positive cases. We report an informatics solution proven to minimize these issues.
Methods
The Kentucky Department for Public Health outsourced testing for mucopolysaccharidosis type I (MPS I) and Pompe disease, conditions recently added to the recommended uniform screening panel, plus Krabbe disease, which was added by legislative mandate. A total of 55,161 specimens were collected from infants born over 1 year starting from February 2016. Testing by tandem mass spectrometry was integrated with multivariate pattern recognition software (Collaborative Laboratory Integrated Reports), which is freely available to newborn screening programs for selection of cases for which a biochemical second-tier test is needed.
Results
Of five presumptive positive cases, one was affected with infantile Krabbe disease, two with Pompe disease, and one with MPS I. The remaining case was a heterozygote for the latter condition. The false-positive rate was 0.0018% and the positive predictive value was 80%.
Conclusion
Postanalytical interpretive tools can drastically reduce false-positive outcomes, with preliminary evidence of no greater risk of false-negative events, still to be verified by long-term surveillance.
Similar content being viewed by others
Introduction
Newborn screening is a public health program aimed at the identification of conditions for which early intervention can prevent mortality, morbidity, and disabilities,1 but is not without its challenges.2 In recent years, expansions of testing panels have been proposed and adopted either as national standards3 or as nonbinding recommendations4 by the US Department of Health and Human Services (HHS). Currently, the recommended uniform screening panel encompasses 34 conditions. The HHS Secretary’s Advisory Committee on Heritable Disorders in Newborns and Children is tasked with overseeing the process5 of adding emerging conditions6,7 to the recommended panel. The most recent additions are acid α-glucosidase (GAA) deficiency (Pompe disease),8 α-l-iduronidase (IDUA) deficiency (MPS I),9 and X-linked adrenoleukodystrophy.10 Other lysosomal disorders, particularly galactocerebrosidase (GALC) deficiency (Krabbe disease),11 have been turned down by the committee because they lacked evidence of net benefits. However, advocacy efforts and legislative mandates have propelled six states to begin screening for Krabbe disease and other lysosomal disorders,12,13,14 but reports of outcomes and performance have not been encouraging.15,16,17
In 2015, the legislature of the Commonwealth of Kentucky passed bill KRS 214.155, mandating screening for Krabbe disease. This action was driven by an advocacy campaign led by the parents of a child affected with the infantile form of the disease. To accelerate implementation, the Kentucky Department for Public Health reached out to the Biochemical Genetics Laboratory at Mayo Clinic (Rochester, Minnesota) to negotiate outsourcing of screening for Krabbe disease by measuring only GALC activity. This assay had to be performed in parallel to local testing for all other conditions and be completed within 24 hours to avoid any further delays in the care of early-onset cases.18 This request was not fulfilled because conventional interpretation methods of a single marker (cutoff, percent of daily mean) are not suited to differentiation of affected patients from individuals who are either heterozygous or carry pseudo-deficiency alleles.12,13,14,15 The laboratory made a counter-proposal to perform a profile of six lysosomal enzyme activities using commercially available substrates, inclusive of the primary markers of Pompe disease and MPS I, integrated with additional analytes and second-tier tests as needed. This new plan, characterized by the expanded scope of testing and reporting of three conditions, was agreeable to both Kentucky and Mayo Clinic. The solution adopted here as primary screening was to rely on six covariate-adjusted enzyme activities integrated by all informative permutations of calculated ratios among them.
Materials and methods
Reference population and patients
Supplementary Figure 1 online shows the size of the cumulative reference population with and without covariate (age at collection and body weight) correction for GALC activity, which was compiled by the end of the first year of testing. At the beginning of prospective testing, reference data from the pilot study mentioned below were also used, but were phased out when approximately 10,000 prospective cases had been tested. The data sets for acid sphingomyelinase, GAA, α-galactosidase, β-glucocerebrosidase, and IDUA were of comparable sizes. To create postanalytical interpretive tools, enzyme profiles of true- and false-positive cases were available from previous clinical testing (confirmed true positives) and unpublished research studies (presumed true positives). Confirmed true positives were all proven by genotyping to carry known pathogenic variants in trans but were not sorted by predicted genotype–phenotype correlation. Cases categorized as presumed false positives were either heterozygous for pathogenic variants or carrying known pseudodeficiency alleles. With approval from the Mayo Clinic’s institutional review board (protocol 09-006866, “Development and validation of newborn screening assays to measure biomarkers and enzyme levels in biological samples from patients with a lysosomal storage disorder and their relatives,” replaced by protocol 15-005393, “Development and validation of clinical biochemical genetic assays associated with inborn errors of metabolism and other heritable or congenital conditions using biospecimens collected from affected patients and their immediate family members”), the Biochemical Genetics Laboratory actively seeks parental permission to retrieve and analyze retrospectively the original newborn screening card of all new cases we diagnose clinically as a combination of biochemical and molecular tests, a situation we encounter fairly frequently, being a worldwide referral laboratory. A total of 86 cases have been enrolled. Twelve presumed true positives and all but one of the presumed false positives were identified through a National Institute of Child Health and Human Development–funded pilot study of 100,000 anonymized specimens from the California Department of Public Health (HHSN275201000017C, principal investigator Dietrich Matern, unpublished data) By the end of the first year of prospective testing, the counts were as follows: Krabbe disease N = 11, false-positive GALC N = 61, Pompe disease N = 40, false-positive GAA N = 75, MPS I N = 6, false-positive IDUA N = 99.
Study population and analytical methods
A total of 55,161 Kentucky specimens were received at Mayo Clinic between 17 February 2016 and 18 February 2017. For most cases, a full spot was separated from the original card at the same time as testing was initiated locally. Batches were shipped 6 days per week by overnight service. Residual samples were returned after testing was completed. Six enzyme activities were measured simultaneously by flow-infusion tandem mass spectrometry (six-(multi)plex assay) as recently described.19 When a repeat analysis was indicated to investigate an initial abnormal result, the concentrations of C20–C26 lysophosphatidylcholines were also measured (cumulative 10-plex assay).19 Enzyme substrates were purchased from PerkinElmer Life Sciences (Shelton, CT).20 Testing showed adequate stability and reproducibility over time (coefficient of variation range: 7–15%; Supplementary Figure 2). Two of the biochemical methods used for second-tier tests (Krabbe disease: psychosine concentration, MPS I: dermatan sulfate and heparan sulfate concentrations) have been described previously.21,22 A second-tier test for the evaluation of low GAA activity was developed during this study and has been described separately.23 It is based on a 12-plex panel inclusive of creatine and creatinine, utilized to calculate the (creatine/creatinine)/GAA ratio that is incorporated in expanded CLIR tools. Molecular testing of the GALC common 30-kb deletion and Sanger sequencing of the GALC, IDUA, and GAA genes were performed using clinically available tests.
Multivariate pattern recognition software
Collaborative Laboratory Integrated Reports (CLIR; https://clir.mayo.edu) is a Web application that maintains an interactive database of laboratory results from multiple sites. CLIR was originally developed to support Region 4 Stork (https://www.clir-r4s.org/), a collaborative project that started in 2004, aiming at performance improvement of newborn screening by tandem mass spectrometry.24 In that project cumulative reference intervals of amino acids and acylcarnitines were used to create postanalytical interpretive tools,25,26 which integrate all informative results into a single score and compare it to known confirmed cases to return a percentile ranking as a measure of likelihood of disease. The tools are applicable to either the diagnosis of one condition or, like the dual scatter plots described in this report, to the differential diagnosis between two conditions with overlapping phenotypes (i.e., true positives versus false positives).25,27 Briefly, in the dual scatter plot the relationship to reference ranges becomes irrelevant as the comparison now takes place between two disease ranges. The dual scatter plot is also the combination of two tools, one that targets any nonoverlapping result to increase the score for the first condition and decrease the score for the second one, and another that operates in exactly the opposite way. A result within the overlap range triggers no score variation for either condition; a result above the overlap range increases the score of the first tool and has the opposite effect on the second tool. A result below the overlap range operates in the opposite way. So, a completely normal result could actually provide information that would help to achieve the desired differential diagnosis, even if the same result would not trigger any score in a one-condition tool for either condition. Another characteristic of the dual scatter plot is a different way to express the calculated scores. Instead of absolute values, both axes use a process of normalization that keeps the scores between 0 and 100. Each result is calculated by subtracting from the score the lowest of all scores, dividing the result by the range of values (highest minus lowest), and then multiplying by 100. This formula preserves the relative distance between values and is ideal for achieving consistency among tools comparing any two conditions with different numbers of informative markers. The output of the dual scatter plot is a visual separation of the combined scores in four quadrants. The lower right quadrant includes the cluster of cases with the first condition, those with a high score using one tool and a low score with the other. The upper left quadrant includes the scores of cases with the second condition. A score located in the upper right quadrant is equivalent to an inconclusive result, meaning that both conditions are still possible. Finally, a score in the left bottom quadrant excludes both conditions. When used to investigate an unknown case, the coordinates of the combined scores of a particular case are shown as a red diamond.
CLIR’s defining characteristics are (i) the replacement of analyte cutoff values with condition-specific degree of overlap between cumulative reference and disease ranges, and (ii) the integration of primary markers with all informative permutations of ratios.24 Ratios calculated between markers not directly related at the biochemical level are particularly helpful in correcting for preanalytical factors and potential analytical bias. An additional and unique feature of CLIR is the replacement of conventional reference intervals with continuous, covariate-adjusted, moving percentiles.28 Utilization of CLIR is freely available to qualified users worldwide willing to share reference data and profiles of positive cases in advance of being given access, with the goal of sustaining a constantly evolving, and improving, clinical validation. Code access, however, is not provided to external users.
Covariate adjustment
All lysosomal enzyme activities exhibited marked variation across two continuous covariates within the newborn population: age at collection and birth weight. To incorporate this variation into routine data analysis, a statistical normalization technique was conducted using a regression-based method.28 This process resulted in a transformation that could be applied to the entire (55,161) Kentucky specimen population. Reference data from multiple contributing labs were analyzed for homogeneity (see Supplementary Figure 1), and if within a bias criterion based on biological variation,28 they were integrated into a cumulative data set binned across the covariate range to elicit any trend. If necessary, the marker and/or covariate values were adapted using a Box–Cox power transformation.29 Median and standard-deviation estimates of each bin were then calculated and polynomial regression models were fit to both of the binned estimates. From these, Z-scores were calculated for all reference data and prospective cases data to normalize marker values across the entire covariate range. An example of a regression model for the GALC enzyme and birth weight as covariate is shown in Supplementary Figure 3. Based on the resulting changes in degree of overlap between reference and disease ranges in both populations, true positives and false positives, tools were created using the adjustment for the single covariate that overall showed a greater separation for true positives and a greater overlap for false positives. The chosen single covariate adjustments for GAA, GALC, and IDUA were age, birth weight, and age, respectively.
Results
The ranges of residual enzyme activities in true- and false-positive cases for lysosomal disorders overlap almost completely, even after covariate adjustment (Supplementary Figure 1). A detailed comparison of three enzyme activities (GALC, GAA, and IDUA) in true-positive and false-positive cases is presented in Supplementary Figures 4 and 5. Based on such evidence, a postanalytical interpretive workflow (Figure 1) was implemented prospectively after February 2016 to deliver newborn screening for Krabbe disease, MPS I, and Pompe disease. To minimize the risk of overlooking affected cases, the 6-plex primary screening was set to resolve as negative only cases with a completely normal profile, i.e., all potentially informative markers were within the 1–99% percentile of the respective reference range.24 If an affected patient was to be found later by clinical ascertainment among the cases reported as screen negative, it would represent a true false-negative event.30 The single-condition tools identified as positive 181, 76, and 397 cases for Krabbe disease, MPS I, and Pompe disease, respectively (Figure 1). The dual scatter plot, a previously reported instrument of differential diagnosis,25 was the next step of the workflow designed to accomplish a segregation between potential true-positive and false-positive cases. These tools assigned either “not informative” or “artifact” status to 642 (98%) of the cases picked up initially, leaving only 12 cases requiring further laboratory evaluation of the initial blood-spot specimen. If no further testing was done by repeat analysis and second-tier tests, the false-positive rate would have been 0.01% (8/55,161) and the positive predictive value 33% (4/12).
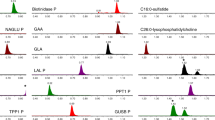
Repeat analysis with the extended 10-plex assay integrated with second-tier tests resolved eight of the remaining cases as false positives (Table 1), with one exception, described below. Figure 2 shows the dual scatter plots (based on either 6 or 10 markers) for the differential diagnosis between Krabbe disease and confirmed false-positive cases. The particular case displayed in this plot shows an indeterminate outcome25 by 6-plex assay, that improves in determinacy to “consistent with Krabbe disease” by 10-plex assay, forming an illustrative example of the incremental value of adding more markers. As a cautionary validation measure, laboratory directors had the discretion to override a not informative resolution by the dual scatter plot and proceed with repeat testing and/or second-tier testing (Table 1), but no discrepancies were observed. 982 samples of this cohort (Supplementary Figure 1) revealed a profile in which more than one of the six enzyme activities tested abnormally low, presumably resulting from preanalytical interferences during sample collection, handling, and/or transport. An added benefit of the 6-plex assay was the ability it afforded to recognize these preanalytical artifacts. even when only a single activity among the three primary targets was low. All cases with a profile showing two or more low enzyme activities were deemed of no clinical relevance and reported as screen negative after verification using dedicated dual scatter plots between a given condition and the cases previously classified as artifacts, which were added to the database on a weekly basis.
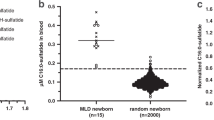
Dual scatter plots for the differential diagnosis between Krabbe disease and false-positive cases. (a) 6-plex dual scatter plot; (b) 10-plex dual scatter plot. Each plot is divided into four quadrants. Lower right: consistent with Krabbe disease (light blue squares and circles); upper right: indeterminate (both conditions are possible); upper left: consistent with false-positive cases (purple squares and circles); lower left: neither condition. Red diamonds indicate the score of one confirmed Krabbe patient not diagnosed as part of this study. Educational material describing the principle of a min-max score and the creation and use of this tool is available online.40 FP GALC, false-positive result for galactocerebrosidase deficiency.
Abnormal cases
Table 2 summarizes the demographic information, results, and genotypes of the five cases reported as abnormal. Patient 2 was an exception made to our protocol: a referral was initiated because of very low IDUA activity, despite normal concentrations of dermatan sulfate (Table 2) measured by the second-tier test. A repeat sample was requested and showed again low IDUA activity and a normal second-tier test. Clinical evaluation still ensued and included genotyping, revealing heterozygosity for a known pathogenic variant (Table 2). Even if no further steps were taken, the fact that additional contact with the patient had occurred to collect a repeat sample meets the designation of a false-positive case.31 On the other hand, the prospective screening of a full sibling of the patient behind the legislative mandate in Kentucky, who was diagnosed prenatally to be a carrier for Krabbe disease, was correctly reported as screen-negative.
In terms of outcome, the case diagnosed with MPS I received a bone-marrow transplant at approximately 6 months of age. The two Pompe cases are evaluated every 3 months at the University of Kentucky in Lexington. While asymptomatic, neither is receiving enzyme replacement therapy and testing of siblings revealed in one family an older sister with the same genotype, clinically asymptomatic. Patient 4 exemplifies what may be construed as the ideal scenario of newborn screening for Krabbe disease, a condition where timely intervention is the key to achieve the best possible outcome.16 Despite transfer of the blood-spot sample at the beginning of a weekend, all testing was completed by the following Monday morning and the patient was admitted to Duke University Medical Center (Durham, NC) early on Tuesday morning, day 7 of life. His confirmatory workup was completed rapidly and a bone-marrow transplant took place on day 24 of life. The patient was developing normally as of 9 months of age but with some complications, attributed to the transplant itself.
Discussion
We report for the first time that automated integration of covariate-adjusted reference intervals and population results combined with second-tier tests can improve the false-positive rate of newborn screening for lysosomal disorders to a sustainable, near-zero level. An additional novel element of this work is the curation of condition ranges for false-positive cases with heterozygous or pseudo-deficiency genotypes. False-positive cases are actually a precious resource available to pave the way to an objective, evidence-based segregation from affected patients using CLIR tools and second-tier tests instead of the traditional approach, employing one condition-one marker and the use of arbitrary cutoff values,32 which is not adequate for lysosomal disorders. The workflow proposed here resolved 98% of the cases with an initial tool score greater than zero. For the sake of argument, dismissal of marginally abnormal results as clinically irrelevant may raise concerns about harboring false-negative events in the population studied. While only continued clinical surveillance could answer this question with confidence, extensive precautionary measures were in place. Foremost among them was the use of all possible permutations of calculated ratios to define a unique biochemical fingerprint of true and false-positive cases that are recognizable and separable by the software. Subjective escalation to the 10-plex panel and liberal utilization of the second-tier tests produced no additional benefit. On the other hand, perceivable limitations of the proposed approach are the higher reagent cost of the 6-plex profile as primary screening, instead of a repeat analysis; the reluctance to forsake cutoff values and break away from established procedures, as often happens in diverse medical fields,33 and an anecdotal concern that results matched to four covariates may breach anonymization and therefore data could not be shared without infringement of existing policies. Furthermore, daily public-health practice cannot depend on manual data entry. The automated creation of.csv files from diverse commercial laboratory information systems now inclusive of covariates, often extracted from separate databases, is still a challenge that has not been adequately resolved by commonly used platforms and requires further development.
Although the pretransplant evaluation of patient 4 was state-of-the-art and inherently expensive, it was fully justified by the circumstances. On the other hand, similar or even partial workups of false-positive cases should be a source of concern, but the financial impact of current practices is seldom discussed. One author (D.S.P.) recapitulated the charges actually billed to health insurance plans on behalf of patients with whom she personally interacted while working in a state offering newborn screening for lysosomal disorders.12 This review showed that just the initial encounter to work up an abnormal referral could cost an average of $6,823 per case (range $4,242–13,438, N = 10) for the combined expenses of office visits, counseling, procedures, laboratory tests, and comprehensive molecular genetic testing. A linear extrapolation of these charges to a hypothetical nationwide implementation (~4.2 million births) when the false-positive rate is not better than a 0.1-0.5% range may translate into $28–143 million of unnecessary expenses per year. Even a smaller portion of these costs would add up to significant expenditures when applied broadly to all other conditions with obligatory genotyping and a record of suboptimal performance.34
With our proposed workflow, a given program has full control of what conditions are targeted and incidental findings can safely be avoided. In other words, a patient with Fabry disease, Gaucher disease, or Niemann-Pick A and B diseases will not be recognized by deploying tools for Krabbe disease, MPS I, and Pompe disease. Screening for more lysosomal disorders, as already mandated by a few states, merely involves activating the tool for the additional condition(s) of interest. The inclusion of X-linked adrenoleukodystrophy could be accomplished by switching the primary 6-plex screening to the 10-plex mode.19
Work is in progress to create even more complex regression models that simultaneously correct for two continuous covariates (age and birth weight), one categorical covariate (sex), and location. This type of statistical modeling will require big data,35 and a readiness to evaluate the concept that covariate-adjusted reference intervals could be defined by “recycling” vast amounts of normal clinical test results from multiple sources.28 Once a critical mass of data is reached, two-dimensional percentiles of reference and disease intervals in a three-dimensional space will become routinely available as the foundation of the next generation of postanalytical interpretive tools. At the same time, it will also be feasible to explore their clinical applicability to a much broader landscape of laboratory tests and panels. It is worth mentioning that CLIR is a collaborative forum and not a commercial product, access is freely available to legitimate newborn screening programs and other laboratories, with the sole requirement of sustained collaboration and data sharing to the benefit of all users.
At a time when clinical practice is pressed to reinvent itself through utilization management and precision-medicine initiatives, public health, and specifically the performance of newborn screening, should not be exempted from public scrutiny.32,36 Newborn screening is based on laboratory tests performed on a growing proportion of ~130 millions of newborns worldwide every year. Poor performance on a mass scale distresses a multitude of patients37 and exposes their families and providers to high risk of psychosocial harm.38 In the case of lysosomal disorders, genomic medicine has struggled to bring closure to the evaluation of presumptive positive cases with an abundance of inconclusive molecular results.34,39 In the United States, the financial costs of open-ended monitoring of these patients could eventually add up to hundreds of millions of dollars per year. As status quo performance is neither scalable nor sustainable in our health-care system, integrated biochemical phenotyping offers an opportunity to reach the goal of meaningful delivery of precision medicine. After 50 years of preparation,1 the era of precision screening could start now.
References
Howell RR. Fifty years of newborn screening. Mol Genet Metab 2014;113:4–5.
Lyon J. Concerns over newborn screening. JAMA 2017;317:576.
US Department of Health and Human Services. Advisory Committee on Heritable Disorders in Newborns and Children. Adoption by all state newborn screening programs of the recommended uniform screening panel. Response by the HHS Secretary. https://www.hrsa.gov/advisorycommittees/mchbadvisory/heritabledisorders/recommendations/correspondence/uniformpanelsecre052110.pdf. Accessed 24 April 2017.
US Department of Health and Human Services. Advisory Committee on Heritable Disorders in Newborns and Children. Pompe disease. Secretary's final response RE Committee's Recommendation to add Pompe Disease to the RUSP. https://www.hrsa.gov/advisorycommittees/mchbadvisory/heritabledisorders/recommendations/correspondence/secretaryfinalresponse.pdf. Accessed 24 April 2017.
Goldenberg AJ, Comeau AM, Grosse SD et al. Evaluating harms in the assessment of net benefit: A framework for newborn screening condition review. Matern Child Health J 2016;20:693–700.
Sanjiv Harpavat S, Garcia-Prats JA, Benjamin L, Shneider BL. Newborn bilirubin screening for biliary atresia. N Engl J Med 2016;375:605–606.
Moat SJ, Korpimäki T, Furu P et al. Characterization of a blood spot creatine kinase skeletal muscle isoform immunoassay for high-throughput newborn screening of Duchenne Muscular Dystrophy. Clin Chem 2017;63:908–914.
van der Ploeg AT, Reuser AJ. Pompe’s disease. Lancet 2008;372:1342–1353.
Clarke LA MPS I. 31 Oct 2002 (updated 11 February 2016). In: Pagon RA, Adam MP, Ardinger HH et al,, (eds). GeneReviews. Seattle, WA: University of Washington, 1993–2017.
Kemper AR, Brosco J, Comeau AM et al. Newborn screening for X-linked adrenoleukodystrophy: evidence summary and advisory committee recommendation. Genet Med 2017;19:121–126.
Escolar ML, Poe MD, Provenzale JM et al. Transplantation of umbilical-cord blood in babies with infantile Krabbe’s disease. N Engl J Med 2005;352:2069–2081.
Hopkins PV, Campbell C, Klug T, Rogers S, Raburn-Miller J, Kiesling J. Lysosomal storage disorder screening implementation: findings from the first six months of full population pilot testing in Missouri. J Pediatr 2015;166:172–177.
Orsini JJ, Kay DM, Saavedra-Matiz CA et al. Newborn screening for Krabbe disease in New York State: the first eight years’ experience. Genet Med 2016;18:239–248.
Burton BK, Charrow J, Hoganson GE et al. Newborn screening for lysosomal storage disorders in Illinois: the initial 15-month experience. J Pediatr; e-pub ahead of print 17 July 2017.
Navarrete-Martínez JI, Limón-Rojas AE, Gaytán-García MJ et al. Newborn screening for six lysosomal storage disorders in a cohort of Mexican patients: three-year findings from a screening program in a closed Mexican health system. Mol Genet Metab 2017;121:16–21.
Wasserstein MP, Andriola M, Arnold G et al. Clinical outcomes of children with abnormal newborn screening results for Krabbe disease in New York State. Genet Med 2016;18:1235–1243.
Goodman B, Miller A Would you want your baby tested for this disease? 9 March 2018. http://www.webmd.com/parenting/baby/news/20170309/newborn-screening-krabbe-disease?ecd=soc_fb_170310_cons_news_newbornscreeningkrabbedisease. Accessed 24 April 2017.
Position Statement of the Society for Inherited Metabolic Disorders (SIMD). Identifying abnormal newborn screens requiring immediate notification of the health care provider. http://www.simd.org/Issues/SIMD%20NBS%20Critical%20Conditions%20policy%20statement.pdf Accessed 24 April 2017.
Tortorelli S, Turgeon CT, Gavrilov DK et al. Simultaneous testing for six lysosomal storage disorders and X-adrenoleukodystrophy in dried blood spots by tandem mass spectrometry. Clin Chem 2016;62:1248–1254.
Chennamaneni NK, Kumar AB, Barcenas M et al. Improved reagents for newborn screening of mucopolysaccharidosis types I, II, and VI by tandem mass spectrometry. Anal Chem 2014;86:4508–4514.
Turgeon CT, Orsini JJ, Sanders KA et al. Measurement of psychosine in dried blood spots: A possible improvement to newborn screening programs for Krabbe disease. J Inherit Metab Dis 2015;38:923–929.
de Ruijter J, de Ru MH, Tom Wagemans T et al. Heparan sulfate and dermatan sulfate derived disaccharides are sensitive markers for newborn screening for mucopolysaccharidoses types I, II and III. Mol Gen Metab 2012;107:705–710.
Tortorelli S, Eckerman JS, Orsini JJ et al. Moonlighting newbornscreening markers: the incidental discovery of a second tier test for Pompe disease. Genet Med, in press.
McHugh DMS, Cameron CA, Abdenur JE et al. Clinical validation of cutoff target ranges in newborn screening of metabolic disorders by tandem mass spectrometry: a worldwide collaborative project. Genet Med 2011;13:230–254.
Marquardt G, Currier R, McHugh DMS et al. Enhanced interpretation of newborn screening results without analyte cutoff values. Genet Med 2012;14:648–654.
Hall PL, Marquardt G, McHugh DMS et al. Post-analytical tools improve performance of newborn screening by tandem mass spectrometry. Genet Med 2014;16:889–895.
Merritt JL 2nd, Vedal S, Abdenur JE et al. Infants suspected to have very-long chain acyl-CoA dehydrogenase deficiency from newborn screening. Mol Genet Metab 2014;111:484–492.
Mørkrid L, Rowe AD, Elgstoen KBP et al. Continuous age- and gender-adjusted reference intervals of urinary markers for cerebral creatine deficiency syndromes: A novel approach to the definition of reference intervals. Clin Chem 2015;61:760–768.
Horn PS, Feng L, Li Y, Pesce AJ. Effect of outliers and non-healthy individuals on reference interval estimation. Clin Chem 2001;47:2137–2145.
Sarafoglou K, Matern D, Redlinger-Grosse K et al. Siblings with mitochondrial acetoacetyl-CoA thiolase deficiency not identified by newborn screening. Pediatrics. 2011;128:E246–250.
Rinaldo P, Zafari S, Tortorelli S, Matern D. Making the case for objective performance metrics in newborn screening by tandem mass spectrometry. Ment Retard Dev Disabil Res Rev 2006;12:255–261.
Gabler E The price of being wrong. Milwaukee Journal Sentinel. 9 December 2016. http://projects.jsonline.com/news/2016/12/11/the-price-of-being-wrong.html. Accessed 24 April 2017.
Herasevich V, Pickering BW, Dong Y, Peters SG, Gajic O. Informatics infrastructure for syndrome surveillance, decision support, reporting, and modeling of critical illness. Mayo Clin Proc 2010;85:247–254.
Hubbard HB. Expanded newborn screening for genetic and metabolic disorders: Modeling costs and outcomes. Nurs Econ 2007;25:345–352.
Myers SR, Carr BG, Branas CC. Uniting big health data for a national learning health System in the United States. JAMA Pediatr 2016;170:1133–1134.
USA Today Editorial Board. When a child is born, tests should be uniform: our view. http://www.usatoday.com/story/opinion/2016/12/22/newborn-tests-screening-hospitals-editorials-debates/95769812/. Accessed 24 April 2017.
Tarini BA, Christakis DA, Welch HG. State newborn screening in the tandem mass spectrometry era: more tests, more false-positive results. Pediatrics 2006;118:448–456.
Morrison DR, Clayton EW. False positive newborn screening results are not always benign. Public Health Genomics 2011;14:173–177.
Elliotta S, Burokera N, Cournoyerb JJ, Potierb AM, Trometerb JD, Elbinb C et al. Pilot study of newborn screening for six lysosomal storage diseases using tandem mass spectrometry. Mol Gen Metab 2016;118:304–309.
Rinaldo P The Region 4 Stork (R4S) collaborative project: Laboratory quality improvement of newborn screening. Part III—Post-analytical interpretive tools. https://news.mayomedicallaboratories.com/2013/06/18/new-hot-topic-the-region-4-storkr4s-collaborative-project-part-3-post-analytical-interpretive-tools/ Accessed 24 April 2017.
Acknowledgments
This work was supported in part by the T. Denny Sanford professorship, Mayo Clinic. We thank Patricia L. Hall (Emory University, Atlanta, GA), Rosemary Hage (Ohio Department of Health Laboratory, Reynoldsburg, OH), and Joseph J. Orsini (New York State Department of Health, Albany, NY) for giving permission to display their anonymized reference intervals in Supplementary Figure 1.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare no conflict of interest.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Minter Baerg, M.M., Stoway, S.D., Hart, J. et al. Precision newborn screening for lysosomal disorders. Genet Med 20, 847–854 (2018). https://doi.org/10.1038/gim.2017.194
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/gim.2017.194
Keywords
This article is cited by
-
Application of tandem mass spectrometry in the screening and diagnosis of mucopolysaccharidoses
Orphanet Journal of Rare Diseases (2024)
-
Global carrier frequency and predicted genetic prevalence of patients with pathogenic sequence variants in autosomal recessive genetic neuromuscular diseases
Scientific Reports (2024)
-
Epidemiology of mucopolysaccharidoses (MPS) in United States: challenges and opportunities
Orphanet Journal of Rare Diseases (2021)
-
Consensus guidelines for newborn screening, diagnosis and treatment of infantile Krabbe disease
Orphanet Journal of Rare Diseases (2018)
-
Status of newborn screening and follow up investigations for Mucopolysaccharidoses I and II in Taiwan
Orphanet Journal of Rare Diseases (2018)