Abstract
Over the past few years, the application of whole-genome scanning array technologies has catalyzed the appreciation of a new form of submicroscopic genomic imbalances, referred to as copy number variants. Copy number variants contribute substantially to genetic diversity and result from gains and losses of genomic regions that are 1000 base pairs in size or larger, sometimes encompassing millions of bases of contiguous DNA sequences. As genome-wide scanning techniques become more widely used in diagnostic laboratories, a major challenge is how to accurately interpret which submicroscopic genomic imbalances are pathogenic in nature and which are benign. Herein, we review the literature from the past 3 years on this new source of genomic variability and comment on factors that should be considered when trying to differentiate between a pathogenic and a benign copy number variant.
Similar content being viewed by others
INTRODUCTION
Genomic variability in humans exists on widely different scales. Microscopic variants, 5 Mb or greater in size, have been identified since 1959 by using standard cytogenetic analysis (e.g., G-banded karyotyping).1 At this level of analysis, it is possible to survey the entire human genome for gains, losses, or rearrangements of genetic material in a single test, but in practice, imbalances smaller than 10 to 20 Mb are often not readily detected. Over the years, classical cytogenetic studies have uncovered several heteromorphisms and euchromatic variants that do not seem to have clinical significance.2,3
With the advent of molecular cytogenetic techniques, such as fluorescence in situ hybridization (FISH), it became possible to more precisely define the extent and the actual DNA sequences involved in these chromosomal variants at a much higher resolution. However, FISH uses probes specifically targeting a given chromosomal locus and assessment of genomic imbalances at multiple chromosomal loci using this technique rapidly becomes labor intensive.
On the other side of the spectrum, genotyping technologies have allowed us to detect smaller and more abundant forms of genomic variability (e.g., single nucleotide polymorphisms [SNPs]). In fact, SNPs were long considered the largest source of genomic variation in humans, with estimates of at least 10 million SNPs within the human population, averaging 1 SNP for every 300 nucleotides in an individual.4
With the development of array-based comparative genomic hybridization (CGH) technologies, a large number of submicroscopic genomic imbalances have now also being identified.5,6 These genomic imbalances are referred to as copy number variants (CNVs) and are defined as deletions and duplications of DNA segments larger than 1000 bases (1 kb) and up to several Mb in size that are present in variable copy number compared with a reference genome.7–9
Over the past few years, the term CNV has been broadly used,10 going beyond the clinical definition of a variant, which usually implies a benign genetic change that does not cause a clinically recognizable phenotype.2,11 However, with increased genotype-phenotype correlations, CNVs that were once thought to be benign or of unknown clinical significance are now known to be associated with and definitive of specific genomic syndromes.12–14 Such associations are appreciated when a particular CNV is observed recurrently among unrelated individuals with similar clinical presentations and/or when the genomic imbalance is found to cosegregate with the clinical presentation in families containing multiple affected individuals. Our limited understanding of the phenotypic impact of the hundreds of CNVs that have already been discovered warrants the use of qualifiers to minimize confusion (especially in a diagnostic setting). Hence, in this perspective, we refer to CNVs as being pathogenic, benign, or of unknown clinical significance—definitions that are based on our current understanding of the structure and function of the human genome.
AN ABUNDANCE OF CNVs IN THE HUMAN GENOME
In 2004, two independent studies screened the human genome of healthy individuals by using array CGH and reported the widespread presence of CNVs.5,6 Iafrate and coworkers5 used a bacterial artificial chromosome (BAC)-based array, with clones chosen at approximately 1-Mb intervals throughout the human genome to identify more than 200 variable loci among 39 unrelated healthy individuals. Sebat and colleagues6 used a microarray platform containing oligonucleotides spaced at 35-kb intervals and detected 76 CNVs among 20 individuals. Although both studies used slightly different approaches to study the genome of unrelated individuals, they reached the same conclusion: phenotypically normal individuals have an unexpectedly high number of genomic imbalances throughout their genomes. However, because of the small number of individuals examined and the limited resolution of both platforms, neither study provided a comprehensive evaluation of CNVs in the human genome. Indeed, the number of CNVs identified by these two studies seemed likely to be an underestimation of the true number of CNVs in humans.15
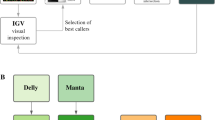
A few years later, Redon and coworkers16 published a more comprehensive CNV map for the human genome. In this study, the DNAs of 270 healthy individuals from four populations (the HapMap collection) were analyzed using two different array platforms: a high-density, genome-wide SNP array (the Affymetrix 500k EA genotyping chips)17 and a whole genome tilepath (WGTP) BAC array containing clones that together represented 94% of the euchromatic portion of the human genome.18 Both methods were capable of detecting CNVs and, in many ways, were complementary to each other. The SNP arrays tended to detect smaller CNVs in regions that had good probe coverage and provided better definition of the structure of CNVs at these regions. The WGTP platform seemed to be more useful in detecting larger and more complex CNVs. Incidentally, these were often regions of the human genome that were overlapping segmental duplications (also known as low copy repeats), which have been found to be regions sparsely covered by SNP genotyping probes.19 Overlapping and juxtaposed CNVs identified by both platforms were merged together into 1447 discrete CNV regions (CNVRs) (discrete CNVRs can be seen for the 1p36.33 chromosome region in Figure 1). The CNVRs identified in this study represented 12% (360 Mb) of the human genome. A whole-genome view of the distribution of CNVRs revealed that they are ubiquitously distributed throughout the genome, with approximately 24% of the CNVRs located near previously known segmental duplications. CNVs that are located in close proximity to segmental duplications are thought to be generated and maintained via nonallelic homologous recombination (NAHR) mechanisms that result from recombination events between flanking segmental duplications.20
A schematic representation of a copy number variant region (CNVR) and CNVs for a part of chromosome region 1p36.33. CNVs called using two different array CGH platforms (i.e., a high-density genome-wide SNP array Affymetrix 500k EA chip and a whole-genome tilepath [WGTP] BAC array) in four different HapMap individuals are represented in colored boxes (Individuals A, B, C, and D). The relative size and position of BAC clones on this WGTP array and the relative position of SNP-detecting oligonucleotide on the Affymetrix 500k EA chip are shown above the CNV regions. Figure adapted from the Database of Genomic Variants (http://projects.tcag.ca/variation).
Based on current information, CNVs tend to be preferentially located outside of genes and ultra-conserved elements in the human genome, with as much as 40% of CNVs lying within gene deserts.16,21 Nevertheless, a substantial number of genes still lie within these CNV regions. Redon et al.16 found that among the 1447 identified HapMap CNVRs, 2908 RefSeq genes (i.e., protein-coding genes taken from the NCBI mRNA reference sequence collection) and 285 OMIM genes (i.e., genes associated with human disorders and that are listed in the Online Mendelian Inheritance in Man database, www.ncbi.nlm.nih.gov/omim) were present, suggesting a possible relationship between certain CNVs and complex diseases/Mendelian disorders. CNV genes do not usually encode for proteins that are critical for development or viability, but instead encode gene products that influence the way that we interact with the environment. Referred to as “environmental sensor genes” by some, they often play a role in cell adhesion, sensory perception, chemical stimuli, and neurophysiological processes. Non-CNV genes are usually genes that are likely to be dosage sensitive and are more critical for cellular maintenance and proper development.16,22 These include genes related to cell signaling, proliferation, and kinase and phosphorylation processes. Interestingly, there have been data suggesting that some CNV regions may overlap with genomic regions corresponding to noncoding RNAs, including microRNAs (miRNAs).16,22 miRNAs regulate gene expression post-transcriptionally and play a critical role in developmental and physiological processes. They have also been implicated in the pathogenesis of several human diseases including cancer.23 Although the effect of DNA copy number variability for miRNAs is not well understood, evidence for disregulated miRNAs expression via copy number changes on chromosome region 13q14 have already been noted for certain hematological malignances.24–26
There are different ways in which a CNV can affect gene expression levels: (1) a CNV can directly affect gene expression levels by altering the actual dosage of a particular gene, or (2) CNVs may indirectly affect gene transcriptional regulatory elements, leading to altered gene expression levels via a positional effect.27–29 For example, a deletion of a repressor element may lead to increased transcriptional levels of the associated gene, whereas duplications of DNA sequences, 3′ to a promoter, may lead to decreased gene expression levels because of suboptimal placement of the promoter with respect to the gene. In an attempt to estimate the relative contribution of CNVs to gene expression variability, Stranger and colleagues28 correlated HapMap CNV data with gene expression data and found that CNVs were correlated with 17.7% of the observed gene expression variability. Most correlations were positive in nature (i.e., increased copy number of a genomic region led to increased expression levels of an overlapping or nearby gene). However, as much as 15% of the associations had an inverse relationship where increased copy number of a genomic region (e.g., duplication of a putative repressor element) led to decreased expression levels of an overlapping or nearby gene. An example for this is a small duplication (<150 kb) downstream of the proteolipid protein gene (PLP1) that silences PLP1 gene expression and results in a spastic paraplegia type 2 phenotype that is also observed when no PLP1 protein is produced.30 Amazingly, some CNVs can exert transcriptional regulatory effects on a gene over extremely large genomic distances, as much as 6 Mb.28
Taken together, these and many other studies31–34 have revealed that the genomes of healthy individuals contain a substantial number of CNVs, and these CNVs likely contribute significantly to human phenotypic diversity. Moreover, over the past 2 years, a dozen or more CNVs have been shown to be associated with differential susceptibility to common human diseases (recently reviewed in ref.35). For example, Fanciulli and colleagues36 recently showed that reduced copy number of the FCGR3B gene is associated with increased susceptibility to systemic autoimmunity. Because CNVs represent a substantial component of natural genetic variation, future disease linkage and association studies should incorporate an evaluation of CNVs. Although some disease-related CNVs may be detected via SNP-based linkage or association analysis, many others are either not in linkage disequilibrium to nearby SNPs or in genomic regions that have insufficient SNP detection coverage on a given genotyping platform.32,37 To include CNV data in these studies, genotyping platforms may be modified to incorporate strategically placed probes for assessing copy number information at known CNV regions. Alternatively, array CGH platforms may be applied in a complementary experimental fashion to all samples being genotyped in a study.
CNVs AND THEIR IMPACT ON CLINICAL DIAGNOSIS
With the implementation of array CGH technologies as a diagnostic tool in clinical cytogenetic laboratories and with the appreciation for the ubiquitous nature of CNVs in the human genome, it is becoming more difficult to accurately differentiate benign CNVs from pathogenic CNVs. In general, smaller and targeted arrays (those that typically cover the subtelomeric and clinically defined regions)38 tend to have fewer CNVs that can be categorized as benign or of unknown clinical significance.39,40 On the other hand, the application of genome-wide array CGH platforms (with effective resolutions that are often 50–100 times higher than that of routine banded chromosomal analysis)41,42 reveals many more CNVs that are difficult to interpret.
At a research level, studies using genome-wide array CGH have directly led to the association of specific submicroscopic imbalances with certain clinically recognizable congenital disorders.12–14,39,43–49 However, in a clinical setting, when a genome-wide array CGH is applied, CNVs will initially fall into two categories: those clearly associated with a genomic disorder and those of uncertain clinical significance. At this point, the clinical cytogeneticist needs to assess the potential pathogenicity of each CNV with unknown clinical significance. The following are some criteria that could be considered when attempting to assess the potential pathogenicity of a given CNV.
Parental/familial studies
In conventional cytogenetics diagnosis, one of the first steps in assessing whether a novel chromosomal alteration is pathogenic is to try to determine whether the chromosomal alteration is inherited. This is accomplished from parental chromosome analyses. If a chromosomal alteration is observed in the affected individual and in a normal, healthy parent, it suggests that the rearrangement is noncontributing to the clinical phenotype. A similar approach can be used for genome-wide and targeted array CGH studies. If a CNV that is observed in the array CGH profile of the affected individual is also observed in an unaffected parent, it is less likely to be pathogenic. A CNV that seems to be de novo in nature has an increased risk of being disease-causing.50,51 If the CNV seems to be de novo, false paternity should be considered during the interpretation of the results. If the CNV seems to be inherited from an apparently healthy parent, an extensive pedigree evaluation (including siblings and other related individuals) may still be warranted. In some cases, a clinical reexamination of “unaffected” carriers of a CNV may actually reveal an underappreciation for subtle clinical presentations that may alter the pathogenic risk assessment for the CNV in question.3,52 Issues such as incomplete penetrance; variable expression of an inherited phenotype; mosaicism (including gonadal mosaicism) in a parent; and epistatic, epigenetic, or environmental factors that can coincide with a given CNV in the patient should also be noted during CNV pathogenicity risk assessment.3,53
There are some situations in which an apparently inherited CNV may still cause pathogenicity in the proband. For example, some deletion CNVs may unmask a recessive mutation on the other allele in the patient but not in a healthy parent. Similarly, deletion CNVs involving X chromosomal genes may not lead to pathogenicity in the mother or other female relatives (because of the presence of an intact copy of the gene on the other X chromosome) but cause a genomic disorder in the son.8
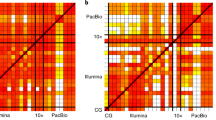
Determining the inheritance of CNVs by array CGH may not always be straightforward. Inheritance patterns of simple CNVs (e.g., biallelic) are easier to interpret compared with multiallelic or complex CNVs. Part of the complication relies on the fact that CNVs identified by array CGH are calculated additively (i.e., based on the diploid genome) and that this technology does not provide allele-specific copy number information. Therefore, one should be cautious when attempting to determine the true inheritance patterns of CNVs solely from array CGH results (Fig. 2).7
A partial array comparative genomic hybridization (CGH) profile (based on the log2 intensity ratios of fluorescence intensities from a dye-swap experiment) and the allele-specific copy number information for the copy number variant (CNV) region is provided for the child as well as both parents. The red line represents results from one dye-swap experiment and the blue line represents results from the other dye-swap experiment. A black horizontal arrow within each partial array CGH profile indicates the clone containing DNA sequences that are copy number variable. The affected child is inheriting the null allele from the mother and the null allele from the father, but the array CGH profile data may erroneously suggest that this is a de novo CNV.
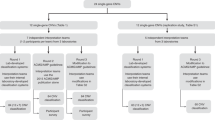
It is also important to note that most current array CGH platforms have technical limitations, including low resolution for defining CNV boundaries, inability to provide information on genomic distributions of CNVs, and inability to provide absolute copy number information. For these reasons (and to minimize false-positive results), any variation found on an array CGH-based clinical test should be ideally confirmed with alternate molecular techniques such as FISH analysis (using a clone within the CNV region), multiplex ligation-dependent probe amplification (MLPA) (with customized probes), or quantitative PCR (qPCR). Among these choices for confirmation tests, FISH is the only one that provides information on the genomic distribution of the copy number variable DNA sequences. Such information could lead to the detection of a cryptic and balanced chromosomal translocation in one of the parents, which in turn carries an increased recurrence risk (Fig. 3). This is clearly important for accurate genetic counseling and indicative for future prenatal testing.
An illustration of a de novo duplication copy number variant (CNV) (based on log2 intensity ratios) that is detected in an affected child using array comparative genomic hybridization (CGH) with a dye-swap strategy. The red line represents results from one dye-swap experiment and the blue line represents results from the other dye-swap experiment. Array CGH studies for (A) the father and (B) the mother show no genomic imbalance at this chromosomal region. Confirmation fluorescence in situ hybridization (FISH) studies with chromosome-specific subtelomeric probes for chromosome (Chr.) 1 (green dots represent a 1p subtelomeric probe and red dots represent a 1q subtelomeric probe) in the affected child demonstrate that the duplicated DNA sequence is actually at another chromosome region (i.e., on the short arm of a chromosome 13), which could be the unbalanced product from a parent with a balanced chromosomal rearrangement. Subsequent FISH studies in (A) the father and (B) the mother reveals a balanced rearrangement in the mother, leading to an increased recurrence risk in her future pregnancies.
Comparison with data from other affected individuals
De novo CNVs should be cross referenced to known, pathogenic genomic imbalances. If the observed de novo CNV matches (or overlaps) a known genomic disorder (i.e., a CNV that has been demonstrated to recurrently be associated with a specific clinical phenotype) it is usually assumed to be pathogenic and contributory to the clinical phenotype. Databases have been developed to collect array CGH and clinical phenotype data on patients referred for genetic testing. The Database of Chromosomal Imbalance and Phenotype in Humans using Ensembl Resources (DECIPHER, http://www.sanger.ac.uk/postGenomics/decipher) is one such resource. Other similar initiatives include the Mendelian Cytogenetics Network Online Database, the Chromosome Abnormality Database (CAD, www.ukcad.org.uk/cocoon/ukcad/), and the European Cytogeneticists Association Register of Unbalanced Chromosome Aberration (ECARUCA, www.ECARUCA.net). Since chromosome imbalances occur throughout the genome and are rare in nature, the actual success of such database efforts rely on a collective global responsibility for sharing array CGH and clinical phenotype data.
Reference CNV databases
De novo CNVs that have not been recurrently reported in other patients should then be cross-referenced to CNVs that have been identified among healthy individuals (e.g., Database of Genomic Variants, http://projects.tcag.ca/variation). If an apparently de novo CNV is found in this database of benign CNVs, it reduces the likelihood that the CNV is causative of the clinical phenotype, with the caveat that most CNVs currently in the database have ill-defined boundaries and only a fraction have been independently verified by multiple studies or alternate CNV detection technologies. Furthermore, it has been shown that the frequency of certain CNVs can vary significantly among ethnic populations.16,54 Therefore, the usefulness of the information of such databases could be decreased for a patient whose ethnic population is underrepresented in the databases. Ultimately, the deposition of CNV data from high-resolution assays for a wide variety of human populations into these publicly accessible databases should significantly improve clinical interpretations of CNVs observed in genome-wide diagnostic assays.
All array CGH methods that have been used to identify CNVs rely on a comparison to a reference genome. Unfortunately, no single individual or DNA source has yet been adopted as a standardized control, which can complicate the designation of copy number changes and subsequent standardization of CNV entries on databases. For example, a loss detected by an array CGH assay may represent a deletion in the test sample or a duplication in the reference sample.7 Therefore, not only is the mapping, characterization, and accurate cataloging of all CNVs in the human genome important, but a detailed genomic characterization of one or a few reference genomes may also be warranted.
Genomic architecture of CNVs
To determine the clinical consequences of a CNV that has not been detected in other patients and is not observed among healthy individuals, other factors such as the type (deletion or duplication), the size, and even the number of copies of the CNV may be considered. For example, it is generally accepted that the human genome is less tolerant of haplo-insufficiency compared with having extra copies of a particular DNA sequence.55 Thus, all else being equal, a given genomic region that is deleted (i.e., a deletion CNV) is more likely to result in pathogenicity than a duplication of the same genomic region (i.e., a duplication CNV).
With respect to size of the CNV, it stands to reason that pathogenic imbalances tend to be larger than benign CNVs.52 De Vries and coworkers45 reported that the median size of benign CNVs was 0.43 Mb, whereas clinically relevant CNVs had a median size of 2.76 Mb. More important than the actual size of the CNV is the number and type of genes that lie within the CNV region. For example, a 100-kb deletion that encompasses two developmentally important genes is more likely to contribute to the etiology of a dysmorphic and developmentally challenged patient than a 800-kb deletion in a gene desert portion of the genome. Indeed, large-scale deletions in a gene desert or in gene-poor regions (composed of noncoding DNA) have been shown to be well tolerated in a variety of organisms.27,32,33,56 These criteria should also be weighed with respect to our lack of understanding of how copy number changes of regulatory elements affect transcription levels of nearby and distantly positioned genes.28 As higher-resolution CNV data emerge and are integrated with functional information (i.e., transcriptional and translational levels), we may be able to more accurately predict the functional effects of these CNVs.
Obtaining absolute copy number information may also be clinically important, especially when a dosage-sensitive gene is implicated in the disorder. For example, a CNV duplication that results in three copies of a given DNA sequence per diploid cell may be phenotypically benign until a particular threshold is crossed (e.g., five copies of the same sequence per diploid cell). In such scenarios, it may be hypothesized that excessive protein levels result in a toxic gain-of-function, leading to a clinical phenotype. Similarly, genes that are in multiple copies in healthy individuals may be haplo-sufficient (not critically detrimental in one copy per cell) but pathogenic when homozygously deleted. Because one of the limitations of most array CGH-based assays is the inability to provide absolute copy number information, alternative quantitative assays that determine such copy number information, in an independent manner, may help to identify CNVs with clinically significant copy number threshold levels.57
In conclusion, despite all the different factors that may be considered when determining the pathogenic effect of a CNV, the clinical significance of many CNVs may still remain unknown. The uncertain clinical implications of these CNVs should be well explained in clinical reports and well conveyed during genetic counseling sessions. We have just begun to reveal the complexity of variation in the human genome, and, in many ways, technology has advanced more rapidly than our ability to understand the biological and medical implications of the generated information. Only by combining efforts will we be able to unravel the contribution that these CNVs have in clinical phenotypes, genetic disorders, and normal human phenotypic diversity.
References
Lejeune J, Gautier M, Turpin R . Study of somatic chromosomes from 9 mongoloid children. C R Hebd Seances Acad Sci 1959; 248: 1721–1722.
Wyandt HE, Tonk VS, editors. Atlas of chromosome heteromorphisms. Boston: Kluwer Academic Publishers, 2004: 279.
Barber JCK . Directly transmitted unbalanced chromosome abnormalities and euchromatic variants. J Med Genet 2005; 42: 609–629.
Kruglyak L, Nickerson DA . Variation is the spice of life. Nat Genet 2001; 27: 234–236.
Iafrate AJ, Feuk L, Rivera MN, Listewnik ML, et al. Detection of large-scale variation in the human genome. Nat Genet 2004; 36: 949–951.
Sebat J, Lakshmi B, Troge J, Alexander J, et al. Large-scale copy number polymorphism in the human genome. Science 2004; 305: 525–528.
Freeman JL, Perry GH, Feuk L, Redon R, et al. Copy number variation: new insights in genome diversity. Genome Res 2006; 16: 949–961.
Feuk L, Carson AR, Scherer SW . Structural variation in the human genome. Nat Rev Genet 2006; 7: 85–97.
Kehrer-Sawatzki H . What a difference copy number variation makes. Bioessays 2007; 29: 311–313.
Lee JA, Lupski JR . Genomic rearrangements and gene copy-number alterations as a cause of nervous system disorders. Neuron 2006; 52: 103–121.
Benn PA, Hsu LY, Prenatal diagnosis of chromosomal abnormalities through amniocentesis. In: Milunsky A, editors, Genetic disorders and the fetus, 5th edition. Baltimore: The Johns Hopkins University Press, 2004; 271–274
Koolen DA, Visser LE, Pfundt R, de Leeuw N, Knight SJ, Regan R, et al. A new chromosome 17q21.31 microdeletion syndrome associated with a common inversion polymorphism. Nat Genet 2006; 38: 999–1001.
Shaw-Smith C, Pittman AM, Willatt L, Martin H, Rickman L, Gribble S, et al. Microdeletion encompassing MAPT at chromosome 17q21.3 is associated with developmental delay and learning disability. Nat Genet 2006; 38: 1032–1037.
Redon R, Baujat G, Sanlaville D, Le Merrer M, et al. Interstitial 9q22.3 microdeletion: clinical and molecular characterisation of a newly recognised overgrowth syndrome. Eur J Hum Genet 2006; 14: 759–767.
Carter NP . As normal as normal can be?. Nat Genet 2004; 36: 931–932.
Redon R, Ishikawa S, Fitch KR, Feuk L, et al. Global variation in copy number in the human genome. Nature 2006; 444: 444–454.
Komura D, Shen F, Ishikawa S, Fitch KR, Chen W, Zhang J, et al. Genome-wide detection of human copy number variations using high density DNA oligonucleotide arrays. Genome Res 2006; 16: 1575–1584.
Fiegler H, Redon R, Andrews D, Scott C, et al. Accurate and reliable high-throughput detection of copy number variation in the human genome. Genome Res 2006; 16: 1566–1574.
Wirtenberger M, Hemminki K, Burwinkel B . Identification of frequent chromosome copy-number polymorphisms by use of high-resolution single-nucleotide-polymorphism arrays. Am J Hum Genet 2006; 78: 520–522.
Lupski JA, Stankiewicz P . Genomic disorders: molecular mechanisms for rearrangements and conveyed phenotypes. PLOS Genet 2005; 1: e49.
Derti A, Roth FP, Church GM, Wu CT . Mammalian ultraconserved elements are strongly depleted among segmental duplications and copy number variants. Nat Genet 2006; 38: 1216–1220.
Wong KK, deLeeuw RJ, Dosanjh NS, Kimm LR, et al. A comprehensive analysis of common copy-number variations in the human genome. Am J Hum Genet 2007; 80: 91–104.
Ambros V . The functions of animal microRNAs. Nature 2004; 431: 350–355.
Tagawa H, Seto M . A microRNA cluster as a target of genomic amplification in malignant lymphoma. Leukemia 2005; 19: 2013–2016.
Calin GA, Dumitru CD, Shimizu M, Bichi R, et al. Frequent deletions and down-regulation of micro- RNA genes miR15 and miR16 at 13q14 in chronic lymphocytic leukemia. Proc Natl Acad Sci USA 2002; 99: 15524–15529.
Calin GA, Liu CG, Sevignani C, Ferracin M, et al. MicroRNA profiling reveals distinct signatures in B cell chronic lymphocytic leukemias. Proc Natl Acad Sci USA 2004; 101: 11755–11760.
McCarroll SA, Hadnott TN, Perry GH, Sabeti PC, et al. Common deletion polymorphisms in the human genome. Nat Genet 2006; 38: 86–92.
Stranger BE, Forrest MS, Dunning M, Ingle CE, et al. Relative impact of nucleotide and copy number variation on gene expression phenotypes. Science 2007; 315: 848–853.
Kleinjan DA, van Heyningen V . Long-range control of gene expression: emerging mechanisms and disruption in disease. Am J Hum Genet 2005; 76: 8–32.
Lee JA, Madrid RE, Sperle K, Ritterson CM, et al. Spastic paraplegia type 2 associated with axonal neuropathy and apparent PLP1 position effect. Ann Neurol 2006; 59: 398–403.
Tuzun E, Sharp AJ, Bailey JA, Kaul R, et al. Fine-scale structural variation of the human genome. Nat Genet 2005; 37: 727–732.
Conrad DF, Andrews TD, Carter NP, Hurles ME, et al. A high-resolution survey of deletion polymorphism in the human genome. Nat Genet 2006; 38: 75–81.
Hinds DA, Kloek AP, Jen M, Chen X, Frazer KA . Common deletions and SNPs are in linkage disequilibrium in the human genome. Nat Genet 2006; 38: 82–85.
Khaja R, Zhang J, MacDonald JR, He Y, et al. Genome assembly comparison identifies structural variants in the human genome. Nat Genet 2006; 38: 1413–1418.
Human Genome Structural Variation Working Group, Eichler EE, Nickerson DA, Altshuler D, Bowcock AM, et al. Completing the map of human genetic variation. Nature 2007; 447: 161–165.
Fanciulli M, Norsworthy PJ, Petretto E, Dong R, et al. FCGR3B copy number variation is associated with susceptibility to systemic, but not organ-specific, autoimmunity. Nat Genet 2007; 39: 721–723.
Locke DP, Sharp AJ, McCarroll SA, McGrath SD, et al. Linkage disequilibrium and heritability of copy-number polymorphisms within duplicated regions of the human genome. Am J Hum Genet 2006; 79: 275–290.
Stankiewicz P, Beaudet AL . Use of array CGH in the evaluation of dysmorphology, malformations, developmental delay, and idiopathic mental retardation. Curr Opin Genet Dev 2007; 17: 182–192.
Lu X, Shaw CA, Patel A, Li J, et al. Clinical implementation of chromosomal microarray analysis: summary of 2513 postnatal cases. PLoS ONE 2007; 2: e327.
Subramonia-Iyer S, Sanderson S, Sagoo G, Higgins J, et al. Array-based comparative genomic hybridization for investigating chromosomal abnormalities in patients with learning disability: systematic review meta-analysis of diagnostic and false-positive yields. Genet Med 2007; 9: 74–79.
Solinas-Toldo S, Lampel S, Stilgenbauer S, Nickolenko J, et al. Matrix-based comparative genomic hybridization: biochips to screen for genomic imbalances. Genes Chromosomes Cancer 1997; 20: 399–407.
Pinkel D, Segraves R, Sudar D, Clark S, et al. High resolution analysis of DNA copy number variation using comparative genomic hybridization to microarrays. Nat Genet 1998; 20: 207–211.
Vissers LE, de Vries BB, Osoegawa K, Janssen IM, et al. Array-based comparative genomic hybridization for the genomewide detection of submicroscopic chromosomal abnormalities. Am J Hum Genet 2003; 73: 1261–1270.
Shaw-Smith C, Redon R, Rickman L, Rio M, et al. Microarray based comparative genomic hybridisation (array-CGH) detects submicroscopic chromosomal deletions and duplications in patients with learning disability/mental retardation and dysmorphic features. J Med Genet 2004; 41: 241–248.
de Vries BB, Pfundt R, Leisink M, Koolen DA, et al. Diagnostic genome profiling in mental retardation. Am J Hum Genet 2005; 77: 606–616.
Friedman JM, Baross A, Delaney AD, Ally A, et al. Oligonucleotide microarray analysis of genomic imbalance in children with mental retardation. Am J Hum Genet 2006; 79: 500–513.
Sharp AJ, Hansen S, Selzer RR, Cheng Z, Regan R, Hurst JA, et al. Discovery of previously unidentified genomic disorders from the duplication architecture of the human genome. Nat Genet 2006; 38: 1038–1042.
de Ravel TJ, Balikova I, Thienpont B, Hannes F, et al. Molecular karyotyping of patients with MCA/MR: the blurred boundary between normal and pathogenic variation. Cytogenet Genome Res 2006; 115: 225–230.
Shaffer LG, Kashork CD, Saleki R, Rorem E, et al. Targeted genomic microarray analysis for identification of chromosome abnormalities in 1500 consecutive clinical cases. J Pediatr 2006; 149: 98–102.
Sebat J, Lakshmi B, Malhotra D, Troge J, et al. Strong association of de novo copy number mutations with autism. Science 2007; 316: 445–449.
Autism Genome Project Consortium Szatmari P, Paterson AD, Zwaigenbaum L, Roberts W, et al. Mapping autism risk loci using genetic linkage and chromosomal rearrangements. Nat Genet 2007; 39: 319–328.
Bisgaard AM, Kirchhoff M, Nielsen JE, Brandt C, et al. Transmitted cytogenetic abnormalities in patients with mental retardation: pathogenic or normal variants?. Eur J Med Genet 2007; 50: 243–255.
Feuk L, Marshall CR, Wintle RF, Scherer SW . Structural variants: changing the landscape of chromosomes and design of disease studies. Hum Mol Genet 2006; 15: R57–66.
Kidd JM, Newman TL, Tuzun E, Kaul R, et al. Population stratification of a common APOBEC gene deletion polymorphism. PLoS Genet 2007; 3: e63.
Shaffer LG, Lupski JR . Molecular mechanisms for constitutional chromosomal rearrangements in humans. Annu Rev Genet 2000; 34: 297–329.
Nobrega MA, Zhu Y, Plajzer-Frick I, Afzal V, et al. Megabase deletions of gene deserts result in viable mice. Nature 2004; 431: 988–993.
Lee C, Iafrate AJ, Brothman AR . Copy number variations and clinical cytogenetic diagnosis of constitutional disorders. Nat Genet 2007; 39: 548–554.
Acknowledgements
This work has received financial support from the Leukemia and Lymphoma Society, the Department of Pathology at Brigham and Women's Hospital, and the Centre for Biomedical Research on Rare Diseases (CIBERER). We thank George Perry and Joelle Tchinda for advice and discussion.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Rodriguez-Revenga, L., Mila, M., Rosenberg, C. et al. Structural variation in the human genome: the impact of copy number variants on clinical diagnosis. Genet Med 9, 600–606 (2007). https://doi.org/10.1097/GIM.0b013e318149e1e3
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1097/GIM.0b013e318149e1e3
Keywords
This article is cited by
-
New insights in the interpretation of array-CGH: autism spectrum disorder and positive family history for intellectual disability predict the detection of pathogenic variants
Italian Journal of Pediatrics (2016)
-
Application of array comparative genomic hybridization in 256 patients with developmental delay or intellectual disability
Journal of Applied Genetics (2014)
-
Array comparative genomic hybridization in prenatal diagnosis of first trimester pregnancies at high risk for chromosomal anomalies
Molecular Cytogenetics (2012)
-
Genomic imbalances detected by array-CGH in patients with syndromal ocular developmental anomalies
European Journal of Human Genetics (2012)
-
Genetic variation in phosphodiesterase (PDE) 7B in chronic lymphocytic leukemia: overview of genetic variants of cyclic nucleotide PDEs in human disease
Journal of Human Genetics (2011)