Abstract
Purpose: Clinical geneticists are often asked to evaluate patients with autism spectrum disorders (ASDs) in reference to questions about cause and recurrence risk. Recent advances in diagnostic testing technology have greatly increased the options available to them. It is not currently clear what the overall diagnostic yield of a battery of tests, either collectively or individually, might be. The purpose of this study was to evaluate the diagnostic yield of a stepwise approach we have implemented in our clinics.
Methods: We used a three-tiered neurogenetic evaluation scheme designed to determine the cause of ASDs in patients referred for clinical genetic consultation. We reviewed the results of our diagnostic evaluations on all patients referred with a confirmed diagnosis of autism over a 3-year period.
Results: By using this approach, we found an overall diagnostic yield for ASDs of more than 40%. This represents a significant increase in the diagnostic yield reported just a few years ago.
Conclusions: Given the implications of these diagnoses on recurrence risk and associated medical conditions, a targeted neurogenetic evaluation of all persons with ASDs seems warranted. We discuss the issues in the future implementation of a fourth tier to the evaluation with the potential for an even higher diagnostic yield.
Similar content being viewed by others
Main
Autism spectrum disorders (ASDs) are a collective of neurobehavioral conditions that share in common primary abnormalities of socialization and communication. Diagnostic criteria for autism include impairment of reciprocal social interactions, impairment of verbal and nonverbal communications, restricted educational activities, abnormal interests, and stereotypic behavior. The genetic basis of autism is undeniable. Studies suggest that ASDs best fit into the model of “multifactorial inheritance.” These disorders exhibit complex inheritance with marked etiologic heterogeneity. They occur four times as common in males. There is 70% concordance in monozygotic twins, 90% if a broader phenotype is used. The overall heritability of autism has been stated as approximately 90%.1–5
Clinical geneticists are often asked to evaluate individuals with autism for a possible etiologic diagnosis. Consistently reported diagnoses from such evaluations include chromosome abnormalities, genetic syndromes, metabolic disorders, and teratogenic causes.6–14 Multiple reports over the past several years have evaluated the efficacy (diagnostic yield) of a search for a cause.15–21 Diagnostic yields reported in these studies have ranged from approximately 5% to 25%. The variance in the reported yields in these studies can be attributed to significant differences in the studies themselves. This includes phenotypic definition, test selection, evaluation protocols, and timing of the studies. In fact, one of these studies recently reported an 8% diagnostic yield even for patients with pervasive developmental disorders—children with autistic-like features who do not fully meet diagnostic criteria for autism.15 In review of this collective of reports, several consistent themes emerge. First, a significant number of diagnoses can be made by history and dysmorphology examination alone. Second, carefully considered medical and genetic testing significantly increases diagnostic yield. Third, diagnostic yield increases with the aggressiveness of the evaluation.
There have been several concerted attempts at outlining a standard approach to the genetic evaluation of autism.13,22–24 Review of these documents suggests that there is general agreement that high-resolution (prometaphase) chromosome studies and Fragile X studies should be part of the evaluation of all individuals with ASD. There is, however, no general agreement as to the role of neuroimaging studies in the evaluation of persons with ASDs. Also, it is notable that these cited organizational policy statement documents were written before the advent of many of the newer testing modalities. Although we are aware that updated policy statements are currently being developed, decisions as to an extended evaluation have not been published.
Much has been written about the dramatic increase in the reported occurrence of autism. There is no clear consensus on how much of this increase is the result of changes in diagnostic and reporting practices versus a true biologic increase in numbers. Regardless of the correct answer to this question, the practical impact has been a striking increase in the referrals for genetic evaluation for patients with ASD. It is increasingly common for clinical geneticists to be asked to evaluate patients with ASD in reference to questions about cause and recurrence risk.
For the past 4 years, we have been using a “tiered” neurogenetic approach to the evaluation of autism. The evaluations and testing in the higher (earlier) tiers were selected on the basis of the predicted incidence of the disorder in patients with ASD, the invasiveness of the testing, the potential of intervention, and the overall practicality of obtaining tests. In our experience, this approach has been met with a high level of acceptance from third-party payers and the families. For this study, we reviewed our clinical data for a 3-year period from 2002 to 2004. We evaluated the effectiveness of our diagnostic strategy in patients with ASD and estimated its diagnostic yield.
MATERIALS AND METHODS
Study design
We performed a retrospective search for the patients referred to us for clinical genetic consultation with the primary diagnosis of ASD for the years 2002 to 2004 inclusive. Patients selected for further review all had an Axis I diagnosis of ASD made by a qualified specialist in autism. We identified 32 patients who met these criteria. Because these patients were evaluated by one of the two of us, all had undergone systematic diagnostic testing using the evaluation approach we had developed (see “Evaluation Scheme”). We then reviewed the results of the diagnostic evaluations of patients seen during this time period. These patients were reported as either having no identifiable diagnosis or characterized by an etiologic diagnosis and what part of the tiered evaluation their diagnosis corresponded with.
Evaluation scheme
During the course of seeing and evaluating patients with ASD, we developed a carefully considered diagnostic approach (Table 1). This approach was derived from our clinical experience and published data (reviewed above) on identifiable diagnoses in individuals with ASD.16–21 We decided to evaluate these patients in a stepwise (tiered) manner. For the first two tiers, we chose to organize our studies to correlate with that previously reported by Shevell and colleagues,21 on the basis of our agreement with their diagnostic philosophy.
Tier 1
The initial assessment was a standard clinical genetic consultation (history and physical) to identify known syndromes or associated conditions. This included a dysmorphologic/clinical genetics examination with a Wood's lamp evaluation. If a diagnosis was suspected on the basis of this initial assessment, we proceeded to targeted testing. In addition, “standard” metabolic screening for urine mucopolysaccharides and organic acids, as well as serum lactate, amino acids, ammonia, and acyl-carnitine profiles, was performed. All patients had sensory screening (to rule out a child with suspected autism who actually just had a hearing loss). Last, rubella titers were performed only if clinical indicators were present.
Tier 2
The second tier included prometaphase chromosomes, brain neuroimaging (magnetic resonance imaging), and DNA studies for Fragile X. In addition, an electroencephalogram (EEG) was performed. If the prometaphase chromosome study results were normal, and the patient had been noted to have clinically significant pigmentary changes on physical examination, a skin biopsy was obtained and a karyotype was performed on cultured fibroblasts.
Tier 3
In patients for whom no diagnosis was found in the first two tiers, we proceeded to a third tier of evaluations. In this tier we included chromosome 22q11 fluorescence in situ hybridization (FISH) studies, methyl-CpG-binding protein (MECP)-2 gene testing, chromosome 15 studies (15q11 FISH, interphase centromeric FISH, and methylation), 17p11.2 FISH, subtelomeric FISH panel (if intelligence quotient <50), and serum and urine uric acid testing. If an increased production of uric acid was noted, then hypoxanthine-guanine phosphoribosyl transferase and phosphoribosylpyrophosphate synthetase superactivity testing were performed. If decreased production was found, we performed a purine and pyrimidine metabolic panel (uracil excretion, xanthine, and hypoxanthine levels).
RESULTS
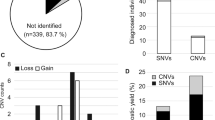
The results of our study are summarized in Table 2. We started with the cohort of 32 patients ascertained as having been referred for genetic evaluation with a diagnosis of autism. In the first two tiers of the evaluation we identified one patient each with a hearing loss (and not autism), neurofibromatosis, tuberous sclerosis, and Sotos syndrome. The patient with tuberous sclerosis is particularly notable in that no external (cutaneous) features were seen on examination. However, neuroimaging identified classic brain findings of tuberous sclerosis. Reexamination (again with a Wood's lamp) after the brain findings were discovered did not identify any overlooked abnormalities. We found two patients with Fragile X syndrome. Two patients had chromosome abnormalities (46 XY, dup22[q12.1 q11.23] and 46 XX, del9[p24.1]). Thus, the diagnostic yield of the first two tiers (8/32 patients) was 25%. Two additional patients were identified in the first two tiers as having cerebral dysgenesis (ventral induction defects) as part of their evaluation. We think these patients also have a positive finding on their diagnostic workup (thus giving us a 10/32 or a 30% yield). However, to avoid controversy over whether these represent a true causal relationship, we chose not to include them in our “positives” for the sake of this report.
In the third tier of the evaluation, another five patients (16%) had a positive diagnostic workup. These diagnoses were a duplication of chromosome 15 [46XY,trp(15)(q11.2q12)], two patients with MECP-2 mutations, one patient with a 17p11 deletion, and one patient with persistently low serum uric acid levels. Reinspection of the patient with the 17p11 deletion showed what was believed to be subtle findings suggestive of Smith-Magenis syndrome not seen on the initial examination. We were not able to identify the primary metabolic defect in the patient with the persistently low serum uric acid levels. However, the levels were significantly below normal on repeated blood draws, leaving little doubt that he has some, as yet unidentifiable, inborn error of purine/pyrimidine metabolism.
The summary of these data gives a calculated total diagnostic yield of 13/32 or 41%.
CONCLUSION
The last several years have produced a dramatic increase in the number of tests available to the clinical geneticist. As these tests have been applied to the etiologic workup of patients with ASDs, a host of new diagnostic associations have been reported. These include subtle (previously undetectable) chromosome abnormalities, atypical expression (expanded phenotypes) of known genetic syndromes, newly discovered inborn errors of metabolism, and specific gene mutations.
Chromosomal abnormalities
Advances in cytogenetic technology including prometaphase and late interphase studies have become the standard for clinical testing. With the use of higher resolution techniques, many chromosome aneuploidies have been reported in individuals with ASDs.6,25–30 In particular, deletions or duplications of 15q and 22q are frequently seen in patients with ASDs.29,31–38 (Of note, interphase FISH has traditionally been needed for confirmation of isodicentric chromosome 15 duplication.) The well-described relationship between pigmentary changes and chromosomal mosaicism has direct application in the evaluation of ASDs.39,40
The application of FISH greatly enhances detection rates in neurogenetic evaluations. Current clinical applications of FISH technology include single-locus FISH, subtelomeric FISH panels, and chromosomal microarray (CMA) studies. Single-locus FISH studies have been reported to be positive in patients with ASDs for 22q11, 22q13, and 17p11.2.37,38,41–44 The addition of a 40-probe FISH panel specific to the subtelomeric regions of the chromosomes has had a quantum effect in the evaluation of mental retardation. More recently, there have been assessments of its applicability in patients with ASDs. One study found 1 of 10 patients with idiopathic autism and moderate mental retardation had a chromosome 2q subtelomeric deletion.45 Not surprisingly, the conclusion from this study was that it was too early to recommend universal screening for subtelomeric deletions in all patients with autism. Other studies have also indicated a link between terminal deletions of 2q37 and autism.46–49 (However, the possible association of this subtelomeric deletion with ASD has been questioned. Deletions, and duplications, of this locus occur frequently as an apparent normal polymorphism in the general population.) Subsequent studies have not found subtelomeric deletions in larger cohorts of children with ASD.50,51 At present, then, there are not enough supporting data to conclusively determine the utility of subtelomeric FISH panels in ASDs. Still, this test panel is absolutely indicated in patients with intelligence quotients less than 50.52 Because mental retardation is a common comorbid condition with ASDs, one could surmise that subtelomeric FISH studies be performed in the subgroup of patients with ASD and severe to profound mental retardation. However, the most recent study suggested that this may actually be a low yield endeavor.50 To our knowledge, CMA studies have not been reported in cohorts with ASD with the exception of studies of 15q.
Genetic syndromes
The association of autism/autistic behaviors with phakomatoses (tuberous sclerosis and neurofibromatosis) has been known for some time.53,54 Similar to the discussions concerning Fragile X syndrome and autism, there is not complete agreement about the nature of the cause–effect relationship. Regardless, the incidence of autism is significantly increased in these two disorders compared with that of the general population.
Newer diagnostic associations have been reported with subtle/atypical expression of defined genetic syndromes and autism. The concept of an “expanded phenotype” of recognizable syndromes has been emerging over the past decade. In recent years, genotypic changes at loci associated with recognizable syndromes have been reported in individuals with ASD, but with few, if any, of the somatic features typically seen with these conditions.55 This includes Rett syndrome (MECP-2 mutations),56–58 Smith-Lemli-Opitz syndrome (7 dehydrocholesterol enzymatic block),7,59 Angelman syndrome (15 methylation and UBE3A abnormalities),60–62 Smith-Magenis syndrome (FISH detectable 17p deletions),7,43 Sotos syndrome (NSD1 mutations),63 Banayaan-Riley-Ruvalcaba syndrome (PTEN mutations),64,65 and ARX gene-associated disorders.66
Metabolic disorders
Several of the “classic” metabolic disorders have been associated with autistic behavior. Individuals with untreated phenylketonuria are often said to be autistic. With current programs in newborn screening, it is hoped that this is a cause of ASDs that we will no longer see. Children with mucopolysaccharide storage disorders look strikingly similar to those with the reported “autistic regressive syndrome” in the early stages of clinical symptoms. Previous studies on diagnostic yields in ASDs have typically included metabolic screening for these and other metabolic disorders such as aminoacidopathies, organic acidurias, urea cycle defects, and fatty acid oxidation disorders.21
The most rapidly expanding diagnostic associations with ASDs have been the identification of a variety of metabolic disorders in which phenotype appears to be simply that of ASDs. This includes disorders of purine and pyrimidine metabolism, sulfation disorders, neurotransmitters, and creatine metabolism (Table 3).67–79 In addition, mitochondrial abnormalities have been reported in association with an autism phenotype.80–82
Single gene disorders
The association of Fragile X syndrome and autism is well documented.28,83–91 Some debate still exists as to the exact nature of autistic behaviors in Fragile X. Yet it is clear that the Fragile X is a frequent finding in persons with ASDs. Other rare folate-sensitive fragile sites have been reported in autistic patients. Some (2q13, 6p23, 12q13) have only been reported in autistic individuals.92–94
This study affirms previous documentation of the utility of a comprehensive clinical genetic evaluation of individuals with the diagnosis of autism. Recent studies indicate that such evaluations are even applicable across the milder aspects of the spectrum of pervasive developmental disorders.15 This study demonstrates a significant increase in the diagnostic yield over earlier reports of the evaluation of persons with ASDs. Our overall diagnostic yield was more than 40% in this report.
This was a retrospective, non-epidemiologic review. Still, it is an accurate reflection of the impact of autism as it presents to a clinical genetics practice. The patients in this study were selected only in that they were referred for a genetic evaluation. It is difficult to imagine what bias this might introduce. Our tiered system of evaluation is practical and reasonable. In our experience, it is well accepted by third-party payers and the patients and their families. Careful explanation of the process before beginning the evaluation seemed to greatly decrease obstacles in completing the evaluation.
On review of our experience and the data from this study, we identified several important insights into this protocol. One of the most important factors we discovered was the importance of accurate diagnostics on the “front side” of the evaluation. Our patient with the hearing loss highlights the importance of documenting normal hearing before making the diagnosis of autism and/or proceeding with a diagnostic workup. In addition, there still remains a large diversity in the way the diagnosis of autism is made. To avoid stigmatization and unnecessary evaluations, we have initiated an autism screening, diagnostic, and triage clinic. Only those patients accurately assessed as having autism are then directed toward our tiered genetic evaluation protocol. We also found that having a clinical geneticist as part of this “first contact” team is invaluable. Many straightforward genetic diagnoses have been made at the time of the initial visit.
For the past year we have used a modified protocol from that used in this study (Table 4). Changes were made by reviewing the outcomes of this study and in incorporating newer technologies. Although it is a little too early to make firm conclusions, our initial look at the numbers for the past year suggests that this will further increase the diagnostic yield to more than 50%. Insights we have gleaned and modifications we have made include the following.
-
1
We found cytogenetic studies to be the most likely to give an answer. This is in agreement with almost all previous reports. Cytogenetic testing should remain a component of the higher tier tests.
-
2
The introduction of CMA studies has tremendously increased the diagnostic yield in mental retardation. We have placed CMA testing in our second tier of our ASD evaluation scheme. This makes sense at all levels. It is in keeping with the high yield of cytogenetic studies as mentioned under point number 1. In addition, there is a good argument for the cost-effectiveness of this decision. Clinical CMA panels would subsume several of the recommended tests in our original protocol including 22q11 FISH, 15q11-13 FISH, 17p11FISH, chromosome 15 centromeric studies, and the subtelomeric FISH panel. In addition, these studies include hundreds to thousands of other probes not otherwise tested. The cost of a CMA study is considerably less than the cost of multiple individual probes. At our institution, in fact, it becomes less expensive to perform CMA studies once three or more individual FISH studies are requested.
-
3
For this study, we chose to include “standard” metabolic testing in the first tier to make a balanced comparison with previous studies. In this study, no patients were identified with metabolic disorders. In reviewing the results of other studies, we could not identify data to support this as a high-yield modality. As such, we modified our approach to perform standard metabolic testing only in the presence of clinical indicators.
-
4
To date, we have found no reports of males with an MECP-2 mutation with the sole phenotypic expression being ASD. In this study, the two patients identified with MECP-2 mutations were female. Thus, we suggest performing MECP-2 gene testing only in females with ASDs until such time that there are data to support otherwise.
Although we are excited about the significant improvements in the diagnostic yield of patients with ASDs, there still remains more than half of the patients without a unifying diagnosis. We predict that the addition of other tests to our evaluation protocol would further increase diagnostic success. Table 5 lists several other clinically available tests for conditions that have been reported in association with an ASD phenotype. It is unclear to what degree the addition of any of these tests would add to diagnostic yield. Ideally, a large-scale, multicenter collaborative study of the utility of these and other potentially informative tests is needed to answer this important question.
We acknowledge several controversies in our recommended approach to the clinical genetic evaluation of patients with ASDs.
-
1
Our testing protocol is reasonably aggressive. Some groups have not recommended as much testing.22 Given the implications of these diagnoses on recurrence risks and associated medical conditions, we believe that an aggressive neurogenetic evaluation of all persons with ASD is warranted. We consider our diagnostic yield more than justification for this approach.
-
2
Many families with children with autism request an EEG. The impetus for these requests is the hope that the underlying diagnosis may actually be Landau-Kleffner syndrome, and thus a “treatable” form of autism. It has been argued that the low incidence of Landau-Kleffner makes using an EEG a standard part of the evaluation of ASDs impractical. Still, the incidence of epileptiform disorders in persons with ASDs has been reported to be as high as 65%.95 Thus, an EEG seems to be medically important for either instance. As such, we have placed this study in the “pre-evaluation” arm of our protocol.
-
3
Likewise, some would question the rationale of routine neuroimaging in patients with ASDs. Our patient with tuberous sclerosis and no cutaneous findings provides a clear answer of the utility of this testing modality. In addition, two of our patients had subtle markers of cerebral dysgenesis. Although the identification of such markers does not provide complete insight into cause, we still find knowledge of such anomalies helpful to clinicians and families.96
This study supports the work of others documenting the utility of a clinical genetic evaluation of patients with ASDs. With our protocol, we expect more than a 40% diagnostic yield. For these patients, detailed diagnosis-specific genetic counseling can be provided. For those in whom no diagnosis is found, empiric recurrence risk counseling can be provided. Currently the literature suggests that the recurrence risk for siblings is 4% if the affected child is a female and 7% if the affected child is a male. If there are two affected children, the recurrence risk is 25%. Thus, clinical geneticists can provide significant and meaningful input into the case management of every individual with ASD/pervasive developmental disorder-not otherwise specified. As such, a genetic consultation should be considered for all such patients.
References
Cook EH . Genetics of autism. Child Adolesc Psychiatr Clin N Am 2001; 10( 2): 333–350.
Lotspeich LJ, Ciaranello RD . The neurobiology and genetics of infantile autism. Int Rev Neurobiol 1993; 35: 87–129.
Muhle R, Trentacoste SV, Rapin I . The genetics of autism. Pediatrics 2004; 113( 5): e472–e486.
Simonoff E . Genetic counseling in autism and pervasive developmental disorders. J Autism Dev Disord 1998; 28( 5): 447–456.
Spence SJ . The genetics of autism. Semin Pediatr Neurol 2004; 11( 3): 196–204.
Castermans D, Wilquet V, Steyaert J, Van de Ven W, et al. Chromosomal anomalies in individuals with autism: a strategy towards the identification of genes involved in autism. Autism 2004; 8( 2): 141–161.
Cohen D, Pichard N, Tordjman S, Bauman C, et al. Specific genetic disorders and autism: clinical contribution towards their identification. J Autism Dev Disord 2005; 35( 1): 103–116.
Folstein SE, Piven J . Etiology of autism: genetic influences. Pediatrics 1991; 87( 5 Pt 2): 767–773.
Mariner R, Jackson AW, Levitas A, Hagerman RJ, et al. Autism, mental retardation, and chromosomal abnormalities. J Autism Dev Disord 1986; 16( 4): 425–440.
Reichelt KL, Saelid G, Lindback T, Boler JB . Childhood autism: a complex disorder. Biol Psychiatry 1986; 21( 13): 1279–1290.
Rutter M . Aetiology of autism: findings and questions. J Intellect Disabil Res 2005; 49( Pt 4): 231–238.
Rutter M, Bailey A, Bolton P, Le Couteur A . Autism and known medical conditions: myth and substance. J Child Psychol Psychiatry 1994; 35( 2): 311–322.
Steiner CE, Guerreiro MM, Marques-de-Faria AP . Genetic and neurological evaluation in a sample of individuals with pervasive developmental disorders. Arq Neuropsiquiatr 2003; 61( 2A): 176–180.
Trottier G, Srivastava L, Walker CD . Etiology of infantile autism: a review of recent advances in genetic and neurobiological research. J Psychiatry Neurosci 1999; 24( 2): 103–115.
Abdul-Rahman OA, Hudgins L . The diagnostic utility of a genetics evaluation in children with pervasive developmental disorders. Genet Med 2006; 8( 1): 50–54.
Battaglia A, Carey JC . Etiologic yield of autistic spectrum disorders: A prospective study. Am J Med Genet C Semin Med Genet 2006; 142( 1): 3–7.
Challman TD, Barbaresi WJ, Katusic SK, Weaver A . The yield of the medical evaluation of children with pervasive developmental disorders. J Autism Dev Disord 2003; 33( 2): 187–192.
Chudley AE, Gutierrez E, Jocelyn LJ, Chodirker BN . Outcomes of genetic evaluation in children with pervasive developmental disorder. J Dev Behav Pediatr 1998; 19( 5): 321–325.
Kosinovsky B, Hermon S, Yoran-Hegesh R, Golomb A, et al. The yield of laboratory investigations in children with infantile autism. J Neural Transm 2005; 112( 4): 587–596.
Miles JH, Hillman RE . Value of a clinical morphology examination in autism. Am J Med Genet 2000; 91( 4): 245–253.
Shevell MI, Majnemer A, Rosenbaum P, Abrahamowicz M . Etiologic yield of autistic spectrum disorders: a prospective study. J Child Neurol 2001; 16( 7): 509–512.
Filipek PA, Accardo PJ, Ashwal S, Baranek GT, et al. Practice parameter: screening and diagnosis of autism: report of the Quality Standards Subcommittee of the American Academy of Neurology and the Child Neurology Society. Neurology 2000; 55( 4): 468–479.
Hillman RE, Kanafani N, Takahashi TN, Miles JH . Prevalence of autism in Missouri: changing trends and the effect of a comprehensive state autism project. Mo Med 2000; 97( 5): 159–163.
Voigt RG, Dickerson CL, Reynolds AM, Childers DO, et al. Laboratory evaluation of children with autistic spectrum disorders: a guide for primary care pediatricians. Clin Pediatr (Phila) 2000; 39( 11): 669–671.
Barrett S, Beck JC, Bernier R, Bisson E, et al. An autosomal genomic screen for autism. Collaborative linkage study of autism. Am J Med Genet 1999; 88( 6): 609–615.
Blackman JA, Selzer SC, Patil S, Van Dyke DC . Autistic disorder associated with an iso-dicentric Y chromosome. Dev Med Child Neurol 1991; 33( 2): 162–166.
Carratala F, Galan F, Moya M, Estivill X, et al. A patient with autistic disorder and a 20/22 chromosomal translocation. Dev Med Child Neurol 1998; 40( 7): 492–495.
Reddy KS . Cytogenetic abnormalities and fragile-X syndrome in autism spectrum disorder. BMC Med Genet 2005; 6: 3.
Vorstman JA, Staal WG, van Daalen E, van Engeland H, et al. Identification of novel autism candidate regions through analysis of reported cytogenetic abnormalities associated with autism. Mol Psychiatry 2006; 11( 1): 1, 18–28.
Weidmer-Mikhail E, Sheldon S, Ghaziuddin M . Chromosomes in autism and related pervasive developmental disorders: a cytogenetic study. J Intellect Disabil Res 1998; 42( Pt 1): 8–12.
Schroer RJ, Phelan MC, Michaelis RC, Crawford EC, et al. Autism and maternally derived aberrations of chromosome 15q. Am J Med Genet 1998; 76( 4): 327–336.
Bundey S, Hardy C, Vickers S, Kilpatrick MW, et al. Duplication of the 15q11-13 region in a patient with autism, epilepsy and ataxia. Dev Med Child Neurol 1994; 36( 8): 736–742.
Bolton PF, Dennis NR, Browne CE, Thomas NS, et al. The phenotypic manifestations of interstitial duplications of proximal 15q with special reference to the autistic spectrum disorders. Am J Med Genet 2001; 105( 8): 675–685.
Wolpert C, Pericak-Vance MA, Abramson RK, Wright HH, et al. Autistic symptoms among children and young adults with isodicentric chromosome 15. Am J Med Genet 2000; 96( 1): 128–129.
Bass MP, Menold MM, Wolpert CM, Donelly SL, et al. Genetic studies in autistic disorder and chromosome 15. Neurogenetics 2000; 2( 4): 219–226.
Manning MA, Cassidy SB, Clericuzio C, Cherry AM, et al. Terminal 22q deletion syndrome: a newly recognized cause of speech and language disability in the autism spectrum. Pediatrics 2004; 114( 2): 451–457.
Prasad C, Prasad AN, Chodirker BN, Lee C, et al. Genetic evaluation of pervasive developmental disorders: the terminal 22q13 deletion syndrome may represent a recognizable phenotype. Clin Genet 2000; 57( 2): 103–109.
Goizet C, Excoffier E, Taine L, Taupiac E, et al. Case with autistic syndrome and chromosome 22q13.3 deletion detected by FISH. Am J Med Genet 2000; 96( 6): 839–844.
Zappella M . Autism and hypomelanosis of Ito in twins. Dev Med Child Neurol 1993; 35( 9): 826–832.
Akefeldt A, Gillberg C . Hypomelanosis of Ito in three cases with autism and autistic-like conditions. Dev Med Child Neurol 1991; 33( 8): 737–743.
Fine SE, Weissman A, Gerdes M, Pinto-Martin J, et al. Autism spectrum disorders and symptoms in children with molecularly confirmed 22q11.2 deletion syndrome. J Autism Dev Disord 2005; 35( 4): 461–470.
Niklasson L, Rasmussen P, Oskarsdottir S, Gillberg C . Attention deficits in children with 22q. 11 deletion syndrome. Dev Med Child Neurol 2005; 47( 12): 803–807.
Park JP, Moeschler JB, Davies WS, Patel PI, et al. Smith-Magenis syndrome resulting from a de novo direct insertion of proximal 17q into 17p11.2. Am J Med Genet 1998; 77( 1): 23–27.
Simon TJ, Bish JP, Bearden CE, Ding L, et al. A multilevel analysis of cognitive dysfunction and psychopathology associated with chromosome 22q11.2 deletion syndrome in children. Dev Psychopathol 2005; 17( 3): 753–784.
Wolff DJ, Clifton K, Karr C, Charles J . Pilot assessment of the subtelomeric regions of children with autism: detection of a 2q deletion. Genet Med 2002; 4( 1): 10–14.
Lukusa T, Smeets E, Vogels A, Vermeesch JR, et al. Terminal 2q37 deletion and autistic behaviour. Genet Couns 2005; 16( 2): 179–180.
Wassink TH, Piven J, Vieland VJ, Jenkins L, et al. Evaluation of the chromosome 2q37.3 gene CENTG2 as an autism susceptibility gene. Am J Med Genet B Neuropsychiatr Genet 2005; 136( 1): 36–44.
Lukusa T, Vermeesch JR, Holvoet M, Fryns JP, et al. Deletion 2q37.3 and autism: molecular cytogenetic mapping of the candidate region for autistic disorder. Genet Couns 2004; 15( 3): 293–301.
Ghaziuddin M, Burmeister M . Deletion of chromosome 2q37 and autism: a distinct subtype?. J Autism Dev Disord 1999; 29( 3): 259–263.
Battaglia A, Bonaglia MC . The yield of subtelomeric FISH analysis in the evaluation of autistic spectrum disorders. Am J Med Genet C Semin Med Genet 2006; 142( 1): 8–12.
Nair-Miranda K, Murch A, Petterson B, Hill W, et al. An investigation into sub-telomeric deletions of chromosome 22 and pervasive developmental disorders. Am J Med Genet B Neuropsychiatr Genet 2004; 125( 1): 99–104.
Baroncini A, Rivieri F, Capucci A, Croci G, et al. FISH screening for subtelomeric rearrangements in 219 patients with idiopathic mental retardation and normal karyotype. Eur J Med Genet 2005; 48( 4): 388–396.
Wiznitzer M . Autism and tuberous sclerosis. J Child Neurol 2004; 19( 9): 675–679.
Mouridsen SE, Andersen LB, Sorensen SA, Rich B, et al. Neurofibromatosis in infantile autism and other types of childhood psychoses. Acta Paedopsychiatr 1992; 55( 1): 15–18.
Artigas-Pallares J, Gabau-Vila E, Guitart-Feliubadalo M . [Syndromic autism: II. Genetic syndromes associated with autism.]. Rev Neurol 2005; 40( Suppl 1): S151–S162.
Carney RM, Wolpert CM, Ravan SA, Shahbazian M, et al. Identification of MeCP2 mutations in a series of females with autistic disorder. Pediatr Neurol 2003; 28( 3): 205–211.
Zappella M, Meloni I, Longo I, Canitano R, et al. Study of MECP2 gene in Rett syndrome variants and autistic girls. Am J Med Genet B Neuropsychiatr Genet 2003; 119( 1): 102–107.
Hammer S, Dorrani N, Dragich J, Kudo S, et al. The phenotypic consequences of MECP2 mutations extend beyond Rett syndrome. Ment Retard Dev Disabil Res Rev 2002; 8( 2): 94–98.
Tierney E, Nwokoro NA, Porter FD, Freund LS, et al. Behavior phenotype in the RSH/Smith-Lemli-Opitz syndrome. Am J Med Genet 2001; 98( 2): 191–200.
Nurmi EL, Bradford Y, Chen Y, Hall J, et al. Linkage disequilibrium at the Angelman syndrome gene UBE3A in autism families. Genomics 2001; 77( 1-2): 105–113.
Lopez-Rangel E, Lewis ME . Further evidence for epigenetic influence of MECP2 in Rett, autism and Angelman's syndromes. Clin Genet 2006; 69( 1): 23–25.
Williams CA, Lossie A, Driscoll D . Angelman syndrome: mimicking conditions and phenotypes. Am J Med Genet 2001; 101( 1): 59–64.
Morrow JD, Whitman BY, Accardo PJ . Autistic disorder in Sotos syndrome: a case report. Eur J Pediatr 1990; 149( 8): 567–569.
Butler MG, Dasouki MJ, Zhou XP, Talebizadeh Z, et al. Subset of individuals with autism spectrum disorders and extreme macrocephaly associated with germline PTEN tumour suppressor gene mutations. J Med Genet 2005; 42( 4): 318–321.
Goffin A, Hoefsloot LH, Bosgoed E, Swillen A, et al. PTEN mutation in a family with Cowden syndrome and autism. Am J Med Genet 2001; 105( 6): 521–524.
Sherr EH . The ARX story (epilepsy, mental retardation, autism, and cerebral malformations): one gene leads to many phenotypes. Curr Opin Pediatr 2003; 15( 6): 567–571.
Fon EA, Sarrazin J, Meunier C, Alarcia J, et al. Adenylosuccinate lyase (ADSL) and infantile autism: absence of previously reported point mutation. Am J Med Genet 1995; 60( 6): 554–557.
Jaeken J, Van den Berghe G . An infantile autistic syndrome characterised by the presence of succinylpurines in body fluids. Lancet 1984; 2( 8411): 1058–1061.
Kohler M, Assmann B, Brautigam C, Storm W, et al. Adenylosuccinase deficiency: possibly underdiagnosed encephalopathy with variable clinical features. Eur J Paediatr Neurol 1999; 3( 1): 3–6.
Maddocks J, Reed T . Urine test for adenylosuccinase deficiency in autistic children. Lancet 1989; 1( 8630): 158–159.
Micheli V, Rocchigiani M, Pompucci G . An HPLC-linked assay of phosphoribosylpyrophosphate synthetase activity in the erythrocytes of adults and children with neurological disorders. Clin Chim Acta 1994; 227( 1-2): 79–86.
Nicholson JF . Metabolic disease. Behavioral aspects. N Y State J Med 1975; 75( 7): 1044–1046.
Page T, Coleman M . Purine metabolism abnormalities in a hyperuricosuric subclass of autism. Biochim Biophys Acta 2000; 1500( 3): 291–296.
Pearl PL, Gibson KM, Acosta MT, Vezina LG, et al. Clinical spectrum of succinic semialdehyde dehydrogenase deficiency. Neurology 2003; 60( 9): 1413–1417.
Pesi R, Micheli V, Jacomelli G, Peruzzi L, et al. Cytosolic 5′-nucleotidase hyperactivity in erythrocytes of Lesch-Nyhan syndrome patients. Neuroreport 2000; 11( 9): 1827–1831.
Salerno C, Siems WG, Crifo C . Succinylpurinemic autism: increased sensitivity of defective adenylosuccinate lyase towards 4-hydroxy-2-nonenal. Biochim Biophys Acta 2000; 1500( 3): 335–341.
Simmonds HA . When and how does one search for inborn errors of purine and pyrimidine metabolism?. Pharm World Sci 1994; 16( 2): 139–148.
Stathis SL, Cowley DM, Broe D . Autism and adenylosuccinase deficiency. J Am Acad Child Adolesc Psychiatry 2000; 39( 3): 274–275.
Stone RL, Aimi J, Barshop BA, Jaeken J, et al. A mutation in adenylosuccinate lyase associated with mental retardation and autistic features. Nat Genet 1992; 1( 1): 59–63.
Segurado R, Conroy J, Meally E, Fitzgerald M, et al. Confirmation of association between autism and the mitochondrial aspartate/glutamate carrier SLC25A12 gene on chromosome 2q31. Am J Psychiatry 2005; 162( 11): 2182–2184.
Clark-Taylor T, Clark-Taylor BE . Is autism a disorder of fatty acid metabolism? Possible dysfunction of mitochondrial beta-oxidation by long chain acyl-CoA dehydrogenase. Med Hypotheses 2004; 62( 6): 970–975.
Lerman-Sagie T, Leshinsky-Silver E, Watemberg N, Lev D . Should autistic children be evaluated for mitochondrial disorders?. J Child Neurol 2004; 19( 5): 379–381.
Hagerman RJ, Chudley AE, Knoll JH, Jackson AW, et al. Autism in fragile X females. Am J Med Genet 1986; 23( 1-2): 375–380.
Vincent JB, Melmer G, Bolton PF, Hodgkinson S, et al. Genetic linkage analysis of the X chromosome in autism, with emphasis on the fragile X region. Psychiatr Genet 2005; 15( 2): 83–90.
Goodlin-Jones BL, Tassone F, Gane LW, Hagerman RJ . Autistic spectrum disorder and the fragile X premutation. J Dev Behav Pediatr 2004; 25( 6): 392–398.
Kaufmann WE, Cortell R, Kau AS, Bukelis I, et al. Autism spectrum disorder in fragile X syndrome: communication, social interaction, and specific behaviors. Am J Med Genet A 2004; 129( 3): 225–234.
Kandil ST, Bilici M, Aksu HB, Celep F, et al. Early infantile autism and fragile X anomaly. Isr J Psychiatry Relat Sci 2004; 41( 1): 70.
Sabaratnam M, Murthy NV, Wijeratne A, Buckingham A, et al. Autistic-like behaviour profile and psychiatric morbidity in Fragile X Syndrome: a prospective ten-year follow-up study. Eur Child Adolesc Psychiatry 2003; 12( 4): 172–177.
Rogers SJ, Wehner DE, Hagerman R . The behavioral phenotype in fragile X: symptoms of autism in very young children with fragile X syndrome, idiopathic autism, and other developmental disorders. J Dev Behav Pediatr 2001; 22( 6): 409–417.
Hagerman RJ, Jackson AW . Autism or fragile X syndrome?. J Am Acad Child Psychiatry 1985; 24( 2): 239–240.
Blomquist HK, Bohman M, Edvinsson SO, Gillberg C, et al. Frequency of the fragile X syndrome in infantile autism. A Swedish multicenter study. Clin Genet 1985; 27( 2): 113–117.
Arrieta I, Nunez T, Gil A, Flores P, et al. Autosomal folate sensitive fragile sites in an autistic Basque sample. Ann Genet 1996; 39( 2): 69–74.
Samadder P, Evans JA, Chudley AE . Segregation analysis of rare autosomal folate sensitive fragile sites. Am J Med Genet 1993; 46( 2): 165–171.
Saliba JR, Griffiths M . Brief report: autism of the Asperger type associated with an autosomal fragile site. J Autism Dev Disord 1990; 20( 4): 569–575.
Reinhold JA, Molloy CA, Manning-Courtney P . Electroencephalogram abnormalities in children with autism spectrum disorders. J Neurosci Nurs 2005; 37( 3): 136–138.
Schaefer GB, Sheth RD, Bodensteiner JB . Cerebral dysgenesis. An overview. Neurol Clin 1994; 12( 4): 773–788.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Schaefer, G., Lutz, R. Diagnostic yield in the clinical genetic evaluation of autism spectrum disorders. Genet Med 8, 549–556 (2006). https://doi.org/10.1097/01.gim.0000237789.98842.f1
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1097/01.gim.0000237789.98842.f1
This article is cited by
-
Detection of autism spectrum disorder-related pathogenic trio variants by a novel structure-based approach
Molecular Autism (2024)
-
Does Every Child With Autism Need Investigations for Inborn Errors of Metabolism?
Indian Pediatrics (2023)
-
Identification of mutations in the PI3K-AKT-mTOR signalling pathway in patients with macrocephaly and developmental delay and/or autism
Molecular Autism (2017)
-
Genetic Studies in Autism
The Indian Journal of Pediatrics (2016)
-
Magnetic Resonance Spectroscopy Studies of Glutamate and GABA in Autism: Implications for Excitation-Inhibition Imbalance Theory
Current Developmental Disorders Reports (2015)