Abstract
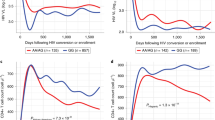
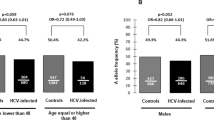
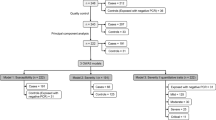
Inflammasomes are multi-protein complexes integrating pathogen-triggered signaling leading to the generation of pro-inflammatory cytokines including interleukin-18 (IL-18). Hepatitis C virus (HCV) and human immunodeficiency virus (HIV) infections are associated with elevated IL-18, suggesting inflammasome activation. However, there is marked person-to-person variation in the inflammasome response to HCV and HIV. We hypothesized that host genetics may explain this variation. To test this, we analyzed the associations of plasma IL-18 levels and polymorphisms in 10 genes in the inflammasome cascade. About 1538 participants with active HIV and/or HCV infection in three ancestry groups are included. Samples were genotyped using the Illumina Omni 1-quad and Omni 2.5 arrays. Linear regression analyses were performed to test the association of variants with log IL-18 including HCV and HIV infection status, and HIV RNA in each ancestry group and then meta-analyzed. Eleven highly correlated single-nucleotide polymorphisms (r2=0.98–1) in the IL-18-BCO2 region were significantly associated with log IL-18; each T allele of rs80011693 confers a decrease of 0.06 log pg ml−1 of IL-18 after adjusting for covariates (rs80011693; rs111311302 β=−0.06, P-value=2.7 × 10−4). In conclusion, genetic variation in IL-18 is associated with IL-18 production in response to HIV and HCV infection, and may explain variability in the inflammatory outcomes of chronic viral infections.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 6 digital issues and online access to articles
$119.00 per year
only $19.83 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Guo H, Callaway JB, Ting JP . Inflammasomes: mechanism of action, role in disease, and therapeutics. Nat Med 2015; 21: 677–687.
Chattergoon MA, Latanich R, Quinn J, Winter ME, Buckheit RW 3rd, Blankson JN et al. HIV and HCV activate the inflammasome in monocytes and macrophages via endosomal toll-like receptors without induction of type 1 interferon. PLoS Pathog 2014; 10: e1004082.
Kanneganti TD . Central roles of NLRs and inflammasomes in viral infection. Nat Rev Immunol 2010; 10: 688–698.
Netea MG, Simon A, van de Veerdonk F, Kullberg BJ, Van der Meer JW, Joosten LA . IL-1beta processing in host defense: beyond the inflammasomes. PLoS Pathog 2010; 6: e1000661.
Dinarello CA . Interleukin-1 in the pathogenesis and treatment of inflammatory diseases. Blood 2011; 117: 3720–3732.
Strowig T, Henao-Mejia J, Elinav E, Flavell R . Inflammasomes in health and disease. Nature 2012; 481: 278–286.
Chattergoon MA, Levine JS, Latanich R, Osburn WO, Thomas DL, Cox AL . High plasma interleukin-18 levels mark the acute phase of hepatitis C virus infection. J Infect Dis 2011; 204: 1730–1740.
Sharma A, Chakraborti A, Das A, Dhiman RK, Chawla Y . Elevation of interleukin-18 in chronic hepatitis C: implications for hepatitis C virus pathogenesis. Immunology 2009; 128 (1 Suppl): e514–e522.
Jia H, Du J, Zhu S, Ma Y, Cai H . Clinical observation of serum IL-18, IL-10 and sIL-2R levels in patients with chronic hepatitis C pre- and post antiviral treatment. Chin Med J 2003; 116: 605–608.
Iannello A, Boulassel MR, Samarani S, Tremblay C, Toma E, Routy JP et al. HIV-1 causes an imbalance in the production of interleukin-18 and its natural antagonist in HIV-infected individuals: implications for enhanced viral replication. J Infect Dis 2010; 201: 608–617.
Samarani S, Allam O, Sagala P, Aldabah Z, Jenabian MA, Mehraj V et al. Imbalanced production of IL-18 and its antagonist in human diseases, and its implications for HIV-1 infection. Cytokine 2016; 82: 38–51.
Matteini AM, Li J, Lange EM, Tanaka T, Lange LA, Tracy RP et al. Novel gene variants predict serum levels of the cytokines IL-18 and IL-1ra in older adults. Cytokine 2014; 65: 10–16.
Melzer D, Perry JR, Hernandez D, Corsi AM, Stevens K, Rafferty I et al. A genome-wide association study identifies protein quantitative trait loci (pQTLs). PLoS Genet 2008; 4: e1000072.
He M, Cornelis MC, Kraft P, van Dam RM, Sun Q, Laurie CC et al. Genome-wide association study identifies variants at the IL18-BCO2 locus associated with interleukin-18 levels. Arterioscler Thromb Vasc Biol 2010; 30: 885–890.
Thompson SR, McCaskie PA, Beilby JP, Hung J, Jennens M, Chapman C et al. IL18 haplotypes are associated with serum IL-18 concentrations in a population-based study and a cohort of individuals with premature coronary heart disease. Clin Chem 2007; 53: 2078–2085.
Niu ZL, Zhang PA, Tong YQ . Association of plasma interleukin-18 levels and polymorphisms in interleukin-18 gene with outcomes of hepatitis C virus infections: a meta-analysis. J Immunoassay Immunochem 2015; 36: 221–232.
Bouzgarrou N, Hassen E, Schvoerer E, Stoll-Keller F, Bahri O, Gabbouj S et al. Association of interleukin-18 polymorphisms and plasma level with the outcome of chronic HCV infection. J Med Virol 2008; 80: 607–614.
Frayling TM, Rafiq S, Murray A, Hurst AJ, Weedon MN, Henley W et al. An interleukin-18 polymorphism is associated with reduced serum concentrations and better physical functioning in older people. J Gerontol A Biol Sci Med Sci 2007; 62: 73–78.
Tiret L, Godefroy T, Lubos E, Nicaud V, Tregouet DA, Barbaux S et al. Genetic analysis of the interleukin-18 system highlights the role of the interleukin-18 gene in cardiovascular disease. Circulation 2005; 112: 643–650.
Xia K, Shabalin AA, Huang S, Madar V, Zhou YH, Wang W et al. seeQTL: a searchable database for human eQTLs. Bioinformatics 2012; 28: 451–452.
Carithers LJ, Ardlie K, Barcus M, Branton PA, Britton A, Buia SA et al. A novel approach to high-quality postmortem tissue procurement: the GTEx Project. Biopreserv Biobank 2015; 13: 311–319.
Zeller T, Wild P, Szymczak S, Rotival M, Schillert A, Castagne R et al. Genetics and beyond–the transcriptome of human monocytes and disease susceptibility. PLoS One 2010; 5: e10693.
Wawrocki S, Druszczynska M, Kowalewicz-Kulbat M, Rudnicka W . Interleukin 18 (IL-18) as a target for immune intervention. Acta Biochim Pol 2016; 63: 59–63.
Puren AJ, Fantuzzi G, Dinarello CA . Gene expression, synthesis, and secretion of interleukin 18 and interleukin 1beta are differentially regulated in human blood mononuclear cells and mouse spleen cells. Proc Natl Acad Sci USA 1999; 96: 2256–2261.
Tone M, Thompson SA, Tone Y, Fairchild PJ, Waldmann H . Regulation of IL-18 (IFN-gamma-inducing factor) gene expression. J Immunol 1997; 159: 6156–6163.
Lamkanfi M, Dixit VM . Inflammasomes and their roles in health and disease. Annu Rev Cell Dev Biol 2012; 28: 137–161.
Franchi L, Eigenbrod T, Munoz-Planillo R, Nunez G . The inflammasome: a caspase-1-activation platform that regulates immune responses and disease pathogenesis. Nat Immunol 2009; 10: 241–247.
Gu Y, Kuida K, Tsutsui H, Ku G, Hsiao K, Fleming MA et al. Activation of interferon-gamma inducing factor mediated by interleukin-1beta converting enzyme. Science 1997; 275: 206–209.
Vlahov D, Munoz A, Anthony JC, Cohn S, Celentano DD, Nelson KE . Association of drug injection patterns with antibody to human immunodeficiency virus type 1 among intravenous drug users in Baltimore, Maryland. Am J Epidemiol 1990; 132: 847–856.
Cox AL, Netski DM, Mosbruger T, Sherman SG, Strathdee S, Ompad D et al. Prospective evaluation of community-acquired acute-phase hepatitis C virus infection. Clin Infect Dis 2005; 40: 951–958.
Kim AY, Kuntzen T, Timm J, Nolan BE, Baca MA, Reyor LL et al. Spontaneous control of HCV is associated with expression of HLA-B 57 and preservation of targeted epitopes. Gastroenterology 2011; 140: 686–696.e1.
Tobler LH, Bahrami SH, Kaidarova Z, Pitina L, Winkelman VK, Vanderpool SK et al. A case-control study of factors associated with resolution of hepatitis C viremia in former blood donors (CME). Transfusion 2010; 50: 1513–1523.
Kuniholm MH, Gao X, Xue X, Kovacs A, Marti D, Thio CL et al. The relation of HLA genotype to hepatitis C viral load and markers of liver fibrosis in HIV-infected and HIV-uninfected women. J Infect Dis 2011; 203: 1807–1814.
Duggal P, Thio CL, Wojcik GL, Goedert JJ, Mangia A, Latanich R et al. Genome-wide association study of spontaneous resolution of hepatitis C virus infection: data from multiple cohorts. Ann Intern Med 2013; 158: 235–245.
Patterson N, Price AL, Reich D . Population structure and eigen analysis. PLoS Genet 2006; 2: e190.
Howie BN, Donnelly P, Marchini J . A flexible and accurate genotype imputation method for the next generation of genome-wide association studies. PLoS Genet 2009; 5: e1000529.
1000 Genomes Project Consortium 1000 Genomes Project Consortium Auton A, 1000 Genomes Project Consortium Brooks LD, 1000 Genomes Project Consortium Durbin RM, 1000 Genomes Project Consortium Garrison EP, 1000 Genomes Project Consortium Kang HM et al. A global reference for human genetic variation. Nature 2015; 526: 68–74.
Roshyara NR, Scholz M . fcGENE: a versatile tool for processing and transforming SNP datasets. PLoS One 2014; 9: e97589.
Normand SL . Meta-analysis: formulating, evaluating, combining, and reporting. Stat Med 1999; 18: 321–359.
Liu JZ, Tozzi F, Waterworth DM, Pillai SG, Muglia P, Middleton L et al. Meta-analysis and imputation refines the association of 15q25 with smoking quantity. Nat Genet 2010; 42: 436–440.
Acknowledgements
We acknowledge support from the National Institutes of Health (NIH), National Institute on Drug Abuse R01DA013324 (Thomas), R01DA12568 (Mehta) and U01DA036297 (Kirk); National Institute of Allergy and Infectious Diseases (NIAID) R01 AI108403 (Cox); NIH K23 AI124913 (Lahiri); NIH U19AI066345, U01AI131314, R01DA033541 and U19AI082630 (Kim). Data in this manuscript were partially collected by the Women’s Interagency HIV Study (WIHS). WIHS (Principal Investigators): UAB-MS WIHS (Saag, Kempf and Konkle-Parker), U01-AI-103401; Atlanta WIHS (Ofotokun and Wingood), U01-AI-103408; Bronx WIHS (Anastos), U01-AI-035004; Brooklyn WIHS (Minkoff and Gustafson), U01-AI-031834; Chicago WIHS (Cohen and French), U01-AI-034993; Metropolitan Washington WIHS (Kassaye), U01-AI-034994; Miami WIHS (Fischl and Metsch), U01-AI-103397; UNC WIHS (Adimora), U01-AI-103390; Connie Wofsy Women’s HIV Study, Northern California (Greenblatt, Aouizerat and Tien), U01-AI-034989; WIHS Data Management and Analysis Center (Gange and Golub), U01-AI-042590; Southern California WIHS (Milam), U01-HD-032632 (WIHS I–WIHS IV). The WIHS is funded primarily by the National Institute of Allergy and Infectious Diseases (NIAID), with additional co-funding from the Eunice Kennedy Shriver National Institute of Child Health and Human Development (NICHD), the National Cancer Institute (NCI), the National Institute on Drug Abuse (NIDA) and the National Institute on Mental Health (NIMH). Targeted supplemental funding for specific projects is also provided by the National Institute of Dental and Craniofacial Research (NIDCR), the National Institute on Alcohol Abuse and Alcoholism (NIAAA), the National Institute on Deafness and other Communication Disorders (NIDCD) and the NIH Office of Research on Women’s Health. WIHS data collection is also supported by UL1-TR000004 (UCSF CTSA) and UL1-TR000454 (Atlanta CTSA).
Disclaimer
The contents of this publication are solely the responsibility of the authors and do not represent the official views of the NIH.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on Genes and Immunity website
Rights and permissions
About this article
Cite this article
Vergara, C., Thio, C., Latanich, R. et al. Genetic basis for variation in plasma IL-18 levels in persons with chronic hepatitis C virus and human immunodeficiency virus-1 infections. Genes Immun 18, 82–87 (2017). https://doi.org/10.1038/gene.2017.2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/gene.2017.2