Abstract
Apoptotic cell death is mediated by caspase activation. Autophagy involves the sequestration of cytoplasmic contents into autophagosomes for traffic to lysosomes for degradation. Although autophagy is antiapoptotic, increased numbers of autophagosomes have been associated with forms of non-apoptotic cell death. Apoptosis and autophagy may be co-regulated in the same directions, as the antiapoptotic Bcl-2 and Bcl-xL proteins negatively regulate autophagy by binding to Beclin 1 (mammalian Atg6), and proapoptotic BH3-only proteins may reverse this effect by displacing these interactions. Here, we show that apoptosis can suppress autophagy. Apoptosis induced by the proapoptotic protein Bax reduced autophagy by enhancing caspase-mediated cleavage of Beclin 1 at D149. After cleavage, both N- and C-terminal Beclin 1 fragments change their localisations and these fragments do not interact normally with Vps34, which is required for autophagy. The cleavage of Beclin 1 is a critical event whereby caspases inhibit autophagy, as a non-cleavable Beclin 1 mutant restored autophagy in cells overexpressing Bax.
Similar content being viewed by others
Main
Macroautophagy, which we will call autophagy, is a bulk degradation system that mediates clearance of long-lived cytoplasmic proteins, including aggregate-prone proteins, certain pathogens (like mycobacteria and herpes viruses) and organelles (like mitochondria).1, 2, 3, 4 In mammalian cells, autophagosome formation begins with a nucleation step, where membranes of unknown origins form phagophores, which then expand and fuse to form completed double-membrane vesicles called autophagosomes. Autophagosomes are formed at random sites in the cytoplasm, then traffic along microtubules in a dynein-dependent fashion towards the microtubule-organising centre where they are more likely to encounter lysosomes.5 After fusion with the lysosomes, the contents of the autophagosomes are degraded.6
Autophagy is controlled by two ubiquitin-like conjugation processes. The first involves conjugation of Atg12 to Atg5. Atg5–Atg12 conjugates are localised to phagophores and dissociate after mature autophagosomes are formed.7 Atg5–Atg12 conjugation is regulated by the activity of the class III PI-3-kinase, Vps34.8 Vps34 activity is positively regulated by the mammalian orthologue of yeast Atg6, called Beclin 1.9 The second modification involves conjugation of microtubule-associated protein 1 light chain 3 (MAP–LC3/Atg8/LC3) to phosphatidylethanolamine (PE). LC3 (cytosolic) is cleaved at its C terminus by Atg4 to form LC3-I. LC3-I is covalently conjugated to PE to form LC3-II. LC3-II (membrane associated) is specifically targeted to Atg5–Atg12-associated, expanded phagophores and remains associated with autophagosomes even after fusion with lysosomes, after which LC3-II can be delipidated and recycled. LC3 is the only known protein that specifically associates with autophagosomes and not with other vesicular structures. Thus, LC3-II levels correlate with autophagosome number, which can also be assessed by scoring LC3-positive vesicles.10
Apoptosis, also called type 1 cell death, is a form of cell suicide mediated by caspase activation.11 Although autophagy has been demonstrated as a cytoprotective and antiapoptotic process, increased numbers of autophagosomes have been associated with forms of non-apoptotic cell death, called type 2 cell death.11, 12, 13, 14 Both apoptosis and autophagy can be regulated by Bcl-2 family proteins. Bcl-2 family proteins have been divided into subgroups based on the presence of their Bcl-2 homology (BH) domain(s).15 The related antiapoptotic proteins Bcl-2 and Bcl-xL contain four BH domains (BH1–BH4). Bad, Noxa, Bim, Bid, Puma, and Bnip3/Nix each contain a single BH3 domain, and are a group of proapoptotic proteins. Bax and Bak contain three BH domains (BH1–BH3), and either protein is required for mitochondrial outer membrane permeabilisation leading to cytochrome c release, which, in turn, results in the activation of a cascade of caspase enzymes that ultimately execute apoptosis.16, 17 BH3-only proteins appear to kill cells by binding to the BH3 receptor, the groove formed by the BH1, BH2 and BH3 domains. This process may either activate Bax/Bak, or may induce death by displacing antiapoptotic BH1-4 (Bcl-2/Bcl-xL) proteins, which inhibit Bax/Bak when bound to them.16, 17, 18
The Bcl-2 family proteins also regulate autophagy, because Beclin 1 has a BH3 domain that binds to a hydrophobic groove in Bcl-2/Bcl-xL.19 The binding of Bcl-2/Bcl-xL to Beclin 1 impairs autophagy, as the complex containing Beclin 1 and Vps34 has less PI-3-kinase activity when associated with Bcl-2/Bcl-xL.20 The inhibitory effect of Bcl-2/Bcl-xL on autophagy can be suppressed if Beclin 1 is dissociated from these proteins by BH3-only proteins like Bad.21 Thus, apoptosis and autophagy may be co-regulated in the same directions. However, BH1-3 proteins, like Bax, have not been implicated in autophagy regulation.
Results
Bax inhibits autophagy
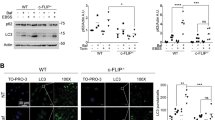
We initially tested whether Bax could disrupt the Beclin 1–Bcl-xL interaction, analogous to what was observed with BH3-only proteins, and confirmed that this was the case (Figure 1a). We then tested if Bax overexpression enhanced autophagy. In contrast to what was seen with Bad overexpression,21 we found that Bax significantly reduced LC3-II levels (Figure 1b), and found that this effect was Bax-dose-dependent (Figure 1c, Supplementary Figure S1a–b). Bax also reduced LC3-II or GFP-LC3-II levels in cells treated with Bafilomycin A1 (Baf), which blocks LC3-II degradation, suggesting that Bax decreased autophagosome synthesis (Figure 1b, Supplementary Figure S1a–b). We also tested whether Bax decreased autophagy in conditions where it was stimulated. Supplementary Figure S2 shows that Bax still attenuated LC3-II synthesis in cells treated with the potent autophagy inducer, trehalose.22 The effects of Bax on autophagy appeared to be caspase-dependent, as they were largely reversed by the pan-caspase inhibitor, z-VAD-fmk. This effect was observed both by LC3-II immunoblotting (Figure 1c) and by scoring GFP–LC3 vesicle numbers (Figure 1d). High concentrations (100 μM) of zVAD-fmk may inhibit cathepsins in cells23 and calpain in vitro.24 The potential inhibitions of cathepsins and calpain are unlikely to have a major impact on our results, because we used a much lower concentration in the experiments (20 μM). We confirmed that the decrease of LC3-II levels caused by Bax was dependent on caspase-3 activity, as a caspase-3 inhibitor reversed this effect (Figure 1e).
Bax reduces autophagosome formation. (a) Bcl-xL (0.6 μg), Bcl-xL (0.6 μg)/Beclin 1 (0.6 μg), Bcl-xL (0.6 μg)/Beclin 1 (1.2 μg)/Bad (0.6 μg) and Bcl-xL (0.6 μg)/Beclin 1 (1.8 μg)/Bax (0.6 μg) were transfected into HeLa cells. To achieve similar expression levels of Bcl-xL and Beclin 1, we used different amounts of Bcl-xL and Beclin 1 in each transfection. Empy vectors were used to keep the amount of DNA in each transfection constant. After 20 h, cells were lysed and anti-Flag was used for immunoprecipitation. Immunoprecipitates were detected with anti-Bcl-xL and anti-Flag (Beclin 1), respectively. Total lysates were detected with anti-Bcl-xL, anti-Flag, anti-Bad and anti-Bax, respectively. (b) HeLa cells were transfected with empty vector (lanes 1, 3), or Bax (lanes 2, 4). After 20 h, one of the two sets (lanes 3–4) was treated with Bafilomycin A1 (Baf) for 4 h. This treatment blocks LC3-II degradation and is saturating; thus, the changes in LC3-II reflect altered autophagosome synthesis.8 The cells were then lysed and subjected to SDS-PAGE and blotting with anti-LC3 (top), anti-tubulin (bottom), respectively. Note that lighter exposures were used in the Baf conditions to enable assessment of changes in LC3-II levels, as Baf dramatically increases LC3-II levels. Endogenous LC3-I levels appear very low in HeLa cells as LC3 antibody has stronger immunoreactivity to LC3-II than LC3 I.33 (c) GFP-LC3 was co-transfected with increasing levels of Bax plasmid, as indicated in duplicate. (In all experiments of this type, the total amount of transfected DNA is kept constant by using empty vector.) After transfection, one set was treated with DMSO (lanes 1–6) and the other set was treated with z-VAD-fmk (20 μM) (lanes 7–12). After 20 h, cell lysates were subjected to western blot with anti-GFP (top and middle) and anti-tubulin (bottom). (d) GFP-LC3/vector, or GFP-LC3/Bax were transfected into HeLa cells. The GFP-LC3/Bax transfections were treated with either DMSO or z-VAD-fmk (20 μM), and GFP–LC3 transfection was also treated with DMSO following transfection. After 20 h, cells were fixed and the numbers of GFP puncta were scored. Pictures were taken under fluorescent microscope. These transfections do not give rise to GFP–LC3 aggregates 5. *P1<0.05; P2<0.01. (e) Vector or Bax were transfected into HeLa cells. The Bax transfections were treated with either DMSO or caspase-3 inhibitor (Ac-DEVD-CHO) (20 μM), and vector transfection was also treated with DMSO following transfection. In a parallel experiment, cells were treated with Bafilomycin A1 for the last 4 h before harvesting. After 20 h, cell lysates were subjected to western blots and probed with anti-LC3, anti-tubulin antibodies. LC3-II densitometry is indicated (versus tubulin)
Caspases cleave and inactivate Beclin 1
We tried to identify the mechanism whereby Bax reduces autophagy, and noted that Bax reduced Beclin 1 expression (Figure 2a). As, pan-caspase inhibition stabilised Beclin 1 in the presence of Bax (Figure 2a, Supplementary Figure S3a–b) and also rescued the Bax-mediated inhibition of autophagy (Figure 1c and d, Supplementary Figure S3b), we considered that Beclin 1 may be a caspase substrate and that this may be the mechanism for these Bax effects.
Beclin 1 is cleaved by caspase-3 at D149. (a) Beclin 1-Flag was co-transfected with increasing levels of Bax plasmid as indicated. After transfection, one set (lanes 1–6) was treated with DMSO and the other set was treated with z-VAD-fmk (lanes 7–12). After 20 h, cell lysates were subjected to western blot with anti-Flag (top), anti-Bax (middle) and anti-tubulin (bottom). Note that Beclin 1 band intensities (versus tubulin) for 0 Bax are 1 (lane 1) and 1.04 (lane 7). (b) Carboxyl terminal Flag-tagged Beclin 1 was transfected into HeLa cells in triplicate. The first set of transfected cells was treated with cycloheximide (CHX) for 5 h (lane 1); the second set of transfected cells was treated with TNF-α+ CHX for 5 h (lane 2); the third set of transfected cells was treated with TNF-α +CHX and z-VAD-fmk for 5 h (lane 3). The cell lysates were subjected to western blot with anti-Flag (10% gel). (c) HeLa cells were treated with DMSO, staurosporine (STS), and STS+z-VAD-fmk for 5 h, respectively. The cell lysates were subjected to blotting with anti-Beclin 1 (C-term) (top) and anti-tubulin (bottom). *A 50 kDa band that may be a cleavage intermediate. (d) Beclin 1, Beclin 1-D142E, Beclin 1-D149E or Beclin 1-D142 149E (C-terminal Flag tagged) were transfected into HeLa cells in duplicate. After 20 h, one set of cells (lanes 5–8) was treated with TNF-α+CHX to induce caspases activation. As controls, another set of cells (lanes 1–4) was treated CHX only (which does not cause effective apoptosis by itself). Samples were subjected to 10% SDS-PAGE and blotting with anti-Flag. (e) For caspase-3 in vitro cleavage assay, Beclin 1, or Beclin 1-149E (C-terminal Flag tagged) were transfected into HeLa cells. After 20 h, cells were lysed. The cell lysates were split into two parts. Caspase-3-containing bacterial lysate (lanes 3, 4) and control (lanes 1, 2) bacterial lysate was used to cleave Beclin 1 or Beclin 1-149E containing cell lysates. Samples were subjected to 12% SDS-PAGE and blotting with anti-Flag. Note that the extent of the Beclin 1 fragment migration differs in Figure 2b, d and e as different percentage gels were used. (f) Alignment of Beclin 1 from different species, showing caspase consensus sequence
The treatment of cells with both TNF-α and cycloheximide (CHX) is an established method for caspase activation.25 Consistent with the Bax overexpression data, TNF+CHX reduced Beclin 1 levels by at least 50% and this effect was inhibited by caspase inhibition (Figure 2b). This is likely to be significant as a 50% reduction of Beclin 1 affects autophagic activity.26 The caspase-dependent reduction in full-length Beclin 1 levels was associated with the appearance of a fragment of this protein that was recognized by a tag on the C terminus of exogenous Beclin 1 (Figure 2b). TNF-α +CHX treatment also caused a reduction in the levels of endogenous Beclin 1. We were unable to detect a Beclin 1 fragment using a Beclin 1 N-terminal antibody, suggesting rapid turnover of this fragment (Supplementary Figure S4a). However, the caspase inhibitor, z-VAD-fmk abolished the ability of TNF+CHX to reduce Beclin 1 levels (Supplementary Figure S4b). Autophagic degradation is unlikely to have a significant role in the Beclin 1 turnover under these conditions, because the potent chemical autophagy inducer trehalose failed to alter Beclin 1 levels significantly in both Atg5 wild-type (autophagy-competent) and Atg5 null (autophagy-deficient) mouse embryonic fibroblasts (MEFs) (Supplementary Figure S5). Consistent with these data, staurosporine, another apoptosis-inducing treatment, also resulted in Beclin 1 cleavage detected with an antibody directed to the C-terminus of endogenous Beclin 1 (Figure 2c). Interestingly, we observed an extra 50 kDa band in this assay, however, this size band was not detected in the assay with Beclin 1-Flag. We cannot exclude the possibility that this 50 kDa band is an intermediate fragment.
To identify the caspase cleavage site, we screened the potential caspase consensus sequences within Beclin 1 and found that Asp149 (149D) was a preferred caspase cleavage site. To verify this caspase cleavage site, we mutated 149D to Glu (149E). Figure 2d shows, that, in contrast to wild-type Beclin 1, Beclin 1-149E is no longer cleaved. As caspase-3 is a major caspase acting in apoptotic execution pathways, we tested if it was able to cleave Beclin 1 in vitro. Figure 2e shows that caspase-3 is capable of cleaving wild-type Beclin 1 but not mutated Beclin 1-149E. These data revealed that Beclin 1 is a substrate of caspase-3 and that 149D is the major cleavage site. This caspase recognition sequence is well conserved in Beclin 1 orthologues in many species (Figure 2f). Notably, this site is not conserved in yeasts that are thought to lack caspases.
We tested if the N-terminal and C-terminal caspase cleavage products of Beclin 1 (Beclin 1 N and C) were still able to induce autophagy. We have established that autophagy regulates the clearance of mutant huntingtin (htt) fragments and other aggregate-prone proteins, and that the percentage of cells with aggregates of mutant htt correlates inversely with autophagic activity, all other things being equal.27, 28 When Beclin 1 was co-transfected with a GFP-tagged exon 1 mutant htt fragment (GFP-74Q) in SK-N-SH human neuroblastoma cells, the aggregation of mutant htt was markedly reduced, compared with vector-cotransfected cells, as previously reported.29 However, neither Beclin 1 N or C by themselves, nor together, had any significant effect on the clearance of mutant htt aggregation, suggesting that both Beclin 1-N and C have lost autophagic activity (Figure 3a). Similarly, full-length Beclin 1 overexpression consistently increased autophagosome numbers (Figure 3b), whereas its N-terminal and C-terminal caspase cleavage products had no significant effects. The effects of Beclin 1 on htt aggregation is autophagy dependent as it is lost in cells treated with the autophagy inhibitor, 3-methyl adenine (3MA) (Figure 3a) and in autophagy-deficient Atg5 null MEFs (Figure 3c). Consistently, Beclin 1, rather than Beclin 1-N or -C, reduced mutant httEx1 aggregation in Atg5 wild-type MEFs (Figure 3c). The expression levels of Myc-Beclin 1, Myc-Bec-N and Myc-Bec-C are shown in Supplementary Figure S6.
Caspases-cleaved Beclin 1 fragments lose autophagic activity. (a) GFP-tagged huntingtin exon 1 with 74 polyglutamines (GFP-74Q) was cotransfected with empty vector, Myc-Beclin 1 (Myc-Bec), Myc-Beclin 1 N-terminal fragment (Myc-Bec-N), Myc-Beclin 1 C-terminal fragment (Myc-Bec-C) and the combination of Myc-Bec-N and Myc-Bec-C (Myc-Bec-N+C) into HeLa cells in two sets. One set of experiments was treated with the autophagy blocker 3-methyl-adenine (3-MA) following transfection. After 24 h, aggregates were scored under a fluorescent microscope. *P<0.05. (b) GFP-LC3 was cotransfected with empty vector, Beclin 1, Beclin 1-N, Beclin 1-C into neuroblastma SK-N-SH cells. After 24 h, GFP puncta were scored under a fluorescent microscope. *P<0.05. (c) GFP-74Q was cotransfected with empty vector, Myc-Beclin 1 (Myc-Bec), Myc-Beclin 1 N-terminal fragment (Myc-Bec-N), Myc-Beclin 1 C-terminal fragment (Myc-Bec-C) and the combination of Myc-Bec-N and Myc-Bec-C (Myc-Bec-N+C) into Atg5−/− MEFs (mouse embryonic fibroblasts). After 48 h, cells were fixed and aggregation was evaluated. GFP-74Q was cotransfected with empty vector, myc-Beclin 1 (myc-Bec), myc-Beclin 1 N-terminal fragment (myc-Bec-N), myc-Beclin 1 C-terminal fragment (myc-Bec-C), or the combination of myc-Bec-N and myc-Bec-C (myc-Bec-N+Bec-C) into Atg5 wild-type MEFs. After 48 h, cells were fixed and aggregation was evaluated. *P<0.05
Non-caspase-cleavable Beclin 1 rescues autophagy in Bax-expressing cells
We tested if Beclin 1 cleavage was the major determinant of the inhibition of autophagy by caspases and confirmed that this was the case, as the Bax reduced autophagosome numbers in vector-transfected cells, but the autophagosome numbers were restored in Beclin 1-149E-transfected cells. The autophagosome numbers were partially restored by cleavable wild-type Beclin 1 (Figure 4a). We confirmed that the expression of Beclin 1-149E in the presence of Bax was similar to those of wild-type Beclin 1 in otherwise normal cells not exposed to Bax overexpression (Figure 4b). We then confirmed that Bax also reduced autophagosome numbers in starvation conditions, which normally increase the numbers of LC3 vesicles per cell (Supplementary Figure S7a). Supplementary Figure S7b shows that Beclin 1-149E exerted a protective effect against the reduction of autophagosomes induced by Bax in starvation conditions, in a way that was analogous to what we observed in full media. Similarly, Beclin 1 D149E restored LC3-II levels in Bax-transfected cells in both Baf or non-Baf-treated cells in full media (Figure 4c).
Non-cleavable Beclin 1 restores Bax-reduced autophagy. (a) GFP-LC3, GFP-LC3/Bax, GFP-LC3/Bax/Beclin 1-149E, GFP-LC3/Bax/Beclin 1 were transfected into HeLa cells. After 20 h, cells were fixed and the numbers of GFP puncta per cell were scored. *P<0.05. (b) HeLa cells were transfected with empty vector, Bax, Bax/Beclin 1-149E and Bax/Beclin 1 respectively. After 20 h, cell lysates were subjected to blotting, and probed with anti-Beclin 1 (N-) and tubulin antibodies. (c) HeLa cells were transfected with vector, Bax, Bax/Beclin 1 and Bax/Beclin 1-149E, respectively, in duplicate. After 20 h, one set was treated with DMSO and the other set was treated Baf for 4 h. Cell lysates were subjected to blotting with anti-LC3 and tubulin, respectively
To exclude the possibility that Beclin 1-149E acts as a caspase-3 inhibitor, we assayed caspase-3 activity in Bax-overexpressing cells in the presence or absence of wild-type on Beclin 1-149E. As expected, caspase-3 activity was enhanced by Bax overexpression. However, this effect was not affected in cells overexpressing either wild-type Beclin 1 and Bax or Beclin 1-149E and Bax, compared with cells overexpressing Bax alone (Supplementary Figure S8a). These data were confirmed by the cleavage assay of poly(ADP-ribose) polymerase (PARP), a classical caspase-3 substrate. Similar levels of the 85-kDa PARP caspase fragment were detected in cells with Bax alone, Bax and Beclin 1 or Bax and Beclin 1-149E (Supplementary Figure S8b).
Previously, we showed that Beclin 1 regulates Atg5–Atg12 conjugation, a step that precedes LC3-II formation.8 Supplementary Figure S9 confirms that the levels of the endogenous Atg5–Atg12 conjugate are decreased in the Beclin 1-knockdown cells. Consistent with these findings, Bax decreased Atg5–Atg12 conjugation in vector-transfected cells, whereas Beclin 1-149E partially restored the levels of the conjugate (Supplementary Figure S10a–b).
Mechanism of the loss-of-function of Beclin 1 cleavage fragments
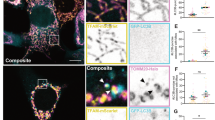
Beclin 1 is a predominantly cytoplasmic protein, enriched in mitochondria and ER.20, 30.These localisations are important for its function(s). We tested if the localisations of caspase-cleaved fragments of Beclin 1 are different from full-length Beclin 1. We generated Beclin 1–GFP (Bec–GFP), N-terminal Beclin 1–GFP (1-149aa, Bec-N-GFP), and C-terminal Beclin 1–GFP (150–450aa, Bec-C-GFP). Bec–GFP was predominantly localised in the cytoplasm, as previously reported. However, Bec-N-GFP was mainly localised in the nucleus, and Bec-C-GFP was localised in both cytoplasm and nucleus (Figure 5(a–b)). Similar effects were observed with these constructs when tagged with Myc, rather than with GFP (Figure 5c). As the interaction between Beclin 1 and Vps34 is essential for autophagy, we tested whether the cleaved Beclin 1 fragments still retained the ability to bind Vps34. Figure 5d and Supplementary Figure S11 show that Beclin 1 N-terminal bound only weakly to Vps34, whereas this interaction is almost lost for Beclin 1 C-terminal. These data, along with the mislocalisation of cleaved Beclin 1, can explain why the cleavage results in a loss-of-function.
Mechanism of the loss-of-function of cleaved Beclin 1 fragments. (a, b) Beclin 1–GFP (Bec–GFP), N-terminal caspase-cleaved fragment of Beclin 1 ( Beclin 1-N)-GFP (Bec-N-GFP), C-terminal caspase-cleaved fragment of Beclin 1 (Beclin 1-C)-GFP (Bec-C-GFP) were transfected into HeLa cells, respectively. After 18 h, cells were fixed. The cells were later viewed under a fluorescent microscope (a) and confocal microscope (b). (c) Myc-Beclin 1 (Myc-Bec), Myc-Beclin 1-N (Myc-Bec-N), Myc-Beclin-C (Myc-Bec-C) were transfected into HeLa cells. After 20 h, cells were fixed and stained with anti-Myc (9E10). The cells were viewed under a confocal microscope. (d) Vps34/vector, Vsps34/Beclin 1-Flag (Bec-Flag), Vps34/Beclin 1-N-Flag (Bec-N-Flag), Vps34/Beclin 1-C-Flag (Bec-C-Flag) were transfected into HeLa cells, respectively. After 20 h, cells were harvested and lysed. The cell lysates were subjected to anti-Flag antibody immunoprecipitation. The immunoprecipitates were used for SDS-PAGE and western blotting with anti-Vps34 antibody (top panel) and anti-Flag antibody (middle panel), respectively. The total lysates were detected with anti-Vps34 (bottom). Three independent experiments were performed with similar results
Bcl-xL rescues autophagy in bax-expressing cells
Then we asked whether antiapoptotic Bcl-2/Bcl-xL antagonizes the ability of Bax to reduce autophagy. Interestingly, Figure 6a shows that Bcl-xL (column 3) dramatically increased autophagosome number in cells where Bax is overexpressed, compared with the cells overexpressing Bax but no Bcl-xL (column 2). Bcl-xL (column 4) decreased autophagosome number in cells without overexpression of Bax, in comparison with the cells without overexpression of Bax and Bcl-xL (column 1). However, we also observed that Bax (column 3) restored the loss of autophagosomes caused by Bcl-xL (column 4). This effect may be because of Bax disrupting the Bcl-xL–Beclin 1 interaction (Figure 1a), similar to the effect of BH3-only proteins on autophagy.31 The pictures of autophagosomes in these cells are shown in Figure 6b. Figure 6c shows LC3-II levels in the cells where vector, Bax, Bax–Bcl-xL, or Bcl-xL overexpressed. The LC3-II levels are consistent with autophagosome numbers in the cells overexpressing Bax, Bax–Bcl-xL or Bcl-xL (Figure 6a, b). Finally, we observed that Bcl-xL rescued both the loss of Beclin 1 and the increased cell death caused by Bax overexpression (Figure 6d, Supplementary Figure S12).
Bcl-xL rescues the inhibitory effects of Bax on autophagosome formation. (a) GFP–LC3 was transfected with empty vector, Bax, Bax–Bcl-xL or Bcl-xL into HeLa cells. After 20 h, cells were fixed and the numbers of GFP puncta per cell were scored. *P<0.05. (b) Pictures were taken under a fluorescent microscope ( × 100). Bar: 5 μm. (c) HeLa cells were transfected with empty vector, Bax, Bax–Bcl-xL or Bcl-xL in duplicate. After 20 h, one set of cells was treated with Baf (lanes 5–8). Cells were harvested and blots were probed with anti-LC3, Bcl-xL, Bax and tubulin, respectively. Note that endogenous LC3-I gives a very weak signal in HeLa cells compared with LC3-II. (d) HeLa cells were transfected with empty vector, Bax, Bax–Bcl-xL or Bcl-xL. After 20 h, cells were harvested and blots were probed with anti-Beclin 1 and tubulin, respectively
Discussion
We find that autophagosome synthesis is inhibited after caspase activation, and this appears to be predominantly mediated by caspase cleavage of Beclin 1. Our finding that Beclin 1 is a caspase substrate is consistent with a recent study by Cho et al.32 that made the same initial observation,32 but did not identify the cleavage site. However, the main conclusion of our study is that caspases/apoptosis inhibit autophagy. This is novel and has not been shown in the Cho paper, which reported that Atg6/Beclin 1 modulates sensitivity of cells to TRAIL-induced cell death, an issue distinct from our study. Furthermore, Cho et al.32 did not examine if the caspase cleavage of Beclin 1 was critical for autophagy inhibition (as caspases could potentially cleave many other autophagy modulators). Although caspase-cleavage of Beclin 1 appears to be a major determinant of the effects we have observed (because they are abrogated by non-caspase cleavable Beclin 1 mutants and pan-caspase or caspase 3 inhibitors), we cannot exclude additional factors influencing Beclin 1 expression under these conditions.
We believe that our data provide new and important insights into the relationship between apoptosis and autophagy. As autophagic flux will be downregulated in cells after caspase activation, the antiapoptotic effects of autophagy will not compete with apoptosis once the latter process is established, and may explain why autophagic and apoptotic morphologies generally do not coexist.
We assayed autophagosome numbers both by western blotting of LC3-II and by counting LC3 dots in cells. The reductions of either parameter induced by Bax were similar (about 50% – see Figure 1e and 6a), although perhaps the effects on LC3 dots were slightly more obvious. One possible explanation for the slightly larger magnitude of effect on Bax on GFP–LC3 dots than on LC3-II expression as measured by western blot analysis is that Atg6 and Beclin 1 may not be directly required for Atg8/LC3 lipidation as an isolated process, although they are required for the localisation of lipidated LC3 to autophagosomal membranes. Therefore, the assays used in this study may be underrepresenting the inhibitory effects of Bax overexpression on autophagy.
Our data also reveal another role for Bcl-2 family members in autophagy regulation. In contrast with BH3-only proteins that can act to enhance autophagy by relieving the inhibition of the inhibitory effects of the Bcl-2–Bcl-xL interaction with Beclin 1, Bax/caspase induction results in Beclin 1 cleavage and decreased autophagy (Supplementary Figure S13). This model is also supported by the observation that when autophagosome numbers are decreased by either Bax (by cleaving Beclin 1), or Bcl-xL (by binding to Beclin 120), co-transfection of Bcl-xL with Bax normalizes autophagosome numbers (Figure 6, Supplementary Figure S13). This may occur because Bcl-xL antagonizes the proapoptotic effects of Bax, whereas Bax, in turn, disrupts the Beclin 1–Bax interaction (see Figure 1a). Thus, there are two scenarios that appear to be operating with Bax overexpression, depending on the antiapoptotic state of the cells (Figure 1a versus Figure 6). In cells without increased antiapoptotic capacity (e.g., without Bcl-xL overexpression or caspase inhibitor treatment), Bax overexpression will lead to decreased Beclin 1 levels by caspase cleavage and this will decrease autophagic flux. However, in cells with sufficient antiapoptotic capacity (e.g., with Bcl-xL overexpression or caspase inhibitor treatment) then Bax will disrupt the Bcl-xl–Beclin interaction and induce autophagy. Another important factor that will regulate the differential effects of Bcl-2 family members is subcellular localisation. For instance, ER-localised, but not mitochondrial Bcl-2 appears to be relevant for autophagy inhibition.20 Conversely, mitochondrial Bcl-2 is relevant for its antiapoptotic effects. As these protective roles of Bcl-2 would reduce caspase activity after proapoptotic insults, they would also be predicted to preserve autophagy in cells exposed to various toxic agents. Thus, in certain circumstances, Bcl-2/Bcl-xL may have opposite effects on autophagy, depending on whether it is associated with mitochondria or the ER.
Materials and Methods
DNA construction
Beclin 1 or its truncated forms were generated by PCR and subcloned into pCMV-5a vector (C-terminal Flag-tagged expression vector), pCMV-6 M (N-terminal Myc-tagged expression vector), pEGFP-N3 (C-terminal GFP-tagged expression vector). Beclin 1 point mutants were generated with Stratagene Quikchange Mutagenesis kit (Stratagene, Cheshire, UK). Mouse Bad was purchased from Cell Signaling (Hitchin, UK) and subcloned into pcDNA3. Bax and Bcl-xL were the products from Addgene (Cambridge, MA, USA) and cloned into pcDNA3. Vps34 was cloned into pcDNA3. All the DNA constructs were confirmed by DNA sequencing.
Antibodies and reagents
Rabbit polyclonal antibodies: anti-Beclin 1 (C-) (1 : 1000) (Sigma, Gillingham, UK); anti-Beclin 1 (N-) (1 : 1000) (Sigma); anti-Myc (1 : 1000) (Sigma); anti-PARP p85 (1 : 1000) (Promega, Southampton, UK). Anti-mouse monoclonal antibodies: anti-Flag (M2) (1 : 1000), anti-tubulin (1 : 5000) (Sigma), anti-Myc (9E10) (1 : 1000) (Sigma). Anti-Flag M2-agarose affinity gel Sigma). TNF-α, recombinant caspase-3 were purchased from Chemicon International (Dundee, UK). Cycloheximide was from Sigma. Bafilomycin A1 (Baf) was purchased from Millipore (Dundee, UK) (400 nM). Caspase-3 inhibitor (Ac-DEVD-CHO) and Pan-caspase inhibitor (z-VAD-fmk) were from Calbiochem (Nottingham, UK). Cells were treated with 20 μM Ac-DEVD-CHO and 20 μM z-VAD-fmk. Both 3-methyladenine (3-MA) and trehalose were from Sigma.
Cell culture
HeLa, SK-N-SH cells and Atg5 knockout MEFs were cultured with standard methods in DMEM supplemented with 10% FCS (Sigma). Transfection was performed with Lipofectamine 2000 (Invitrogen, Paisley, UK) according to standard methods. In cellular Beclin 1 cleavage assays, cells were treated with 10 ng/ml TNF-α and 30 μg/ml CHX or Staurosporine (STS, 1 μM) for indicated times. To inhibit autophagy, cells were treated with 10 mM 3-MA for 24 h following transfection. To stimulate autophagy, we treated cells with 100 mM trehalose for the indicated times.
siRNA transfection
HeLa cells were split 1 day before transfection to 50% confluence and left overnight in antibiotic-free DMEM containing 10% FBS. Beclin 1 siRNAs (Ambion, Warrington, UK) were transfected with DharmaFect 1 according to the manufacturer's instructions. Non-targeting siRNA (Dharmacon, Lafayette, CO, USA, cat no: D-001210-01) was the control siRNA. HeLa cells were maintained in 10% FBS DMEM containing no antibiotics for 48 h after transfection.
In vitro caspase cleavage
Cells were lysed in lysis buffer (50 mM HEPES, 5 mM DTT, 0.1 mM. EDTA, 0.1% CHAPS, pH 7.4, supplied by Calbiochem in Caspase-3 activity assay kit). Beclin 1 in the lysates was then cleaved with 1 U caspase-3 in cleavage buffer (50 mM HEPES, 50 mM NaCl, 0.1% CHAPS, 10 mM DTT, 1 mM EDTA, 5% Glycerol) for 1 h at 37°C. The cleaved products were subjected to western blot analysis.
Caspase-3 activity assay
Caspase-3 activity assay was performed according to the instructions in the Caspase-3 cellular activity assay kit (Calbiochem, cat no: 235419). Briefly, caspase substrate Ac-DEVD-pNA in assay buffer was added to the same amount of cellular lysates (15 μl) as for in vitro caspase cleavage and the mixtures were incubated for 1 h at 37°C. Absorbance was read at 405 nm.
Estimation of aggregates
To measure aggregates, approximately 200 transfected cells were counted in multiple random visual fields per slide. All coverslips were scored with the observer blinded to the identity of the slides. Cells were analysed using a fluorescent microscope (Eclipse E600, Nikon, Japan). The figures show data from representative experiments in triplicate. Cells were counted as aggregate-positive if one or several aggregates were visible within a cell.
Estimation of numbers of autophagosome per cell
GFP-LC3 puncta were scored under a fluorescent microscope ( × 100). Approximately 50 GFP-positive cells were counted in multiple random visual fields per slide. All coverslips were scored with the observer blinded to the identity of the slides. The figures show data from experiments in triplicate, at least.
Quantification of autoradiographs
To quantify western blot band densitometry, the relevant specified bands were analysed using ChemiImager (Alpha Innotech Co., San Leandro, CA, USA).
Statistics
T-test was used and P-values were determined by unconditional logistical regression analysis by using the general loglinear option of SPSS 9.1 software (SPSS, Chicago, IL, USA).
Immunoprecipitation
Immunoprecipitation (IP) was performed using Buffer A (20 mM Tris-HCl, pH 7.2, 2 mM MgCl2, 150 mM NaCl, 5 mM NaF, 1 mM Na3VO4, 0.5% NP-40, protease inhibitor cocktail (Roche)). Cells were lysed in Buffer A for 20 min on ice, followed by centrifugation at 13 000 × g for 15 min. Then, 500 μg–1 mg total protein were used as the starting material for IPs. Anti-Flag M2-agarose affinity gel was added to a final concentration of 5 μg/ml and incubated for 2 h to overnight at 4°C. IP products were either directly boiled in Laemmli buffer and subjected to PVDF membrane transfer and western blot.
Abbreviations
- Baf:
-
Bafilomycin A1
- BH:
-
Bcl-2 homology
- CHX:
-
cycholoheximide
- GFP-74Q:
-
GFP-tagged exon 1 mutant htt fragment
- htt:
-
huntingtin
- MEFs:
-
mouse embryonic fibroblasts
- 3MA:
-
3-methyl adenine
- PE:
-
phosphatidylethanolamine
- STS:
-
staurosporin
- TNF-α:
-
tumour necrosis factor alpha
References
Klionsky DJ . The molecular machinery of autophagy: unanswered questions. J Cell Sci 2005; 118: 7–18.
Levine B, Klionsky DJ . Development by self-digestion: molecular mechanisms and biological functions of autophagy. Dev Cell 2004; 6: 463–477.
Mizushima N . The pleiotropic role of autophagy: from protein metabolism to bactericide. Cell Death Differ 2005; 12 (Suppl 2): 1535–1541.
Rubinsztein DC, Gestwicki JE, Murphy LO, Klionsky DJ . Potential therapeutic applications of autophagy. Nat Rev Drug Discov 2007; 6: 304–312.
Jahreiss L, Menzies FM, Rubinsztein DC . The itinerary of autophagosomes: from peripheral formation to kiss-and-run fusion with lysosomes. Traffic 2008; 9: 574–587.
Huang WP, Klionsky DJ . Autophagy in yeast: a review of the molecular machinery. Cell Struct Funct 2002; 27: 409–420.
Mizushima N, Noda T, Yoshimori T, Tanaka Y, Ishii T, George MD et al. A protein conjugation system essential for autophagy. Nature 1998; 395: 395–398.
Ravikumar B, Imarisio S, Sarkar S, O’Kane CJ, Rubinsztein DC . Rab5 modulates aggregation and toxicity of mutant huntingtin through macroautophagy in cell and fly models of Huntington disease. J Cell Sci 2008; 121: 1649–1660.
Liang XH, Jackson S, Seaman M, Brown K, Kempkes B, Hibshoosh H et al. Induction of autophagy and inhibition of tumorigenesis by beclin 1. Nature 1999; 402: 672–676.
Kabeya Y, Mizushima N, Ueno T, Yamamoto A, Kirisako T, Noda T et al. LC3, a mammalian homologue of yeast Apg8p, is localized in autophagosome membranes after processing. EMBO J 2000; 19: 5720–5728.
Levine B, Yuan J . Autophagy in cell death: an innocent convict? J Clin Invest 2005; 115: 2679–2688.
Lum JJ, DeBerardinis RJ, Thompson CB . Autophagy in metazoans: cell survival in the land of plenty. Nat Rev Mol Cell Biol 2005; 6: 439–448.
Mizushima N, Levine B, Cuervo AM, Klionsky DJ . Autophagy fights disease through cellular self-digestion. Nature 2008; 451: 1069–1075.
Yorimitsu T, Klionsky DJ . Eating the endoplasmic reticulum: quality control by autophagy. Trends Cell Biol 2007; 17: 279–285.
Danial NN, Korsmeyer SJ . Cell death: critical control points. Cell 2004; 116: 205–219.
Wei MC, Zong WX, Cheng EH, Lindsten T, Panoutsakopoulou V, Ross AJ et al. Proapoptotic BAX and BAK: a requisite gateway to mitochondrial dysfunction and death. Science 2001; 292: 727–730.
Willis SN, Fletcher JI, Kaufmann T, van Delft MF, Chen L, Czabotar PE et al. Apoptosis initiated when BH3 ligands engage multiple Bcl-2 homologs, not Bax or Bak. Science 2007; 315: 856–859.
Certo M, Del Gaizo Moore V, Nishino M, Wei G, Korsmeyer S, Armstrong SA et al. Mitochondria primed by death signals determine cellular addiction to antiapoptotic BCL-2 family members. Cancer Cell 2006; 9: 351–365.
Oberstein A, Jeffrey PD, Shi Y . Crystal structure of the Bcl-XL-Beclin 1 peptide complex: Beclin 1 is a novel BH3-only protein. J Biol Chem 2007; 282: 13123–13132.
Pattingre S, Tassa A, Qu X, Garuti R, Liang XH, Mizushima N et al. Bcl-2 antiapoptotic proteins inhibit Beclin 1-dependent autophagy. Cell 2005; 122: 927–939.
Maiuri MC, Le Toumelin G, Criollo A, Rain JC, Gautier F, Juin P et al. Functional and physical interaction between Bcl-X(L) and a BH3-like domain in Beclin-1. EMBO J 2007; 26: 2527–2539.
Sarkar S, Davies JE, Huang Z, Tunnacliffe A, Rubinsztein DC . Trehalose, a novel mTOR-independent autophagy enhancer, accelerates the clearance of mutant huntingtin and alpha-synuclein. J Biol Chem 2007; 282: 5641–5652.
Rozman-Pungercar J, Kopitar-Jerala N, Bogyo M, Turk D, Vasiljeva O, Stefe I et al. Inhibition of papain-like cysteine proteases and legumain by caspase-specific inhibitors: when reaction mechanism is more important than specificity. Cell Death Differ 2003; 10: 881–888.
Waterhouse NJ, Finucane DM, Green DR, Elce JS, Kumar S, Alnemri ES et al. Calpain activation is upstream of caspases in radiation-induced apoptosis. Cell Death Differ 1998; 5: 1051–1061.
Hsu H, Huang J, Shu HB, Baichwal V, Goeddel DV . TNF-dependent recruitment of the protein kinase RIP to the TNF receptor-1 signaling complex. Immunity 1996; 4: 387–396.
Qu X, Yu J, Bhagat G, Furuya N, Hibshoosh H, Troxel A et al. Promotion of tumorigenesis by heterozygous disruption of the beclin 1 autophagy gene. J Clin Invest 2003; 112: 1809–1820.
Ravikumar B, Duden R, Rubinsztein DC . Aggregate-prone proteins with polyglutamine and polyalanine expansions are degraded by autophagy. Hum Mol Genet 2002; 11: 1107–1117.
Ravikumar B, Vacher C, Berger Z, Davies JE, Luo S, Oroz LG et al. Inhibition of mTOR induces autophagy and reduces toxicity of polyglutamine expansions in fly and mouse models of Huntington disease. Nat Genet 2004; 36: 585–595.
Shibata M, Lu T, Furuya T, Degterev A, Mizushima N, Yoshimori T et al. Regulation of intracellular accumulation of mutant Huntingtin by Beclin 1. J Biol Chem 2006; 281: 14474–14485.
Liang XH, Yu J, Brown K, Levine B . Beclin 1 contains a leucine-rich nuclear export signal that is required for its autophagy and tumor suppressor function. Cancer Res 2001; 61: 3443–3449.
Maiuri MC, Criollo A, Tasdemir E, Vicencio JM, Tajeddine N, Hickman JA et al. BH3-only proteins and BH3 mimetics induce autophagy by competitively disrupting the interaction between Beclin 1 and Bcl-2/Bcl-X(L). Autophagy 2007; 3: 374–376.
Cho DH, Jo YK, Hwang JJ, Lee YM, Roh SA, Kim JC . Caspase-mediated cleavage of ATG6/Beclin-1 links apoptosis to autophagy in HeLa cells. Cancer Lett 2008; 274: 95–100.
Mizushima N, Yoshimori T . How to interpret LC3 immunoblotting. Autophagy 2007; 3: 542–545.
Acknowledgements
We are grateful to Brinda Ravikumar for helpful comments and to the MRC (programme grant) and the Wellcome Trust (Senior Clinical Fellowship) for funding.
Author information
Authors and Affiliations
Corresponding author
Additional information
Edited by D Klionsky
Supplementary Information accompanies the paper on Cell Death and Differentiation website (http://www.nature.com/cdd)
Supplementary information
Rights and permissions
About this article
Cite this article
Luo, S., Rubinsztein, D. Apoptosis blocks Beclin 1-dependent autophagosome synthesis: an effect rescued by Bcl-xL. Cell Death Differ 17, 268–277 (2010). https://doi.org/10.1038/cdd.2009.121
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cdd.2009.121
Keywords
This article is cited by
-
ERRα protects against sepsis-induced acute lung injury in rats
Molecular Medicine (2023)
-
Targeted Inhibition of Hsp90 in Combination with Metformin Modulates Programmed Cell Death Pathways in A549 Lung Cancer Cells
Applied Biochemistry and Biotechnology (2023)
-
Identification and validation of key biomarkers and potential therapeutic targets for primary open-angle glaucoma
Science China Life Sciences (2023)
-
Paradoxical downregulation of LPAR3 exerts tumor-promoting activity through autophagy induction in Ras-transformed cells
BMC Cancer (2022)
-
Cytochrome B5 type A alleviates HCC metastasis via regulating STOML2 related autophagy and promoting sensitivity to ruxolitinib
Cell Death & Disease (2022)