Abstract
Population genetic structure of the circum-Mediterranean caddisfly Mesophylax aspersus (Trichoptera, Limnephilidae) on the Canary Islands was investigated by studying allozyme variation at nine putative loci in five populations. Genetic variability, population structure and gene flow were compared with data in the literature for continental taxa to assess the effect of isolation of island populations on the genetic structure. Larvae were collected from streams on the islands of Tenerife (one population), La Gomera (two populations in the same catchment) and La Palma (two populations in different catchments). Genetic variability within populations was high relative to that recorded previously for continental Trichoptera, e.g. mean heterozygosity was 0.119–0.336 (0.035–0.15 in continental taxa). Highly significant population structuring was observed (mean FST=0.250), and there was significant within-population structuring (mean F IS = 0.098). Populations from the same catchment or island were no more similar than populations from different islands, which suggests that occasional long-distance dispersal, both between and within islands, is the predominant influence on the population structure. This dispersal ability has contributed to the colonization of most permanent streams on the Canary Islands by M. aspersus.
Similar content being viewed by others
Introduction
Drainage networks can be viewed as ‘habitat islands’ surrounded by a ‘sea’ of land inhospitable to freshwater invertebrates. Colonization of streams on oceanic islands is more problematic because of the dispersal barrier of the sea and, often, the scarcity of streams, resulting in aquatic taxa often being poorly represented on isolated islands (Wallace, 1880). The community present is strongly influenced by the dispersal abilities of the species in the archipelago species pool, their niche requirements and stochastic colonization processes (Bunn & Hughes, 1997; Belyea & Lancaster, 1999). The archipelago species pool, in turn, is influenced by the chance dispersal of suitable species from a continental source pool (MacArthur & Wilson, 1967). The isolation and age of the Canary Islands, situated off the coast of the western Sahara, have resulted in a high degree of endemism in their flora and fauna. This is due to both the presence of taxa of Tertiary origin, which have become extinct elsewhere in their range, and to post-colonization speciation (Juan et al., 2000).
Freshwater insects possess a wide variety of active and passive dispersal mechanisms. In-stream dispersal by active or passive drift, crawling and swimming typically takes place at the reach scale but, over longer time scales, may allow colonization of a whole stream system. Most freshwater insects can also disperse over land as actively flying adults, allowing colonization of other stream systems (Sheldon, 1984). Long-distance dispersal of winged adults can additionally occur by passive drift in air currents (e.g. Clarke, 1903; Ashmole & Ashmole, 1988). The freshwater taxa occurring on the Canary Islands exhibit a range of dispersal abilities, mechanisms and distributions from extremely localized to ubiquitous (Malmqvist et al., 1995). Widespread species may have greater dispersal ability than species with more restricted distributions, as patch occupancy is often related to dispersal ability (Plague & McArthur, 1998). The relative lack of single-island endemic species within the Canarian freshwater fauna, compared to terrestrial invertebrates, is an indication that inter-island dispersal is substantial in most freshwater taxa. In the Coleoptera, for example, 4% of Dytiscidae are single-island endemics, compared to 54% of Carabidae (Machado, 1992; Alarie & Bilton, in press).
The dispersal ability of individual taxa determines the geographical scale of recruitment and, in combination with historical factors, the scale of population genetic differentiation (Slatkin, 1985). Conversely, the degree of population differentiation observed at a particular scale can be used to infer the amount of dispersal (Bohonak, 1999). Interpopulation dispersal reduces the genetic differentiation of populations that would otherwise occur through founder events, genetic drift and natural selection (Wright, 1943).
The island-like nature of stream habitats can potentially lead to genetic structuring of populations, which is likely to be enhanced by the distribution of the species across real islands. Several studies have used population genetic structure estimates to infer dispersal patterns from allozyme variation in stream invertebrates. Some workers have found no evidence for isolation-by-distance and conclude that stochastic processes such as founder events and fluctuating population sizes are sufficient to explain the population genetic structure (e.g. Jackson & Resh, 1992; Bunn & Hughes, 1997). Others have demonstrated isolation-by-distance, suggesting an additional influence of ongoing distance-dependent dispersal (e.g. Varvio-Aho & Pamilo, 1979; Dillon & Wethington, 1995; Hughes et al., 1996).
In the present study we made a survey of allozyme variation in Mesophylax aspersus Rambur 1842 (Trichoptera, Limnephilidae) from five populations on three islands in the Canary archipelago. We tested three hypotheses about genetic variability, population structure and gene flow in M. aspersus. We hypothesized that genetic variability would be lower than in continental Trichoptera, as island populations are likely to have undergone more marked founder events, and as the sea may be a significant barrier to long-distance gene flow (Pashley et al., 1985). The second hypothesis predicted that genetic structure would be significant, because of the patchy nature of the stream habitat and the effect of the islands in isolating populations (e.g. Schug et al., 1998; Thomas et al., 1998). It was expected that populations would be nested by island and within island by watershed (e.g. Jackson & Resh, 1992; Hughes et al., 1996; Bunn & Hughes, 1997). In addition we expected that interpopulation gene flow would be lower than in continental species, because of the greater difficulty of trans-oceanic dispersal (e.g. Mulvey et al., 1988). Our final hypothesis predicted that genetic differentiation of populations would increase with geographical distance regardless of island boundaries (e.g. Varvio-Aho & Pamilo, 1979; Dillon & Wethington, 1995).
Materials and methods
Study species
Mesophylax aspersus has a circum-Mediterranean distribution, occurring from the Canary Islands to the Near East (e.g. Schmid, 1957; Botosaneanu, 1974; Dakki, 1987). The species is common on the western Canary Islands of Tenerife (Nybom, 1948; Malmqvist et al., 1993), La Gomera (Nybom, 1954; L.C.K., unpublished data) and La Palma (L.C.K., unpublished data). M. aspersus is found in most first and second order streams at altitudes of 200–2150 m in a range of habitats including dense laurisilva woodland, open pine forest and agricultural land (Malmqvist et al., 1995).
Localities and sampling
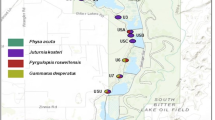
In April 1999 late-instar larvae of Mesophylax aspersus were collected from shallow pools in a set of five streams on three islands (Tenerife, La Gomera and La Palma), chosen to allow comparisons within and between catchments and islands (Fig. 1). The study streams are referred to as P1, P2, T, G1 and G2. They are located in Barranco Taburiente, La Palma, Barranco del Rio, La Palma, Barranco del Rio, Tenerife, and a tributary and the main channel at El Cedro, La Gomera, respectively. In an attempt to sample from a single population individuals were collected from two to three shallow pools in a 5–10 m stretch of stream (minimum sample size 24). Specimens were kept alive in insulated flasks of stream water then transferred to individual cryotubes within 2–3 h, for storage at −196°C until analysis.
Electrophoretic analysis
Fourteen enzyme systems were successfully screened using cellulose acetate gel electrophoresis (protocol modified from Hebert & Beaton, 1991), revealing nine putative loci (eight enzyme systems) which could be scored reliably in all five populations. The eight enzymes, the abbreviations used and their Enzyme Commission numbers (International Union of Biochemistry Nomenclature Committee, 1984) were: esterase, EST (E.C. 3.1.1._) with α-naphthyl acetate substrate; fumarate hydratase, FUM (E.C. 4.2.1.2); glucose-6-phosphate isomerase, GPI (E.C. 5.3.1.9); isocitrate dehydrogenase, IDH (E.C. 1.1.1.42); leucine aminopeptidase, LAP (E.C. 3.4._._); glycyl-L-leucine peptidase, PEP C (E.C. 3.4._._); proline dipeptidase, PEP D (E.C. 3.4._._); and phosphoglucomutase, PGM (E.C. 5.4.2.2).
Larvae were removed from their cases and homogenized in 200 μL of grinding buffer (Peakall & Beattie, 1991). Running buffers and stains were adapted from Richardson et al. (1986), Easteal & Boussy (1987), Hillis & Moritz (1990) and Hebert & Beaton (1991). 0.025 M tris-glycine buffer, pH 8.5, was used for running the EST enzyme system; 0.01 M citrate-phosphate, pH 6.4, was used for running FUM and PEP C; the other enzyme systems were run with 0.04 M tris-citrate, pH 7.6. Specific methods are available from D.T.B. on request. Rat liver tissue (adult male Sprague–Dewley rats) was run in one lane on each gel as a positive control. Loci and alleles were labelled numerically and alphabetically, respectively, in ascending order from the least to the most mobile.
Statistical analysis
The data were summarized as allele frequencies at each locus in each population with the BIOSYS-1 package (Swofford & Selander, 1989). As measures of genetic variability, the mean number of alleles (MNA) per locus, the percentage of polymorphic loci (P) at the 95% levels and expected heterozygosity (H) (Nei’s 1978 unbiased estimate) were calculated with BIOSYS-1.
Population differentiation and structure was investigated with F-statistics (Wright, 1943) estimated by the formulae of Weir & Cockerham (1984) with the GENETIX package (Université de Montpellier II, 1999). Standard deviations of the multilocus F statistic estimates were obtained by jack-knifing over loci. Comparing the observed means to the outcomes generated from permutation tests estimated significance: to test FIS, alleles were randomized within populations; to test FST, individual genotypes were randomly allocated to populations. A sequential Bonferroni correction for the analysis of multiple tests was used (Rice, 1989), calculated by hand. Multilocus FST was calculated for each pair of sites. Pairwise site comparisons were also performed using Rogers’ (1972) genetic distance, calculated with GENETIX. Significance was estimated by comparing the observed distances with a null distribution generated by recalculating the distance matrix after 1000 random reassignments of individuals to sites. A dendrogram showing the relationships between the sites was constructed by the distance Wagner (Farris, 1972) procedure with BIOSYS-1.
For each pair of sites, multilocus FST and Rogers’ genetic distance were regressed against geographical distance between sites and minimum inter-island distances, both directly and with log transformations, using Microsoft Excel. Distances were defined as the shortest measurements on the map. The relationships between the genetic and geographical distances were tested formally with Mantel tests (Mantel, 1967) in the GENETIX package.
Results
Genetic variability measures
All loci but FUM were polymorphic in at least one population, and EST, LAP-1 and PEP C were polymorphic in every population (Table 1). There was large variation in allele frequencies between populations, and at only two loci was the most common allele constant across populations. However there were only one site-specific (allele B of FUM locus at P1) and no island-specific alleles. Populations at the five sites showed different amounts of variability, with the Tenerife sample showing particularly little: MNA, P and mean H were all lowest at T, however, H was not significantly lower.
Population differentiation and structure
A summary of F-statistics is provided in Table 2. FIS was found to be very variable. P1 and G1 showed a significant excess of heterozygotes whilst P2 and G2 showed a significant deficiency. T had a nonsignificant deficiency. LAP-1 had a particular excess of heterozygotes, whilst for PEP D there was an excess of heterozygotes at site G1 but a deficiency at P2 and none at all at G2. The multilocus estimates of FIS and FIT were significantly positive. The multilocus FST was 0.250, which implies substantial population structuring (P < 0.001). All the pairwise genetic distances (both FST and Rogers’ distance) were significant (P < 0.001). The most distant pair of sites was T–G2, and the closest P1–G1 (Table 3). The branching order and relative branch lengths of the distance Wagner network showed that sites within an island were not more similar than sites on different islands.
Genetic distance and geographical isolation
Regressions of pairwise FST and Rogers’ genetic distance against geographical distances were nonsignificant. Mantel tests on each pair of matrices confirmed that there was no significant pattern of isolation by distance.
Discussion
Genetic variability compared to continental species
Levels of genetic variability in Mesophylax aspersus were generally high, except at site T (Table 1). The mean H and P were higher than any previously recorded in Trichoptera (Plague & MacArthur, 1988; Jackson & Resh, 1992; Guinand, 1994), falsifying the first stated hypothesis. MNA of M. aspersus was more typical of Trichoptera. The lack of site- or island-specific alleles suggests that the populations are not of independent origin.
The most likely cause of the high genetic variability in M. aspersus is occasional interpopulation dispersal of individuals between populations with different genetic composition, despite their geographical isolation and the dispersal barrier of the sea. If populations are of reasonable size and longevity then genetic variability can accumulate. It is also possible that balancing selection and temporal and spatial variation in selection pressures may maintain some of the genetic diversity. The genetic variability estimates are likely to be inflated by the lack of monomorphic loci in the data set; however, further work (L.C.K., unpublished data) found that of an additional 10 loci none were monomorphic.
The lower heterozygosity found in other studies of Trichoptera might also be due in part to the sampling methods employed. Attracting adults to a light trap may inadvertently sample individuals from more than one population, leading to the Wahlund effect (e.g. Plague & McArthur, 1998). On the other hand, larvae collected from a small area of a stream may represent only one or a few sibling groups (e.g. Jackson & Resh, 1992; Guinand, 1994; Bunn & Hughes, 1997).
Population structure: genetic and geographical isolation
Mesophylax aspersus has substantial population structuring on the Canary Islands (Table 2), as predicted. FIS was significantly positive overall but varied in sign from locus to locus. Possible explanations for the variability in FIS are that null alleles confounded the scoring of gels, or that selection is acting upon some loci (e.g. against homozygotes at LAP-1) whilst others are subject to genetic drift (e.g. Giles et al., 1998). The Wahlund effect may have produced significant FIS in the study by Plague & McArthur (1998) but is not likely to have operated alone in the present study, as heterozygote excess as well as deficiency was found.
The hypothesis of hierarchical population structure in M. aspersus was not supported, as population subdivision was as significant within as between islands, and same-island pairs had genetic distances in the mid-range of the pairwise distances (Table 3). The patchy nature of suitable stream habitat may make dispersal between streams on the same island as unlikely as dispersal over the sea. In contrast, Jackson & Resh (1992) found that genetic variation in Helicopsyche was hierarchically structured, with smaller differences in allele frequencies observed among sites within a stream and larger differences between catchments and regions.
Similar values of multilocus FST are reported for M. aspersus and the continental species, when populations are separated by comparable geographical distances: FST=0.425 over 200 km in Helicopsyche borealis (Jackson & Resh, 1992); FST=0.015 over 25 km in Hydropsyche exocellata (Guinand, 1994). Thus the prediction that interpopulation gene flow would be lower in M. aspersus was not supported. However single-locus FST in M. aspersus varied by two orders of magnitude, and this heterogeneity means that the multilocus estimator should be interpreted with caution (Guinand, 1994).
The final hypothesis was that FST would increase with geographical distance, whether within or between islands. This was not supported. This implies that streams will not necessarily be colonized by the nearest neighbouring population. Dispersal between sites in close proximity could be prevented by: prevailing wind direction; topography, particularly when streams are in deep gorges (as are P1, P2, and T); dense forest (as surrounds P2, G1, and G2); and low stream density (as on Tenerife). Passive dispersal over longer distances could occur if an airborne insect became caught in a wind current, as studies of insect fallout on the snowfields of Mount Teide, Tenerife (Ashmole & Ashmole, 1988), on ships and over the sea (Clarke, 1903) have demonstrated. A number of similar studies have failed to find isolation-by-distance (e.g. Jackson & Resh, 1992; Bunn & Hughes, 1997), and the stochastic effect of recruitment, random dispersal, population history and environmental structure are invoked. In this case we have shown that the division of the species’ range into an archipelago of islands does not determine its genetic structure, and the genetic variability within populations suggests that the stochastic effect of recruitment is also not the cause of the population structure.
Conclusion: genetic differentiation and dispersal
Bohonak (1999) found that there is a robust relationship between population structure and dispersal ability: genetic distance estimates are informative and patterns of dispersal do, in the majority of comparisons, make a measurable contribution to observed population genetic structure. This study makes use of this paradigm to infer dispersal ability from genetic differentiation in order to investigate the relationship between dispersal ability and distribution of Mesophylax aspersus. It is likely that M. aspersus is the strongest flier of the Canarian trichopteran fauna (Gothberg, 1973; Svensson, 1974), and adult flight is the principal mechanism of dispersal in Trichoptera (Bunn & Hughes, 1997). We conclude that a small amount of distance-independent dispersal of individuals between populations occurs, which has allowed M. aspersus to colonize almost all the permanent streams in the archipelago. Whilst the paucity of streams on the Canary Islands leaves freshwater fauna isolated in an otherwise arid environment, populations of M. aspersus appear to be large and persistent enough, and receive enough genetically distinct immigrants, to maintain high levels of genetic variability within them.
References
Alarie, Y. and Bilton, D. T. Larval morphology of Hydrotarsus Falkenström, 1938: Generic characteristics, description of H. compunctus (Wollaston, 1865), and analysis of relationships with other members of the tribe Hydroporini (Coleoptera: Dytiscidae, Hydroporinae). Coleopterists Bull, in press.
Ashmole, N. P. and Ashmole, M. J. (1988). Insect dispersal on Tenerife, Canary Islands: high altitude fallout and seaward drift. Arctic Alpine Res, 20: 1–12.
Belyea, L. R. and Lancaster, J. (1999). Assembly rules within a contingent ecology. Oikos, 86: 402–416.
Bohonak, A. J. (1999). Dispersal, gene flow, and population structure. Q Rev Biol, 74: 21–45.
Botosaneanu, L. (1974). Notes descriptives, faunistiques, écologiques, sur quelques trichoptères du ‘trio subtroglophile’ (Insecta: Trichoptera). Travaux Institut Spéologie Emil Racovitza, 13: 61–75.
Bunn, S. E. and Hughes, J. M. (1997). Dispersal and recruitment in streams: evidence from genetic studies. J North Am Benthol Soc, 16: 338–346.
Clarke, W. E. (1903). Vanessa cardui and other insects at the Kentish Knock lightship. Entomol Monthly Mag, 39: 289–290.
Dakki, M. (1987). Ecosystèmes d’eau courante du haut Sebou (Moyen Atlas): études typologiques et analyses écologiques et biogéographiques des principaux peuplements entomologiques. Travaux Institut des Sciences, Rabat, Serie de Zoologie, 42: 25–38.
Dillon, R. T. and Wethington, A. R. (1995). The biogeography of sea islands: clues from the population genetics of the freshwater snail, Physa heterostropha. Syst Biol, 44: 400–408.
Easteal, S. and Boussy, I. A. (1987). A sensitive and efficient isozyme technique for small arthropods and other invertebrates. Bull Entomol Res, 77: 407–415.
Farris, J. S. (1972). Estimating phylogenetic trees from distance matrices. Am Nat, 106: 645–668.
Giles, B. E., Lunqvist, E. and Goudet, J. (1998). Restricted gene flow and subpopulation differentiation in Silene dioica. Heredity, 80: 715–723.
Gothberg, A. (1973). Dispersal of lotic Trichoptera from a north Swedish stream. Aquilo Series Zoologica, 14: 99–104.
Guinand, B. (1994). Investigations on the genetic differentiation of two populations of Hydropsyche exocellata Dufour (Trichoptera) in the upper Loire River (France) and ecological implications. Zool Anz, 232: 1–14.
Hebert, P. D. N. and Beaton, M. J. (1991). Methodologies for Allozyme Analysis using Cellulose Acetate Electrophoresis. Helena Laboratories, Beaumont, TX.
Hillis, D. M. and Moritz, C., eds. (1990). Molecular Systematics. SinauerAssociates, Sunderland, MA.
Hughes, J. M., Bunn, S. E., Hurwood, D. A. and Choy, S. et al (1996). Genetic differentiation among populations of Caridina zebra (Decapoda: Atyidae) in tropical rainforest streams, northern Australia. Freshwater Biol, 36: 289–296.
International Union of Biochemistry Nomenclature Committee. (1984). Enzyme Nomenclature. Academic Press, Orlando, FL.
Jackson, J. K. and Resh, V. H. (1992). Variation in genetic structure among populations of the caddisfly Helicopsyche borealis from three streams in northern California. Freshwater Biol, 27: 29–42.
Juan, C., Emerson, B. C., Oromi, P. and Hewitt, G. M. (2000). Colonisation and diversification: towards a phylogeographic synthesis for the Canary Islands. Trends Ecol Evol, 15: 104–109, 10.1016/s0169-5347(99)01776-0.
Macarthur, R. H. and Wilson, E. O. (1967). The Theory of Island Biogeography. Princeton University Press, Princeton, NJ.
Machado, A. (1992). Monographie de los Carabìdos de las Islas Canarias. Instituto de Estudios Canarios, La Laguna, Tenerife.
Malmqvist, B., Nilsson, A. N., Baez, M. and Armitage, P. D. et al (1993). Stream macroinvertebrate communities in the island of Tenerife. Arch Hydrobiologie, 128: 209–235.
Malmqvist, B., Nilsson, A. N. and Baez, M. (1995). Tenerife’s freshwater macroinvertebrates – status and threats (Canary Islands, Spain). Aquatic Conserv – Mar Freshwater Ecosystems, 5: 1–24.
Mantel, N. (1967). The detection of disease clustering and a generalised regression approach. Cancer Res, 27: 209–220.
Mulvey, M., Newman, M. C. and Woodruff, D. S. (1988). Genetic differentiation among West Indian populations of the schistosome-transmitting snail Biomphalaria glabrata. Malacologia, 29: 309–317.
Nei, M. (1978). Estimation of average heterozygosity and genetic distance from a small number of individuals. Genetics, 89: 583–590.
Nybom, O. (1948). The Trichoptera of the Atlantic Islands. Commentationes Biologicae Societas Scientarum Fennica, 8: 1–19.
Nybom, O. (1954). Entomological results of the Finnish expedition to the Canary Islands 1947–51. No. 9. Some additions to the Trichopterous fauna of the Canary Islands. Commentationes Biologicae Societas Scientarum Fennica, 14: 1–3.
Pashley, D. P., Rai, K. S. and Pashley, D. N. (1985). Patterns of allozyme relationships compared with morphology, hybridisation, and geologic history in allopatric island-dwelling mosquitoes. Evolution, 39: 985–997.
Peakall, R. and Beattie, A. J. (1991). The genetic consequences of worker ant pollination in a self-compatible, clonal orchid. Evolution, 45: 1837–1848.
Plague, G. R. and Mcarthur, J. V. (1998). Genetic diversity vs. geographic distribution of five congeneric caddisflies. Hydrobiologia, 362: 1–8, 10.1023/a:1003185225656.
Rice, W. R. (1989). Analyzing tables of statistical tests. Evolution, 43: 223–225.
Richardson, B. J., Baverstock, P. R. and Adams, M. (1986). Allozyme Electrophoresis: a Handbook for Animal Systematics and Population Studies. Academic Press, London.
Rogers, J. S. (1972). Measures of genetic similarity and genetic distance. Studies Genet VII, University Texas Publ, 7213: 145–153.
Schmid, F. (1957). Les genres Stenophylax Kol., Micropterna St. et Mesophylax McL. (Trichoptera, Limnephilidae). Trabajos del Museo de Zoología, Barcelona, Nueva Serie Zoológica, 2: 1–51.
Schug, M. D., Downhower, J. F., Brown, L. P. and Sears, D. B. et al (1998). Isolation and genetic diversity of Gambusia hubbsi (mosquitofish) populations in blueholes on Andros Island, Bahamas. Heredity, 80: 336–346.
Sheldon, A. L. (1984). Colonisation dynamics of aquatic insects. In: Resh, V. H. and Rosenberg, D. M. (eds). The Ecology of Aquatic Insects, pp. 401–429. Praeger Scientific, New York.
Slatkin, M. (1985). Gene flow in natural populations. Ann Rev Ecol Syst, 16: 393–430.
Svensson, B. W. (1974). Population movements of adult Trichoptera at a South Swedish stream. Oikos, 25: 157–175.
Swofford, D. L. and Selander, R. B. (1989) BIOSYS-1. A computer programme for the analysis of allelic variation in population genetics and biochemical systematics. Release 1.7. University of Illinois, Urbana, IL.
Thomas, E. P., Blinn, D. W. and Keim, P. (1998). Do xeric landscapes increase genetic divergence in aquatic ecosystems? Freshwater Biol, 40: 587–593.
UNIVERSITÉ DE MONTPELLIER II. (1999). GENETIX 4.0. Logiciel sous WindowsTMpour la génétique des populations. Laboratoire Génome, Populations, Interactions, University of Montpellier II, Montpellier.
Varvio-Aho, S. -L. and Pamilo, P. (1979). Genic differentiation of Gerris lacustris populations. Hereditas, 90: 237–249.
Wallace, A. R. (1880) Island Life. Macmillan, London.
Weir, B. S. and Cockerham, C. C. (1984). Estimating F-statistics for the analysis of population structure. Evolution, 38: 1358–1370.
Wright, S. (1943). Isolation by distance. Genetics, 28: 114–138.
Acknowledgements
This study was financially supported by a studentship from the University of Plymouth. Marcos Baez, University of La Laguna, Tenerife, and Björn Malmqvist, University of Umeå, Sweden, are thanked for their kind assistance in the field. The manuscript was improved by the comments of two anonymous referees.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kelly, L., Bilton, D. & Rundle, S. Population structure and dispersal in the Canary Island caddisfly Mesophylax aspersus (Trichoptera, Limnephilidae). Heredity 86, 370–377 (2001). https://doi.org/10.1046/j.1365-2540.2001.00839.x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1046/j.1365-2540.2001.00839.x