Abstract
Wolbachia are maternally transmitted endocellular bacteria infecting several arthropod species. In order to study Wolbachia segregation rate, Drosophila simulans females from an Indo-Pacific population (Seychelles) bi-infected by the two Wolbachia variants wHa and wNo were backcrossed to uninfected males in two conditions. In the first case, Seychelles males from a stock cured from its Wolbachia by tetracycline treatment were used. In the second case, the males came from a naturally uninfected Tunisian population. It was found that (i) the two Wolbachia variants can segregate, so that bi-infected females can produce a few offspring infected only by wHa or wNo. This occurs in both backcross conditions. (ii) Segregation leads more frequently to wHa than to wNo mono-infection. (iii) Wolbachia transmission is lower when the Seychelles genome is introgressed by the Tunisian genome, suggesting that host genomic factors might influence infection fate.
Similar content being viewed by others
Main
Wolbachia are endocellular bacteria infecting numerous species of arthropods (O’Neill et al., 1997). Found in the germline of both sexes, they are only transmitted maternally through the cytoplasm of the egg. The infection can result in various alterations of sexuality and reproduction such as feminization, thelytokous parthenogenesis, and cytoplasmic incompatibility. Recently, male-killing has also been described (Hurst et al., 1999). All these phenomena enhance the spread of infected cytoplasmic lines. The most common phenomenon, cytoplasmic incompatibility (CI), is an embryonic mortality occurring among the progeny of crosses between infected males and females uninfected or infected by another Wolbachia strain (Werren, 1997).
The presence of Wolbachia can be detected through their effect on the host or by PCR amplification of Wolbachia genes. Wolbachia can be separated, using molecular data, into two groups (named A and B) that diverged 58–67 Ma (Werren et al., 1995b). The existence of double infections (the presence of two different Wolbachia strains in the same individuals) was first suspected by Breeuwer et al. (1992) in the parasitoid wasp Nasonia vitripennis because individual DNA extracts yielded two different bacterial 16S sequences. Using specific primers for group A and B based on the FtsZ gene, Werren et al. (1995b) found 7 double infections among 29 arthropod species known to be infected. The same assay was used to screen a random sample of 154 neotropical insect species (Werren et al., 1995a). Of these, 26 harboured Wolbachia, including 9 species in which individuals harboured two different Wolbachia 16S and ftsZ sequences simultaneously. It was then apparent that double infections were not exceptional, and indeed were more frequent than expected by chance, considering the frequency of A and B mono-infections (Werren et al., 1995a).
One well-known Wolbachia host is Drosophila simulans, where a double infection was first suspected based on 16S molecular data (Rousset & Solignac, 1995) and confirmed experimentally by segregation of the two variants involved (Merçot et al., 1995). This double infection is a result of the simultaneous presence of a group A and a group B Wolbachia, called, respectively, wHa and wNo (Rousset & Solignac, 1995). In the wild, bi-infected flies are found in the Seychelles archipelago and in New Caledonia (Merçot et al., 1995; Rousset & Solignac, 1995). In other pacific islands, only flies harbouring wHa have been found (O’Neill & Karr, 1990; Merçot et al., 1995; Rousset & Solignac, 1995). Moreover, a few individuals infected only by wHa can be found in bi-infected strains, while individuals infected only by wNo have not been found in the wild although they can be generated in the laboratory (Merçot & Poinsot, 1998a).
As a result of the double infection, males infected by both wNo and wHa induce CI in crosses with females infected either by wHa or wNo only (Merçot et al., 1995; Merçot & Poinsot, 1998a). Accordingly, should Wolbachia segregation take place in a bi-infected population, the resulting mono-infected female offspring would be incompatible with most males, and the mono infections would be eliminated. It thus remains that in bi-infected populations, rare individuals carrying a wHa mono-infection can be found, while the wNo mono-infection seems completely absent. This might be because CI expressed by bi-infected males reaches 90% embryonic mortality against females mono-infected by wNo, but only 65% against females mono-infected by wHa (Merçot et al., 1995; Merçot & Poinsot, 1998a). Another cause of the absence of the wNo mono-infection might be a very rare occurrence of segregation toward wNo mono-infected individuals.
In this study, we present an experiment where cytoplasmic lineages naturally bi-infected by wHa and wNo were subjected to repeated backcrosses with uninfected males following two different procedures. The first backcross procedure did not change the genetic background since the males used had the same origin as the female foundresses, except that their stock had been cured from Wolbachia infection by an antibiotic treatment. The second backcross procedure used males from a naturally uninfected population, carrying a different genetic background. We observed that: (i) segregation leads preferentially to wHa rather than wNo mono-infection; and (ii) host nuclear background seems to influence the efficiency of Wolbachia maternal transmission.
Materials and methods
Drosophila strains
The Seychelles strain was derived from flies collected on Mahe Island (Seychelles archipelago) in 1981. Most individuals from this strain are bi-infected by both Wolbachia variants, wHa and wNo, but a few are infected only by wHa (Merçot et al., 1995; Rousset & Solignac, 1995). Seychelles-TC is a stock from the Seychelles strain, cured from its Wolbachia following a Tetracycline treatment. Nasr’allah is a naturally uninfected strain collected in the Nasr’allah oasis (Tunisia) in 1983. The strains were reared at 25°C on standard axenic medium (David, 1962). All tests also took place at 25°C.
Backcrosses
Three bi-infected females (defining three original cytoplasmic lineages noted A, B and C) were crossed individually with two Seychelles males. Twelve F1 females from each lineage were then chosen at random and placed individually with two Seychelles males. In each F2 progeny obtained, two females were chosen at random. One was crossed with two Seychelles-TC males to initiate an S-line. The other was crossed with two Nasr’allah males, to initiate an N-line. These individuals constitute back-cross generation G0. The lines were then maintained through 18 generations by crossing at each generation a single female chosen at random with two Seychelles-TC males (for the 36 S-lines, corresponding to series AS, BS and CS) or with two Nasr’allah males (for the 36 N-lines, corresponding to series AN, BN, CN). The name of a line is formed by the letter corresponding to the original lineage (A, B or C), the number of the F1 female (1–12) and the type of backcross (S or N).
Male incompatibility mass test
Fifteen 2–3-day-old virgin males of the lines under test were allowed to mate with 10 4–5-day-old virgin Nasr’allah females for 8 h in a vial containing standard axenic medium. Flies were transferred for oviposition on fresh axenic medium darkened with vegetal charcoal powder. After 24 h, the adults were discarded and the eggs kept at 25°C for at least another 24 h before hatch rate was estimated on 96–200 eggs per line with a mean of 167.
Individual male incompatibility test
Individual crosses were carried-out using 24–36-h-old virgin males under test and 4–5-day-old virgin Nasr’allah females. Each cross was initiated by placing one male and one female in a vial containing darkened axenic medium until mating was observed. The male was then removed and the female left to oviposit, with three further transfers to fresh tubes over 48 h. Hatch rate was estimated as above. Only the crosses yielding at least 20 eggs have been considered (with a mean of 85 eggs).
PCR-RFLP
The presence and nature of the Wolbachia was determined as previously described by PCR amplification of 16S Wolbachia DNA followed by digestion with VspI (Merçot et al., 1995). The PCR product is uncut in the case of wHa (897 bp) and yields two fragments (531 and 365 bp) for wNo; bi-infected flies exhibiting a three band pattern. Each PCR was performed with a Wolbachia-positive control extract. Each VspI digestion experiment included a PCR product from a wNo-infected control to check that digestion was complete.
Results
Among the 36 S-lines, one (A9S) was lost in G3, and two others (A1S and A10S) were discarded because the G0 females were found to be mono-infected, respectively, by wHa and wNo. The corresponding N-lines (A1N, A9N and A10N) were also discarded to maintain the paired structure of the data.
Wolbachia segregation
Starting from bi-infected lineages, lines mono-infected by wHa or wNo were been obtained by segregation in both backcross procedures throughout the generations (Fig. 1). In addition, some lines have completely lost the infection. For each original lineage (A, B, or C), we have compared the frequencies of the four possible infection statuses between the backcrosses (Table 1). A significant backcross effect is observed in lineages A and B, but not in lineage C. In lineage A, this effect is a result of the apparition of a significantly higher number of uninfected lines in series AN compared to series AS, from the G12 onwards. In G12, all 9 lines were still infected in series AS but 6/9 were uninfected in series AN. In lineage B, the backcross effect appeared from the G6 onward: this is because segregation was rarer in series BS than in series BN (e.g. in G9, 11 BS lines were still bi-infected and one was uninfected while in the BN series, only 5 lines were still bi-infected, 5 were mono-infected by wHa and 2 were mono-infected by wNo). By the end of the experiment, most BN lines (8/12) were devoid of Wolbachia, whereas only 2/12 BS lines had lost all infection.
Sequence of events leading to complete infection loss
For each line found infected at a given generation, we checked the infection status three generations later. The frequencies of each possible event were then estimated by pooling the observations over the 18 generations (Fig. 2). Bi-infected lines became totally uninfected only 7 times out of 221 possibilities, vs. 26 times out of 98 possibilities for mono-infected lines (χ2=39.94, 1d.f., P < 0.001). In consequence, the status preceding total infection loss was nearly always a mono-infection. The usual sequence of events leading to uninfected cytoplasm in our experiment was therefore that of bi-infected lines becoming first mono-infected and then uninfected.
Infection transmission rate
The fidelity of bi-infection transmission was significantly higher in backcross S than in backcross N (χ2=8.16, 1d.f., P < 0.01). This difference was a result of bi-infected females producing mono-infected lines less frequently in backcross S (χ2=8.85; 1d.f., P < 0.01). However, bi-infected females did not produce significantly more uninfected lines in backcross N (χ2=0.004, 1d.f., NS). Overall, when considering mono- and bi-infected females, the production of uninfected lines was significantly more frequent in backcross N than in backcross S (17.69% vs. 6.02%; χ2=7.97, 1d.f., P < 0.01).
Wolbachia segregation led to wHa mono-infection twice as often as it did to wNo mono-infection (33 vs. 16 events when considering the 33 S-lines and all the 36 N-lines where this information could be searched for: χ2=5.89, 1d.f., P < 0.05). In both backcrosses, wNo mono-infected lines seemed to lose their infection more frequently than wHa mono-infected lines (Fig. 2), but this difference is not significant (χ2=1.56, 1d.f., NS).
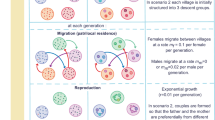
In backcross S (i.e. without any perturbation resulting from the introgression of foreign genes), we have estimated the following parameters (per generation): the transmission rates of bi-infection, wHa mono-infection and wNo mono-infection (pBi, pHa and pNo, respectively), and the segregation rates of bi-infection toward wHa or wNo mono-infection (segHa and segNo, respectively). Assuming all parameters to be constant over time, the probabilities of each possible observation over three generations can be described as follows:

Resolving these equations using the observed data yields pBi=0.939, pHa=0.943, pNo=0.909, segHa=0.035 and segNo = 0.018. Then, by difference, the rate of total infection loss by bi-infected flies can be estimated at 0.008 per generation (as compared to 0.057 in wHa mono-infected flies and 0.091 in wNo mono-infected flies).
Cytoplasmic incompatibility tests
We have measured the level of CI induced by males (in mass test) of the original lineages (A, B, C) at the beginning of the experiment. The results were as follows: A: 95.0%; B: 92.0%; C: 92.5% (percentage of unhatched eggs out of 200 eggs). It has been shown that several generations of backcross with uninfected males decreased the level of CI induced by males from infected backcrossed lines, correlatively with a decreased bacteria load in male gonads (Merçot et al., 1995). We have measured CI in males from our lines after 13 (mass crosses) and 16 (mass and individual crosses) generations of backcross, to see if the CI decrease would depend on their infection status. Strictly speaking, this comparison only applies between bi-infection and the wHa mono-infection. This is because males from strains mono-infected by wHa have been shown to induce levels of CI similar to those of bi-infected males (Merçot et al., 1995; Rousset & Solignac, 1995), whereas males from wNo mono-infected strains have been shown to induce significantly lower levels of CI (Merçot et al., 1995; Merçot & Poinsot, 1998a).
For the mass crosses, the results are shown in Table 2 and have been analysed, after arcsine transformation, using an ANOVA (SAS, 1989: GLM procedure, type IIISS). The infection type effect is significant (F230= 9.05; P < 0.001), but not the backcross effect nor the interaction. We then compared the means of each infection type (LSMEANS/TDIFF statement in SAS, 1989). This comparison shows that the CI induced by bi-infected males (77.8 ± 3.0%) was significantly higher than in males mono-infected by wHa (54.7 ± 5.4%; P < 0.001) or by wNo (50.5 ± 3.7%; P < 0.01), while the wHa and wNo mono-infections did not differ significantly in that respect. Among the bi-infected lines tested in G13, four were found to be mono-infected in G16, with a significant decrease of CI in three cases (Table 3). On the contrary, out of 10 lines still bi-infected in G16, only two showed a lower CI than in G13, with one significant case (A2S: 96.0% vs. 80.9%, χ2=21.55, 1d.f., P < 0.01; A7S: 88.0% vs. 81.0%, χ2=3.74, 1d.f., NS).
For the individual crosses, Fig. 3 shows for each line the distribution of CI induced by individual males when crossed with uninfected females. The results have been analysed using an ANOVA as above. The backcross effect is significant (F1109=12.44; P < 0.001), which corresponds to a higher CI in the S backcross (this difference was already apparent in mass crosses, [Table 2], although it was not significant). The infection type effect is also significant (F2109=56.36; P < 0.001), but not the interaction backcross × infection type (F2109= 2.61; NS). We then compared the means of each infection type (LSMEANS/TDIFF statement in SAS, 1989). This comparison shows that the CI induced by bi-infected males was significantly higher than that of males mono-infected by wHa or wNo (P < 0.001 in both cases). In addition, the CI induced by males mono-infected by wHa was this time significantly higher than that of males mono-infected by wNo (P < 0.001).
Individual measurements of the cytoplasmic incompatibility induced by males sampled in G16 of the backcross. CI is estimated by the percentage of unhatched eggs when a male is crossed with an uninfected (Nasr’allah) female. The lines tested are ranked on the Y-axis according to the average CI of their males (within brackets). Each dot represents the level of CI induced by one male.
Discussion
Segregation
We have observed the segregation of wHa and wNo in two conditions of backcross in lines derived from Seychelles bi-infected females. The present results confirm those obtained previously in the other D. simulans population (Nouméa, New Caledonia) known to be bi-infected by wHa and wNo (Merçot & Poinsot, 1998a). In particular, we have been able to obtain wNo mono-infected individuals, which have not yet been described in natural populations. It should be noted that two G0 females were found to be mono-infected (one by wHa, the other by wNo), demonstrating that mono-infected individuals can be produced even without backcrossing with uninfected males. The production of mono-infected offspring by bi-infected females can therefore not be attributed to the experimental procedure and can be expected to occur in the wild. The use of uninfected males in our experiments simply prevented the elimination of mono-infected eggs at the embryo stage through incompatibility with sperm from bi-infected males (Merçot et al., 1995).
The general rule in our experiment is clearly the cotransmission by bi-infected females of both Wolbachia variants, showing that the two bacteria coexist and develop within the same cellular lineages. However, the fact that bi-infected females can produce mono-infected eggs (estimated at 3.5% wHa and 1.8% wNo per generation in the backcross S) and rarer uninfected eggs (estimated at 0.8% per generation in the backcross S) suggests a stochastic mechanism of transmission. If one assumes that the number of Wolbachia transmitted from the mother to its eggs varies stochastically from egg to egg, it appears that most mono-infected eggs will be found among those which received a low number of Wolbachia (because the higher the number of Wolbachia transmitted, the lower the probability that they all belong to the same bacterial type). As a result, the mono-infected lines generated during our experiment could be expected to carry a lower bacterial load than average. This would explain why they were significantly more likely to completely lose the infection than lines which had remained bi-infected.
An alternative hypothesis is that Wolbachia do not segregate at random but instead have a significant tendency to remain together in clusters. We consider this hypothesis less likely (indeed, it would not seem adaptive at all from the Wolbachia point of view because it would enhance the probability that some eggs will be uninfected), but this ‘clustering behaviour’ is still a theoretical possibility, and could result from selective pressures completely unrelated to transmission efficiency. For completeness, it should be noted that the opposite ‘scattering behaviour’ (individual Wolbachia moving away one from another) might be especially adaptive during transmission, because it might spread the few Wolbachia there are in weakly infected females more evenly between eggs, perhaps ensuring a maximum transmission rate. The individual and collective behaviour of Wolbachia (if any) remains unknown.
Measurements of CI carried out on males from backcrossed lines support the first hypothesis of a stochastic segregation linked to low infection loads. In Drosophila, the CI induced by a given Wolbachia variant is correlated to its load in the males (Bressac & Rousset, 1993; Rousset & de Stordeur, 1994; Solignac et al., 1994; Bourtzis et al., 1996). If segregation is correlated with a diminution of bacterial load, a decrease of CI is also expected during the backcross when lines become mono-infected. Indeed, our observations show that males from mono-infected lines induce significantly less CI than males from lines that remain bi-infected. However, this comparison is only valid between the wHa mono-infected lines and the bi-infected lines, because it is known that such lines induce similar levels of CI when maintained in brother-sister mating (Merçot et al., 1995; Rousset & Solignac, 1995), whereas the CI induced by males from wNo lines maintained in brother-sister mating is significantly lower (Merçot et al., 1995; Merçot & Poinsot, 1998a,b).
Influence of host nuclear background
Out of our three initially bi-infected cytoplasmic lineages, a significant backcross effect in G18 is observed in lineages A and B. In lineage A, this effect is explained by the appearance of a higher number of uninfected lines in series AN compared to series AS. In lineage B, the effect is found because segregation is lower in series BS than in series BN and because most BN lines became uninfected during the experiment. Moreover, individual tests revealed that S backcross males induced significantly higher CI than N backcross males. Assuming that both transmission and CI are positively correlated with Wolbachia load, these results suggest that the genome of the host influences the fate of the infection. A suitable hypothesis is that indo-pacific Wolbachia have adapted to the genome of their usual host, and suffer from the introgression of foreign genes. This might induce a slower bacterial growth with lower bacterial loads and resulting in lower transmission rates. An alternative hypothesis is that introgressing foreign genes led to a perturbation in host homeostasis, with lower bacterial loads simply as an aspecific by-product. We note in particular that the mtDNA variant of Seychelles D. simulans is SiI, whereas the nuclear genome of Nasr’allah is associated with the SiII mtDNA variant in the original strain (Baba-Aïssa et al., 1988). During a previous isolation of wNo from the Nouméa strain (harbouring the SiI mtDNA variant), the introgression by the Nasr’allah nuclear genome resulted in an abnormally low hatch rate independently of the Wolbachia infection (Merçot et al., 1995). However, these data are not sufficient to determine the precise cause of the backcross effect observed. Nevertheless, the present work does suggest that, within a given species, the genome of the host can influence Wolbachia fate. Such an influence had already been demonstrated clearly at the interspecific level between Drosophila simulans and D. melanogaster (Boyle et al., 1993; Poinsot et al., 1998), between D. simulans and D. serrata (Clancy & Hoffmann, 1997) and between the two hymenopteran parasitoids Nasonia vitripennis and N. giraulti (Bordenstein & Werren, 1998).
Comparison between wHa and wNo
Segregation in the offspring of bi-infected females led twice as often to wHa mono-infections than to wNo mono-infections. Thus, either wHa was more abundant in bi-infected individuals than wNo, or its transmission was intrinsically better on average in our experimental conditions. Our results do not allow us to decide between these two hypotheses. In mono-infected lines maintained in brother-sister mating, wHa-infected males induce a significantly higher CI than wNo-infected males both in mass tests (Merçot et al., 1995; Merçot & Poinsot, 1998a) and in individual tests (Merçot & Poinsot, 1998b). In such cases, the hypothesis of a lower load for wNo is not supported by observations made by Bourtzis et al. (1998). Using individual dot-blot measurements on total DNA extracts from testes tissue, these authors did not find a lower Wolbachia load in a wNo mono-infected strain compared to a strain mono-infected by wHa.
wNo: a parasite of wHa?
When two Wolbachia are present as mono-infections only, the variant inducing the higher level of CI should eliminate the weaker variant from the host population. In such a competition, wHa should wipe wNo out of the host population, because males infected by wHa induce more CI towards females infected by wNo than in the reverse cross (Merçot et al., 1995; Merçot & Poinsot, 1998a). However, bi-infection represents a puzzling case wherein the level of reciprocal CI between the two variants will not allow the stronger to eliminate the weaker. The reason is, of course, that bi-infected females are compatible with all males (Merçot et al., 1995; Perrot-Minot et al., 1996) and thus benefit from the highest fitness. The Seychelles bi-infection would then be an efficient shield, allowing the maintenance of wNo at the expense of wHa, a situation that could almost be described as parasitism of wHa by wNo. Indeed, wNo is only found in the wild in the bi-infection wHa + wNo. This bi-infection predates the divergence between D. simulans and the Seychelles archipelago endemic species D. sechellia, which took place at least 0.5 Ma (Rousset & Solignac, 1995). Yet, several wild populations of D. simulans, probably established during human migrations (Hawaii, French Polynesia), are infected only by wHa (O’Neill & Karr, 1990; Merçot et al., 1995), implying a secondary loss of the wNo variant. Bi-infection is therefore not an absolute guarantee of survival for the weaker partner.
References
Baba-Aïssa, F., Solignac, M., Dennebouy, N. and David, J. (1988). Mitochondrial DNA variability in Drosophila simulans: quasi absence of polymorphism within each of the three cytoplasmic races. Heredity, 61: 419–426.
Bordenstein, S. R. and Werren, J. H. (1998). Effects of A and B Wolbachia and Host genotype on interspecies cytoplasmic incompatibility in Nasonia. Genetics, 148: 1833–1844.
Bourtzis, K., Dobson, S. L., Braig, H. R. and O’neill, S. L. (1998). Rescuing Wolbachia have been overlooked. Nature, 391: 852–853.
Bourtzis, K., Nirgianaki, A., Markakis, G. and Savakis, C. (1996). Wolbachia infection and cytoplasmic incompatibility in Drosophila species. Genetics, 144: 1063–1073.
Boyle, L., O’neill, S. L., Robertson, H. M. and Karr, T. L. (1993). Interspecific and intraspecific horizontal transfer of Wolbachia in Drosophila. Science, 260: 1796–1799.
Breeuwer, J. A. J., Stouthamer, R., Barns, S. M., Pelletier, D. A., Weisburg, W. G. and Werren, J. H. (1992). Phylogeny of cytoplasmic incompatibility micro-organisms in the parasitoid wasp genus Nasonia (Hymenoptera: Pteromalidae) based on 16S ribosomal DNA sequences. Insect Mol Biol, 1: 25–36.
Bressac, C. and Rousset, F. (1993). The reproductive incompatibility system in Drosophila simulans: DAPI-staining analysis of the Wolbachia symbionts in sperm cysts. J Invert Pathol, 63: 226–230.
Clancy, D. and Hoffmann, A. A. (1997). Behavior of Wolbachia endosymbionts from Drosophila simulans in Drosophila serrata a novel host. Am Nat, 149: 975–988.
David, J. (1962). A new medium for rearing Drosophila in axenic conditions. Dros Inf Ser, 93: 28.
Hurst, G. D. D., Jiggins, F. M., von Der Schulenburg, J. H. G., Bertrand, D., West, S. A., Goriacheva, I. I. et al (1999). Male-killing Wolbachia in two species of insects. Proc R Soc B, 266: 735–740.
Merçot, H., Llorente, B., Jacques, M., Atlan, A. and Montchamp-Moreau, C. (1995). Variability within the Seychelles cytoplasmic incompatibility system in Drosophila simulans. Genetics, 141: 1015–1023.
Merçot, H. and Poinsot, D. (1998a). Wolbachia transmission in a naturally bi-infected Drosophila simulans strain from New Caledonia. Entomol Exp Appl, 86: 97–103.
Merçot, H. and Poinsot, D. (1998b). Rescuing Wolbachia have been overlooked and discovered on mount Kilimanjaro. Nature, 391: 853–853.
O’neill, S. L., Hoffmann, A. A. and Werren, J. H. (1997). Influential Passengers: Inherited Microorganisms and Arthropod Reproduction, Oxford University Press, Oxford.
O’neill, S. L. and Karr, T. L. (1990). Bi-directional incompatibility between conspecific populations of Drosophila simulans. Nature, 348: 178–180.
Perrot-Minot, M.-J., Guo, L. R. and Werren, J. H. (1996). Single and double infections with Wolbachia in the parasitic wasp Nasonia vitripennis: effects on compatibility. Genetics, 143: 961–972.
Poinsot, D., Bourtzis, K., Markakis, G., Savakis, C. and Merçot, H. (1998). Wolbachia tranfer from Drosophila melanogaster into D. simulans: host effect and cytoplasmic incompatibility relationships. Genetics, 150: 227–237.
Rousset, F. and de Stordeur, E. (1994). Properties of Drosophila simulans strains experimentally infected by different clones of the bacterium Wolbachia. Heredity, 71: 325–331.
Rousset, F. and Solignac, M. (1995). Evolution of single and double Wolbachia symbioses during speciation in the Drosophila simulans complex. Proc Natl Acad Sci USA, 92: 6389–6393.
SAS (1989). STAT User’s Guide, Version 6 4th edn. SAS Institute, Cary, NC.
Solignac, M., Vautrin, D. and Rousset, F. (1994). Widespread occurrence of the proteobacteria Wolbachia and partial cytoplasmic incompatibility in Drosophila melanogaster. C R Acad Sci Paris, 317: 461–470.
Werren, J. H. (1997). Biology of Wolbachia. Ann Rev Entomol, 42: 587–609.
Werren, J. H., Guo, L. R. and Winsor, D. W. (1995a). Distribution of Wolbachia in neotropical arthropods. Proc R Soc B, 262: 197–204.
Werren, J. H., Zhang, W. and Guo, L. R. (1995b). Evolution and phylogeny of Wolbachia: reproductive parasites of arthropods. Proc R Soc Lond B, 261: 55–63.
Acknowledgements
We thank A. Atlan and D. Higuet for helpful comments on the manuscript. We also thank E. Aspect, V. Delmarre and C. Labellie for technical support. This work was supported by a grant from the French Ministère de l’Enseignement Supérieur et de la Recherche (ACC SV3 n°9503017) and DP was supported by a PhD grant from the MESR.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Poinsot, D., Montchamp-Moreau, C. & Merçot, H. Wolbachia segregation rate in Drosophila simulans naturally bi-infected cytoplasmic lineages. Heredity 85, 191–198 (2000). https://doi.org/10.1046/j.1365-2540.2000.00736.x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1046/j.1365-2540.2000.00736.x
Keywords
This article is cited by
-
Symbiont-Driven Male Mating Success in the Neotropical Drosophila paulistorum Superspecies
Behavior Genetics (2019)
-
High-sensitivity detection of cryptic Wolbachia in the African tsetse fly (Glossina spp.)
BMC Microbiology (2018)
-
Population genomics of Wolbachia and mtDNA in Drosophila simulans from California
Scientific Reports (2017)
-
More than fishing in the dark: PCR of a dispersed sequence produces simple but ultrasensitive Wolbachia detection
BMC Microbiology (2014)
-
New Insight into Wolbachia Epidemiology: Its Varying Incidence During the Host Life Cycle Can Alter Bacteria Spread
Bulletin of Mathematical Biology (2014)