Abstract
We studied the population genetic structure of nine bisexual Artemia sinica populations from the provinces of Inner Mongolia, Shanxi and Qinghai in China, using variation at nine allozyme loci (cellulose acetate electrophoresis). There is a clear-cut tendency for an increase in genetic variation, as measured by heterozygosity, with increasing habitat size. Although we observe a positive relationship between genetic differentiation and geographical distance, overall FST values are low: populations separated by approximately 1000 km show average FST values of 0.05–0.1, whereas populations separated by 100 km show no genetic differentiation at all.
Similar content being viewed by others
Introduction
The brine shrimp Artemia. (Crustacea: Anostraca) is normally restricted to saline inland lakes and coastal salterns (Vanhaecke et al., 1987). In common with other large branchiopods, the brine shrimp Artemia is capable of producing diapausing cysts that can withstand adverse conditions such as anoxia, drying, freezing, mechanical disturbance and digestive enzymes, and these cysts are therefore believed to be very suited for passive dispersal by wind, waterfowl or man (Persoone & Sorgeloos, 1980). The presumed good dispersal capacities of the brine shrimp translate into the expectation that, even though genetic differentiation among populations will increase with geographical distance (Slatkin, 1985), overall levels of genetic differentiation will be low. The island-like nature of inland waters may, however, provide opportunities for genetic differentiation, and several studies have indeed emphasized strong among-populational genetic differentiation in various zooplankton taxa, even though many of these species have attributes (e.g. resting eggs) that promote dispersal (Boileau et al., 1992; Boileau & Taylor, 1994; De Meester, 1996). Being confronted with these conflicting expectations, we set out to analyse genetic differentiation among Chinese Artemia populations inhabiting salt lakes that are separated by up to 1500 km.
Artemia has proven suitable for the study of evolutionary processes such as speciation and genetic or morphometric differentiation (Browne & Bowen, 1991; Pilla & Beardmore, 1994; Triantaphyllidis et al., 1997a,b). Allozyme electrophoresis has so far mainly been used for the resolution of taxonomical status, although a number of studies have also considered genetic differentiation among local populations (Bowen & Sterling, 1978; Beardmore & Abreu-Grobois, 1983; Abatzopoulos et al., 1993; Gajardo et al., 1995). At least five bisexual species and many parthenogenetic species are currently recognized in Artemia (Pilla & Beardmore, 1994; Gajardo et al., 1995). Artemia is distributed in many salt lakes of north-western China, including Inner Mongolia, Xinjiang and Qinghai provinces (Xin et al., 1994). Chinese Artemia are either bisexual or parthenogenetic. A bisexual strain in Xiechi Lake of Yunchen, Shanxi province, has been characterized as A. sinica by Cai (1989), and Hou et al. (1993, 1997) found A. sinica in another 11 Chinese bisexual populations. Pilla & Beardmore (1994) showed a close genetic relationship between one Inner Mongolian population (unknown lake locality) and the Xiechi population (Yuncheng, Shanxi province). Recently, a new species (named as A. tibetiana) has been described from Tibet by Abatzopoulos et al. (1998). So far, genetic studies on Chinese populations have focused on taxonomic problems and the identification of strains. In the present paper, we provide more detailed information on levels of within-population genetic variation and among-population genetic differentiation in A. sinica populations from China, mainly from Inner Mongolia. Artemia populations have been cited from 39 salt lakes in this area (Ren et al., in press), and these lakes vary widely in size and ecological conditions. In total, we studied nine populations: seven from Inner Mongolia, one from Qinghai and one from Shanxi province.
Materials and methods
Cyst collection and Artemia culture
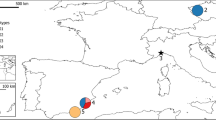
The cysts used to initiate laboratory cultures were collected by staff of the Salt Research Institute (SRI) and the Inner Mongolian Salt company of China, and were stored in the cyst bank of SRI. The localities from which cysts were collected are shown in Fig. 1. More information on the salt lakes sampled is given in Table 1.
Location map showing outline of China and indicating the salt lakes that were sampled. For abbreviations, see Table 1.
The cysts of each population were disinfected and hatched according to the methods described by Sorgeloos et al. (1986). All populations were cultured at a density of 100 nauplii/L in 3-L glass bottles containing 80 ppt artificial seawater (made up with the commercial sea salt Instant Ocean) under fluorescent light providing a photoperiod of 16:8 light:dark, in a temperature-controlled room (20°C ± 1°C). The Artemia were kept under mild aeration and were fed daily with algae (Dunaliella tertiolecta Butcher). The brine was renewed weekly.
Allozyme electrophoresis
Approximately equal numbers of randomly drawn adult males and females (50–80 from each population) from the stock cultures were stored at −80°C and used for allozyme analysis. Cellulose acetate electrophoresis (following Hebert & Beaton, 1989) was used to screen for allelic variation at nine loci. We initially assessed variation for 19 enzymes: AAT (EC 2.6.1.1), ADH (EC 1.1.1.1), AK (EC 2.7.4.3), AMY (EC 3.2.1.1), AO (EC 1.2.3.1), APK (EC 2.7.3.3), FUM (EC 4.2.1.2), G6PDH (EC 1.1.1.49), HEX (EC 2.7.1.1), IDH (EC 1.1.1.42), LDH (EC 1.1.1.27), MDH (EC 1.1.1.37), ME (EC 1.1.1.40), MPI (EC 5.3.1.8), 6PGDH (EC 1.1.1.44), PGI (EC 5.3.1.9), PGM (EC 5.4.2.2.), SOD (EC 1.15.1.1), XDH (EC 1.1.1.204). Nine loci (AAT, APK, IDH-1, IDH-2, LDH, MDH-1, MDH-2, MPI and PGI ) could be reliably stained and proved polymorphic, and they were chosen for our study.
Data analysis
All statistical analyses of allozyme data were performed with TFPGA (Tools for Population Genetic Analyses) version 1.3 (Miller, 1997) and with GENEPOP 1.2 (Raymond & Rousset, 1995). The percentage of polymorphic loci (0.99 criterion) was calculated for each population. Two estimates of heterozygosity were calculated: direct count and expected heterozygosities under Hardy–Weinberg equilibrium. Tests for deviations from H.–W. equilibrium and for linkage disequilibrium were carried out using exact tests (GENEPOP). Genetic structuring within and between populations was estimated using F-statistics. The significance of the among-population genetic differentiation over all loci was calculated by the exact test. To test for a correlation between genetic differentiation and geographical distance, we calculated pairwise FST values for all possible pairs of the eight populations that remained after excluding the one Inner Mongolian population of unknown locality. The correlation of this measure of genetic differentiation with the logarithm of geographical distance between the populations was analysed using a Mantel test for dependent variables (Sokal & Rohlf, 1995). The same analysis was also performed separately for the six populations from Inner Mongolia.
Results
The percentage of polymorphic loci and the mean heterozygosity in the nine populations studied are shown in Table 2. Twelve out of 59 tests for deviations cf from H.–W. equilibrium yielded P-values <0.05, the deviation being significant at the 0.05 level in three cases after sequential Bonferroni correction for table-wide errors. We found no evidence for linkage disequilibrium (nine tests out of 324 yielded P-values <0.05; none significant at the 0.05 level after sequential Bonferroni correction for table-wide errors). The mean heterozygosity (expected) ranged from 0.110 to 0.161. Haolebaoqing (HL) and Xiechi (XC) showed the highest percentage of polymorphic loci, and Xiechi (XC) displayed the highest value of heterozygosity. When we plot expected heterozygosity for each population against the surface area of the habitat, there is a clear tendency for higher heterozygosities with larger surface area (Fig. 2; the regression is significant, P = 0.04).
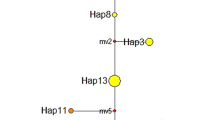
Figure 3 shows the UPGMA dendrogram constructed from a matrix of pairwise values of Nei’s (1972) genetic distance for all nine populations studied. The Xiechi population (XC) from Shanxi province is clearly separated from all Inner Mongolia populations. Within the group of populations from Inner Mongolia, the populations from Yimeng cluster together and are genetically very similar. The only odd feature in the dendrogram is the fact that the Xiaocaidan population (XCD) from Qinghai province clusters among the Ximeng populations (Inner Mongolia). The average distance between the Xiaocaidan and the Inner Mongolia populations is approximately 1200 km.
UPGMA dendrogram of Nei’s (1972) genetic distance for nine Artemia sinica populations sampled from different provinces in China. For abbreviations of lake names, see Table 1.
Single-locus and average values of FIS, FIT and FST for all nine populations are shown in Table 3. The mean value of FST over all nine loci and all populations is 0.083, indicating moderate genetic differentiation. Allele frequencies differ significantly among populations by an exact test (all nine loci combined, P < 0.0001).
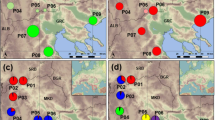
A significant correlation (P < 0.05) was found between pairwise FST values and the logarithm of geographical distance for all eight A. sinica populations for which the exact location is known (Fig. 4a) and for the six Inner Mongolia populations (Fig. 4b) using a Mantel test. There is a clear tendency for an increase in genetic differentiation with increased geographical distance. The value of the correlation coefficient (r) was increased to 0.83 when performing the analysis on the six Inner Mongolia populations only. The most striking feature of Fig. 4, however, is that populations that are separated by as much as 100 km are not genetically differentiated at all (FST ≈ 0).
Correlation between pairwise values of genetic differentiation among populations (FST) and the logarithm of geographical distance between the habitats from which these populations were sampled: (a) for all eight Artemia sinica populations from which the exact locality is known (r = 0.51, P = 0.048); (b) for all six Inner Mongolia populations (r = 0.83, P = 0.035).
Discussion
Average values of heterozygosity and the percentage of polymorphic loci observed in our study are comparable with values from several other studies on bisexual Artemia (Abreu-Grobois, 1987; Pilla & Beardmore, 1994; Gajardo et al., 1995). The mean FST value (0.083) obtained for the nine A. sinica populations studied by us is, however, smaller than the FST values that have been calculated for several conspecific A. franciscana (FST = 0.24–0.38) and A. salina (FST = 0.12) populations (Abreu-Grobis & Beardmore, 1982; Abreu-Grobois, 1987; Gajardo et al. unpubl. data). As these other studies did not span larger geographical distances than ours (>1500 km), we can conclude that genetic differentiation among the A. sinica populations studied by us is rather low. Figure 4 shows that populations that are separated by 100 km show an average FST value approximating 0, whereas populations separated by as much as 1200 km have FST values ranging from as low as 0.02 to 0.25. This low level of genetic differentiation may reflect the good dispersal capacities of the resting stages (cysts) and transport by migratory waterfowl and wind. The FST values obtained here are lower than the values reported for other Crustacea, such as Daphnia (Lynch & Spitze, 1994). However, one should recall that because of linkage effects, average FST values in cyclically parthenogenetic organisms such as Daphnia are expected to be higher than in obligately sexual organisms (see Vanoverbeke & De Meester, 1997). But our values of interpopulational genetic differentiation are also low when compared to obligately bisexual zooplankton (Boileau & Taylor, 1994) and freshwater anostracans (Riddoch et al., 1994; Brendonck et al., in press; these studies observed significant interpopulational genetic differentiation in populations separated by a few tens of metres only). It remains speculative to try to explain this discrepancy, but it is likely that the size of the habitats plays a role. All populations studied by us inhabit rather large lakes (3.5–64 km2; estimated volumes 2.4–39 × 106 m3) and are estimated to consist of >109 individuals, which may have considerably reduced the impact of two random processes. First, large populations reduce the speed with which populations differentiate from each other through genetic drift. Secondly, the impact of founder events may have been reduced because ‘effective’ propagule sizes colonizing the habitat may have been large, thus effectively sampling regional diversity. We hypothesize that in large lakes, the ‘effective’ propagule size is large because colonization success of immigrants remains relatively high for a long time. The latter follows from the fact that it takes longer before a colonizing population approaches carrying capacity in a large than in a small habitat. This argument is based on the idea that effective gene flow is often much lower than dispersal, and that the discrepancy between the two increases with time, as the establishment success of immigrants reduces in habitats in which competition with resident conspecifics is severe (De Meester, 1996).
References
Abatzopoulos, T. J., Triantaphyllidis, C. and Kastrisis, C. (1993). Genetic polymorphism in two parthenogenetic Artemia population from North Greece. Hydrobiologia, 250: 73–80.
Abatzopoulos, T. J., Zhang, B. and Sorgeloos, P. (1998). International study on Artemia LIX: Artemia tibetiana: preliminary characterization of a new Artemia species found in Tibet (People’s Republic of China). Int J Salt Lake Res, 7: 41–44.
Abreu-Grobois, F. A. (1987). A review of the genetics of Artemia. In: Sorgeloos, P., Bengtson, D. A., Decleir, W. and Jaspers, E. (eds) Artemia Research and its Applications, vol. 1, Morphology, Genetics, Strain Characterization, Toxicology, pp. 61–99. Universa Press, Wetteren, Belgium.
Abreu-Grobis, F. A. and Beardmore, J. A. (1982). Genetic differentiation and speciation in the brine shrimp Artemia. In: Barigozzi, C. (ed.) Mechanisms of Speciation, pp. 345–376. Alan R. Liss, New York.
Beardmore, J. A. and Abreu-Grobois, F. A. (1983). Taxonomy and evolution in the brine shrimp Artemia. In: Oxford, G. S. and Rollinson, D. (eds) Protein Polymorphism: Adaptive and Taxonomic Significance, pp. 153–164. Academic Press, London.
Boileau, M. G. and Taylor, B. E. (1994). Chance events, habitat age, and the genetic structure of pond populations. Archiv Hydrobiol, 132: 191–202.
Boileau, M. G., Hebert, P. D. N. and Schwartz, S. S. (1992). Non-equilibrium gene frequency divergence: persistent founder effects in natural populations. J Evol Biol, 5: 25–39.
Bowen, S. T. and Sterling, G. (1978). Esterase and malate dehydrogenase isozyme polymorphism in 15 Artemia populations. Comp Biochem Physiol, 61B: 593–595.
Brendonck, L., De Meester, L. and Riddoch, B. J. Spatial patterns of genetic variation in Branchipodopsis wolfi (Crustacea: Anostraca). Oecologia. in press.
Browne, R. A. and Bowen, S. T. (1991). Taxonomy and population genetics of Artemia. In: Browne, R. A., Sorgeloos, P. and Trotman, C. (eds) Artemia Biology, pp. 221–236. CRC Press, Boca Raton, FL.
Cai, Y. (1989). A redescription of the brine shrimp (Artemia sinica). Wasmann J Biol, 47: 105–110.
De Meester, L. (1996). Local genetic differentiation and adaptation in freshwater zooplankton populations: Patterns and processes. Ecoscience, 3: 385–399.
Gajardo, G. M., Conceicao, M., Weber, L. and Beardmore, J. A. (1995). Genetic variability and interpopulational differentiation of Artemia strains from South America. Hydrobiologia, 302: 21–29.
Hebert, P. D. N. and Beaton, M. J. (1989) Methodologies for Allozyme Analysis using Cellulose Acetate Electrophoresis. Helena Laboratories, Beaumont, TX.
Hou, L., Cai, H. and Zhou, X. (1993). A study on isozymes of ten Artemia strains from China. Acta Zool Sinica, 39: 30–37.
Hou, L., Cai, H., Zhou, X. and Yang, G. (1997). Expression of isozyme genes and taxonomic status of bisexual Artemia from China. Acta Zool Sinica, 43: 184–191.
Lynch, M. and Spitze, K. (1994). Evolutionary genetics of Daphnia. In: Real, L. A. (ed.) Ecological Genetics, pp. 109–128. Princeton University Press, Princeton, NJ.
Miller, M. P. (1997) Tools for Population Genetic Analyses. (Tfpga), version 1.3. Department of Biological Sciences, Northern Arizona University, Flagstaff, AZ.
Nei, M. (1972). Genetic distance between populations. Am Nat, 106: 283–296.
Persoone, G. and Sorgeloos, P. (1980). General aspects of the ecology and biogeography of Artemia. In: Persoone, G., Sorgeloos, P., Roels, O. and Jaspers, E. (eds) The Brine Shrimp Artemia, vol. 3, Ecology, Culturing, Use in Aquaculture, pp. 3–24. Universa Press, Wetteren, Belgium.
Pilla, E. J. S. and Beardmore, J. A. (1994). Genetic and morphometric differentiation in Old World bisexual species of Artemia (the brine shrimp). Heredity, 73: 47–56.
Raymond, M. and Rousset, F. (1995). Genepop (version 1.2): population genetics software for exact tests and ecumenicism. J Hered, 86: 248–249.
Ren, M., Guo, Y., Wang, J., Su, R., Li, H. and Ren, B. Survey of Artemia ecology and researches in inland salt lakes in Northwest China. Heilongjiang. in press.
Riddoch, B. J., Mpoloka, S. W. and Cantrell, M. (1994). Genetic variation and localized gene flow in the fairy shrimp, Branchipodopsis wolfi, in temporary rainwater pools in Southeastern Botswana. In: Beaumont, A. R. (ed.) Genetics and Evolution of Aquatic Organisms, pp. 96–102. Chapman & Hall, London.
Slatkin, M. (1985). Gene flow in natural populations. Ann Rev Ecol Syst, 16: 393–430.
Sokal, R. R. and Rohlf, F. J. (1995) Biometry, 3rd edn. W. H. Freeman, New York.
Sorgeloos, P., Lavens, P., Leger, P., Tackaert, W. and Versichele, D. (1986) Manual for the Culture and Use of Brine Shrimp Artemia in Aquaculture, State University of Gent Press, Gent, Belgium.
Triantaphyllidis, G. V., Criel, G., Abatzopoulos, T. J. and Sorgeloos, P. (1997a). International study on Artemia. LIII. Morphological study of Artemia with emphasis to Old World strains. II. Bisexual populations. Hydrobiologia, 357: 139–153.
Triantaphyllidis, G. V., Criel, G., Abatzopoulos, T. J. and Sorgeloos, P. (1997b). International study on Artemia. LIV. Morphological study of Artemia with emphasis to Old World strains. II. Parthenogenetic populations. Hydrobiologia, 357: 155–163.
Vanhaecke, P., Tackaert, W. and Sorgeloos, P. (1987). The biogeography of Artemia an updated review. In: Sorgeloos, P., Bengtson, D. A., Decleir, W. and Jaspers, E. (eds) Artemia Research and its Applications, vol. 1, Morphology, Genetics, Strain Characterization, Toxicology, pp. 129–155. Universa Press, Wetteren, Belgium.
Vanoverbeke, J. and De Meester, L. (1997). Among-populational genetic differentiation in the cyclical parthenogen Daphnia magna (Crustacea, Anomopoda) and its relation to geographic distance and clonal diversity. Hydrobiologia, 360: 135–142.
Xin, N., Sun, J., Zhang, B., Triantaphyllidis, G. V., Vanstappen, G. and Sorgeloos, P. (1994). International Study on Artemia, resources in the People’s Republic of China. Int J Salt Lake Res, 3: 105–112.
Acknowledgements
X.N. was recipient of a fellowship of the Flemish Government while carrying out this study. We thank the staff of the Salt Research Institute (SRI), the Inner Mongolian Salt company of P. R. China and the ARC (Gent) for providing us with cyst material. This work was financially supported by project BIL/96/43 of the Flemish Government and by projects G.0260.97 of the Fund for Scientific Research, Flanders and OT/96/13 of the Research Fund KULeuven. We thank two anonymous referees for valuable comments on an earlier version of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Naihong, X., Audenaert, E., Vanoverbeke, J. et al. Low among-population genetic differentiation in Chinese bisexual Artemia populations. Heredity 84, 238–243 (2000). https://doi.org/10.1046/j.1365-2540.2000.00664.x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1046/j.1365-2540.2000.00664.x
Keywords
This article is cited by
-
Diapausing egg banks, lake size, and genetic diversity in the rotifer Brachionus plicatilis Müller (Rotifera, Monogononta)
Hydrobiologia (2017)
-
Dormancy and dispersal as mediators of zooplankton population and community dynamics along a hydrological disturbance gradient in inland temporary pools
Hydrobiologia (2017)
-
Phylogeography and genetic structure of the Mediterranean killifish Aphanius fasciatus (Cyprinodontidae)
Marine Biology (2007)
-
Contrasting patterns of genetic variation in the two sympatric geckos Gekko tawaensis and G. japonicus (Reptilia: Squamata) from western Japan, as revealed by allozyme analyses
Heredity (2003)