Abstract
In plant populations alleles often deviate from a random distribution and reveal positive autocorrelation at short distances. In species with both clonal and sexual reproduction, such clustering may be because ramets of the same genet were sampled at nearby locations. Alternatively, clustering may be the result of limited gene flow through pollen or seeds (isolation-by-distance). Here, we modify a conventional spatial autocorrelation analysis using the join-count statistic in order to differentiate between these two causes of genetic structure. We examined the distribution of seven microsatellite loci representing 37 alleles in a 20 × 80 m plot of a perennial population of eelgrass Zostera marina L. In analysing join-counts between all like genotypes we found significant genetic autocorrelation among ramets at distances between 1 and 7 m (P < 0.001). We then excluded joins between clonemates which were identified from the expected likelihood of their seven-locus genotypes. Without joins within genets, no autocorrelation was evident, indicating that most of the significant genetic clustering was caused by clonal spread. At distances up to 27 m, alleles were distributed at random, indicating a panmictic population at this spatial scale. These results illustrate the need for an a priori estimation of genet–ramet structure in clonally reproducing plants in order to avoid erroneous inferences about putative gene flow at various spatial scales.
Similar content being viewed by others
Introduction
Spatial autocorrelation techniques are a powerful tool for detecting the nature and scale of genetic differentiation over a range of spatial scales (e.g. Sokal & Oden, 1978a,b; Cliff & Ord, 1981). Simulations have demonstrated that these methods are able to distinguish between several causes of genetic structure, i.e. directional migration, selection or restricted gene flow (isolation-by-distance, Sokal et al., 1997). In contrast to statistics based on allele frequencies such as FST, autocorrelation techniques make few assumptions regarding the underlying population genetic model (Heywood, 1991). No information is lost by pooling of individuals into arbitrary sampling areas for subsequent comparison of gene frequencies among subpopulations. Instead, in autocorrelation analyses, the explicit information of each spatial location of individuals relative to one another can be utilized (reviewed in Heywood, 1991; Epperson, 1993). Therefore, autocorrelation techniques may have a superior power over methods based on nested samples of gene frequencies for detecting spatial patterns (Epperson & Li, 1996; Epperson, 1997).
The numerical value of an autocorrelation statistic is often examined as a function of the Euclidean distance among pairs of plants, a correlogram. The shape of the correlogram allows inferences about the direction and magnitude of the evolutionary processes at work (Sokal et al., 1997). Isolation-by-distance typically results in a clustered distribution of like genotypes which corresponds to positive autocorrelation values at short distances (Sokal & Wartenberg, 1983). In contrast, directional migration and selection typically lead to clinal patterns in genetic structure (Sokal et al., 1997) which can be further examined with correlograms that include the directionality of the autocorrelation. The latter two processes will not be considered further in this paper.
Clonal growth, the vegetative production of modular individuals (ramets sensuHarper, 1977) from a sexually produced individual (i.e. a genet), is also expected to lead to strong spatial autocorrelation in allele and genotype distribution. Wright (1969) pointed out that in vegetatively reproducing species the genetic similarity should be maximal at short distances if nearby ramets belong to the same genet. Although many sessile animal species and many plants are able to reproduce clonally (overview in Jackson et al., 1985), theoretical studies on expected spatial genetic patterns have not considered the potential contribution of clonal growth (e.g. Sokal & Wartenberg, 1983; Epperson & Li, 1996; Epperson, 1997; Sokal et al., 1997). Likewise, in field studies, the contribution of clonal reproduction to significant genetic autocorrelation has not been properly differentiated from isolation-by-distance, although the importance of both processes is generally acknowledged (e.g. Hossaert-McKey et al., 1996; Caujapé-Castells & Pedrola-Montfort, 1997; Kang & Chung, 1997; but see Montalvo et al., 1997); for example, in predominantly outcrossing populations with large clones, an inclusion of several ramets belonging to the same genet will suggest restricted gene flow and a small genetic neighbourhood if clonemates are not identified a priori.
We present an autocorrelation method that can assess the contribution of vegetative spread to a positive genetic autocorrelation at small distances. Using microsatellite data we demonstrate the usefulness of the method in examining the spatial genetic structure of eelgrass Zostera marina L., a marine angiosperm or seagrass which is widely distributed in coastal waters of the northern hemisphere (Den Hartog, 1970). Many seagrasses, including eelgrass, can reproduce both sexually and vegetatively. Within-population genetic structure in seagrasses may hence be caused either by restricted pollen and seed movement or by vegetative spread.
In plant groups with low allozyme variability such as seagrasses (Les, 1988; Laushman, 1993), the resolution power needed to examine within-population genetic structure, or to differentiate between ramets and genets, is only provided by DNA-based markers such as RAPD (Waycott, 1998), restriction fragment length polymorphism (RFLP; Alberte et al., 1994) or microsatellites (Reusch et al., 1999a). We used the polymorphism displayed by microsatellites, single-locus markers which can be amplified by PCR and scored by size (Queller et al., 1993; Schlötterer & Pemberton, 1994), to examine within-population structure in eelgrass.
Materials and methods
Study site and sampling design
We selected a perennial Zostera marina (eelgrass) population at Falkenstein, Baltic Sea (54°24′N, 10°12′E), in a depth of 2.4 m to 3.2 m (mid-bed). The sample area of 20 m (perpendicular to shore) × 80 m (parallel to shore) represents a small portion of a continuous eelgrass meadow growing from 2 m to 4 m depth. An area of 20 × 80 m was chosen because it was expected that it would contain at least four genetic neighbourhoods based on the available data on pollen and seed dispersal (Orth et al., 1994; Ruckelshaus, 1996). The inclusion of at least four spatial clusters is recommended by Epperson (1997) to maximize the statistical power of an autocorrelation analysis. To avoid sampling of physically connected ramets the minimal sampling distance between neighbouring ramets was 1 m. Within the rectangular plot, we sampled and mapped 80 randomly chosen ramets in August 1997 by means of SCUBA diving. Shoot densities/0.25 m2 (vegetative + reproductive ± SE) ranged from 29 ± 6 in July 1997 to 36 ± 9 in June 1998. The number of reproductive shoots was 1.9 ± 0.55 and 4 ± 1.1, respectively. Our sample represents 0.04% of the total number of ramets within the 1600 m2 sampling area.
Samples were preserved by drying in silica-gel. After a CTAB extraction of the genomic DNA (Doyle & Doyle, 1987) plants were genotyped for seven polymorphic microsatellite loci (Reusch et al., 1999a), representing a total number of 37 alleles. In brief, approximately 20 ng of crude DNA extract was amplified in 10-μL PCR-reactions with incorporation of radioactive dCTP using the microsatellite locus-specific primers. The PCR products were electrophoretically separated on 6% polyacrylamide sequencing gels. Different product sizes corresponding to different microsatellite alleles were scored from autoradiograms.
Autocorrelation analysis of the distribution of alleles
To examine the spatial distribution of genotypes for each microsatellite locus, the autocorrelation analysis was performed based on the genetic similarity found between single ramets. The join-count statistic is the appropriate autocorrelation statistic for such nominal data (Sokal & Oden, 1978a,b; Cliff & Ord, 1981; Epperson, 1997). Here, a join connects two ramets of a specified nominal character (allele or genotype) within a specified distance range.
Because we expected like alleles to cluster at short distances, we focused on the analysis of joins where pairs of plants shared both alleles. In considering like joins between pairs of all possible heterozygous or homozygous genotypes, we were able to summarize information over all alleles of a given locus. Simulations have shown that such summarizing results in an amplification of the autocorrelation pattern (Epperson, 1997). This was also carried out because we found an average of 5.3 alleles per locus. As a consequence the possible number of like (or unlike) joins between pairs of specific genotypes would be exceedingly large (Sokal & Oden, 1978b; Epperson & Allard, 1989). It should be noted that one can also summarize information over alleles of a given locus in using counts of nonidentical genotypes (Epperson, 1993, 1997). In fact, the number of like joins for each locus is exactly the reciprocal of the number of unlike joins suggested by Epperson (1993, 1997).
Because we want to identify the spatial scale at which significant autocorrelations occur we explored changes in the join-count statistic as a function of the Euclidian distance between ramets. For each distance class x, all the pairs of ramets located in an annulus of (x − r ≤ distance < x + r), with r being 1 m were considered. The lowest distance class was from 1 to 3 m because the minimal sampling distance was 1 m between ramets.
No analytical solution is available to determine the expected number of joins under a random distribution if several different genotype pairings are included. Instead, for each distance class, the null hypothesis of spatial randomness was generated by permutation during which the spatial coordinates of the samples were maintained while the multilocus genotypes were swapped 10 000 times. The resulting frequency distribution of joins was used to obtain standard normal deviates (SND) by dividing through the standard deviation (Cliff & Ord, 1981). In the permutation test used in this study, the normality assumption is, strictly speaking, not necessary. The standard normal deviate=SND is retained only to be consistent with other recent publications. For significance testing, the actual SND(x) was compared with the 10 000 SND-values generated by the permutations. If, for example, the actual SND(x) exceeded 99.9% of the randomly generated SND, this is taken as strong evidence against the null hypothesis of no spatial pattern.
It is reasonable to assume that all loci are selectively neutral such that the pattern between autocorrelation and distance for each individual locus is the same, notwithstanding sampling variation among loci. Thus, to increase the statistical power of the analysis, the information was summarized over all seven loci by calculating the SND of the sum of all join-counts for each locus SND (hereafter ΣSND). We expected ΣSND to be larger than the arithmetical mean of the single-locus SND because the standard deviation used in the denominator for calculating ΣSND is approximately √7 smaller (Epperson, 1997).
The spatial autocorrelation was visually examined as a correlogram, plotting SND(x) as a function of distance. For each correlogram, the chance of committing a type I error is increased because we test the autocorrelation for each distance class in a separate statistical test. However, we expected autocorrelation values to be a decreasing function of distance if either of the two causes for genetic structure, i.e. clonal growth and isolation-by-distance, was present. A test at the smallest possible distance class was therefore considered a de facto test of the entire correlogram (Heywood, 1991) and no Bonferroni correction of significance levels was performed. The permutation tests were run on a UNIX workstation using a ‘Scheme’ program with C extensions developed by one of the authors (W. Hukriede, unpubl. computer program).
We also calculated a different autocorrelation coefficient, Moran’s I, using the method suggested by Heywood (1991) for each of the 37 alleles present in the population. Alleles are coded 1 if homozygous, 0.5 if heterozygous, and 0 if absent. Results from these calculations were examined in 37 correlograms and are in general agreement with those from the join-count statistic (results not shown).
Spatial autocorrelation considering clone assignment of ramets
Alleles may cluster spatially because the same genet, and thus an identical genotype, has been repeatedly sampled at nearby locations, or because of restricted gene flow (isolation-by-distance). To differentiate between these sources of spatial genetic structure, we first assigned ramets to genets based on the criteria outlined in Parks & Werth (1993) and Reusch et al. (1999b). In brief, the expected multilocus genotype frequency for genotypes occurring more than once owing to chance had to be <0.05, which was true in all cases. For ramets having identical genotypes which were not contiguous but separated from one another by several other different genotypes, the likelihood of finding one or more of the same genotypes within all the sampled genets was calculated and had to be <0.01 to assign two ramets to a common genet.
We then calculated the SND(x) in which the joins within genets were purposely not counted. For each distance class x, the weighting matrix W (Cliff & Ord, 1981) was modified such that joins between clonemates were set to zero. In addition, the permutation test of spatial randomness was modified as explained in Fig 1. Spatial locations originally belonging to one genet were assigned a common ‘other’ genotype during each permutation step (Fig 1b). In other words, we viewed genets having more than one ramet as extended in space. To determine distance classes in genets having two or more ramets, it was always the ramet closest to a different genotype that formed the join (Fig 1c).
Diagrammatic representation of the permutation procedure by which the null hypothesis of random genotype distribution excluding joins within genets is tested. Each plot has 10 hypothetical individuals. Each shading represents assignment to a genet at a fixed position in the plot (a). The five black genotypes were assigned to a single genet. Joins between ramets of this genet are purposely not counted. In (b), a permutation has been performed. During permutation, the five ramets are assigned a common ‘other’ genotype. Rather than taking the mid-point ramet of the circled cluster occupied by the five duplicate genotypes, joins with neighbours are accomplished as in (c). A distance class is represented by the two circles surrounding the black and the white genotypes. Each neighbour (black or white genotype) is matched by its distance to the nearest ramet of the stippled genet to define a join (straight line).
We also examined whether the exclusion of joins within genets reduced the power of the autocorrelation test. This was important in order to verify that differences in the outcome of both sets of correlograms (with and without joins within genets) were caused only by clonal structure. Alternatively, fewer joins may have decreased the statistical power. To this end, we randomly selected 56 (the number of genets) of the total number of 80 genotypes present. The autocorrelation and the permutation test were then calculated ignoring the genet assigment, i.e. allowing all joins. Thirty-five runs of different random reductions in the number of genotypes were performed.
Results
Autocorrelation analysis of the distribution of alleles
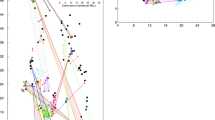
The actual spatial location of the plants in the field together with the representation of the genotypes at one microsatellite locus (Zosmar GA-2) is presented in Fig 2. Alleles at this locus are not distributed at random but cluster in groups approximately 2–9 m across, which are partly identical to the areas covered by ramets belonging to the same genet. Not surprisingly, the spatial autocorrelation analysis revealed positive autocorrelations at small distances for five of seven microsatellite loci (Fig 3a). For five loci, the SNDs of like joins exhibited a maximum at the smallest distances. At least one of the SND(x) was significant at P < 0.05, and crossed the x-axis at approximately 7–9 m. A minimum SND value was attained between 11 and 13 m, possibly reflecting the average diameter of a genetically homogeneous patch. The correlogram of locus Zosmar GA-4 showed the same trend, although it attained no significant autocorrelation at any distance. One locus (Zosmar GA-6) revealed no spatial autocorrelation at all. The single-locus pattern was amplified when summarizing the information by using ΣSND of all like joins over all seven loci. ΣSND exhibited a highly significant positive autocorrelation at short distances (first three distance classes, P < 0.001, Fig 3a). This supports suggestions by Epperson (1997) that the inclusion of several microsatellite loci will considerably increase the statistical power of an autocorrelation analysis.
Genotypes at one representative microsatellite locus and genet assignment of 80 mapped ramets of Zostera marina sampled within an area 20 × 80 m. The inner circle represents the genotype at microsatellite locus Zosmar GA-2, whereas the lighter, outer circle represents genet assignment. Those ramets not possessing a second outer circle possessed unique seven-locus genotypes. In total, the 80 ramets were assigned to 56 genets, 10 of which had more than one ramet.
Correlograms plotting the standard normal deviate (SND) of all like join-counts at seven microsatellite loci as a function of Euclidean distance. The thick lines represent ΣSND which summarizes the information over all seven loci. In (a) all joins are permitted. In (b) joins within genets were excluded as described in Fig 1 to remove the effect of clonal propagation from the correlogram. Outer horizontal lines indicate the ±95% confidence interval of SNDs at ±1.96.
The permutation test revealed that the expected frequency of the number of random runs was very close to the normal expectations, based on the standard deviation of the distribution of counts under the null hypothesis. In general, the deviation was less than ± 0.2 standard deviations of the expected confidence interval under a normal distribution. Therefore, in Fig 3, the ±95% confidence interval is approximated as ±1.96 SND for all eight correlograms.
The contribution of clonal spread to the spatial genetic structure
The 80 ramets sampled were assigned to a total of 56 genets (Fig 2). Following the procedure outlined in Fig 1, joins within genets were excluded in order to subtract the contribution of clonal spread from the calculation of spatial autocorrelation. Then, neither the correlograms of single-locus SND(x) nor of ΣSND revealed any pattern with distance (Fig 3b). At distances up to 27 m, alleles of all seven microsatellite loci were apparently distributed at random. Thus, the significant genetic clustering revealed by autocorrelation at distances from 1 to 7 m prior to the removal of joins within genets results mainly from the clonal spread of eelgrass. The distance at which the correlogram of ΣSND intersects the x-axis in Fig 3(a) can be interpreted as the average size of a genet.
There was little evidence that the significant autocorrelation disappeared because the statistical power was not sufficient for the 56 genets as opposed to the 80 ramets. When calculating the correlograms for ΣSND with 56 randomly chosen plants and ignoring the genet assignment, in 34/35 (97%) of the cases, there was at least one autocorrelation value highly significant (P < 0.001) within the distance classes of 1–7 m (data not shown). In 32/35 (92%) of the cases the shape of the entire correlogram was very similar to the representation based on all 80 ramets.
Discussion
Differentiating between causes leading to spatial clustering in allele or genotype distribution is important if we wish to draw inferences on isolation-by-distance by means of a correlogram. In clonally reproducing plants, vegetative spread will tend to blur genetic structure owing to restricted pollen and seed movement, prohibiting an assessment of neighbourhood size from spatial genetic structure (i.e. Morton, 1982). In the only study known to us, Montalvo et al. (1997) calculated a related autocorrelation statistic, the coefficient of kinship, with and without joins between clonemates in a stand of oaks using allozyme polymorphism. In this study, a positive autocorrelation at small distances decreased after excluding the contribution of joins within genets. However, in contrast to the method outlined in the present paper, only one arbitrarily chosen central ramet in each genet was chosen in Montalvo et al. (1997) to calculate the autocorrelation which changed the distances between pairs of plants present in the original spatial network.
Direct measurements of pollen and seed movement suggest that gene flow in Z. marina is limited to short distances. Orth et al. (1994) found that, even in areas with strong tidal currents (0.5 m/s), most eelgrass seeds are not transported farther than 2 m from their parent plant. In measuring the dispersal distances of eelgrass pollen and seeds, Ruckelshaus (1996) studied the genetic neighbourhood of a Pacific eelgrass population and found that dispersal distances of pollen and seeds were not more than a few metres. Based on the distribution of dispersal distances, she calculated a neighbourhood area of 524 m2 in an area with tidal currents.
At the Falkenstein study site, a subtidal eelgrass bed, which is not subjected to tidal currents (the Baltic Sea is atidal), we found a random distribution of microsatellite alleles at distances >27 m. Because the genetic autocorrelation is a nonlinear function of neighbourhood size (Morton, 1982; Sokal & Wartenberg, 1983), the size of a genetically uniform patch is not identical to the genetic neighbourhood area. Considering these limitations, our data suggest that the neighbourhood size in this eelgrass population is of the order of 572 m2 but may be much larger. We found the 56 genets at the study site to be in Hardy–Weinberg equilibrium at all loci (Reusch et al., 1999b) which further supports panmixis at this scale. Ruckelshaus (1998) also examined the Pacific eelgrass meadow mentioned above, using spatial autocorrelation of five allozyme loci. Results of that study suggested a much higher patch area (4096–6400 m2) with a random distribution of alleles than calculations based on the direct measurements.
Given the short distances found in direct measurements of dispersal it is surprising that we found no evidence of isolation-by-distance. It is unlikely that we missed the scale of autocorrelation when purposely excluding all distances between ramets <1 m. Why direct measurements of dispersal and indirect assessments of genetic stucture are not concordant needs further study. Differences between ecological measurements of seed and pollen dispersal distances and indirect methods using genetic markers have been observed in other plant populations (e.g. Campbell & Dooley, 1992). Both methods may actually measure different processes. Ecological measurements which are typically performed during a single season are subjected to high spatiotemporal variability (Slatkin, 1987). In contrast, estimates based on genetic markers integrate over several generations. In addition, estimation of within-population dispersal in Z. marina may be site- and habitat-specific because the forces, such as currents and turbulence, responsible for the transport of alleles in the aquatic habitat differ strongly between sites. Indirect and direct methods may yield concordant results when performed over several seasons at the same site. Future research will assess differences in Z. marina genetic structure at several sites in order to determine whether or not generalizations can be drawn about the spatial scale of within-population structure.
References
Alberte, R. S., Suba, G. K., Procaccini, G., Zimmerman, R. C. and Fain, S. R. (1994). Assessment of genetic diversity of seagrass populations using DNA fingerprinting: Implications for population stability and management. Proc Natl Acad Sci USA, 91: 1049–1053.
Campbell, D. R. and Dooley, J. L. (1992). The spatial scale of genetic differentiation in a hummingbird-pollinated plant: comparison with models of isolation by distance. Am Nat, 139: 735–748.
Caujapé-castells, J. and Pedrola-montfort, J. (1997). Space–time patterns of genetic structure within a stand of Androcymbium gramineum (Cav.) McBride (Colchicidae). Heredity, 79: 341–349.
Cliff, A. D. and Ord, J. K. (1981). Spatial Processes. Pion, London.
Den Hartog, C. (1970). The seagrasses of the world. Verlumdelingen Koninklije. Nederlands Akademi Wefenschapen Afdeling Natuurkunde II, 59: 1–275.
Doyle, J. J. and Doyle, J. L. (1987). A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem Bull, 19: 11–15.
Epperson, B. K. (1993). Recent advantages in correlation studies of spatial patterns of genetic variation. Evol Biol, 27: 95–155.
Epperson, B. K. (1997). Gene dispersal and spatial genetic structure. Evolution, 51: 672–681.
Epperson, B. K. and Allard, R. W. (1989). Spatial autocorrelation of the distribution of genotypes within populations of lodgepole pine. Genetics, 121: 369–377.
Epperson, B. K. and Li, T. -Q. (1996). Measurement of genetic structure within populations using Moran’s spatial autocorrelation coefficient. Proc Natl Acad Sci USA, 93: 10528–11032.
Harper, J. L. (1977). Population Biology of Plants. Academic Press, New York.
Heywood, J. S. (1991). Spatial analysis of genetic variation in plant populations. Ann Rev Ecol Syst, 22: 335–355.
Hossaert-mckey, M., Valero, M., Magda, D., Jarry, M., Cuguen, J. and Vernet, P. (1996). The evolving genetic history of a population of Lathyrus sylvestris: evidence from temporal and spatial genetic structure. Evolution, 50: 1808–1821.
Jackson, J. B. C., Buss, L. W. and Cook, R. E. (1985). Population Biology and Evolution of Clonal Organisms. Yale University Press, New Haven, CT.
Kang, S. S. and Chung, M. G. (1997). Spatial genetic structure in populations of Chimaphila japonica and Pyrola japonica (Pyrolaceae). Ann Bot Fenn, 34: 15–20.
Laushman, R. H. (1993). Population genetics of hydrophilous angiosperms. Aquat Bot, 44: 147–158.
Les, D. H. (1988). Breeding systems, population structure, and evolution in hydrophilous angiosperms. Ann Mo Bot Gard, 75: 819–835.
Montalvo, A. M., Conard, S. G., Conkle, M. T. and Hodgskiss, P. D. (1997). Population structure, genetic diversity, and clone formation in Quercus chrysolepis (Fagaceae). Am J Bot, 84: 1553–1564.
Morton, N. E. (1982). Estimation of demographic parameters from isolation by distance. Hum Hered, 32: 37–41.
Orth, R. J., Luckenbach, M. and Moore, K. A. (1994). Seed dispersal in a marine macrophyte: implications for colonization and restoration. Ecology, 75: 1927–1939.
Parks, J. C. and Werth, C. R. (1993). A study of spatial features of clones in a population of bracken fern, Pteridium aquilinum (Dennstaedtiaceae). Am J Bot, 80: 537–544.
Queller, D. C., Strassman, J. E. and Hughes, C. R. (1993). Microsatellites and kinship. Trends Evol Ecol, 8: 285–288.
Reusch, T. B. H., Stam, W. T. and Olsen, J. L. (1999a). Microsatellite loci in eelgrass Zostera marina reveal marked polymorphism within and among populations. Mol Ecol, 8: 317–322.
Reusch, T. B. H., Stam, W. T. and Olsen, J. L. (1999b). Size and estimated age of genets in eelgrass Zostera marina L. assessed with microsatellite markers. Mar Biol, 133: 519–525.
Ruckelshaus, M. H. (1996). Estimation of genetic neighborhood parameters from pollen and seed dispersal in the marine angiosperm Zostera marina L. Evolution, 50: 856–864.
Ruckelshaus, M. H. (1998). Spatial scale of genetic structure and an indirect estimate of gene flow in eelgrass, Zostera marina. Evolution, 52: 330–343.
Schlötterer, C. and Pemberton, J. (1994). The use of microsatellites for genetic analysis of natural populations. In: Schierwater, B., Streit, B., Wagner, G.P. and DeSalles, R. (eds) Molecular Ecology and Evolution. Approaches and Application. pp. 203–214. Birkhäuser, Basel.
Slatkin, M. (1987). Gene flow and the geographic structure of natural populations. Science, 236: 787–792.
Sokal, R. S. and Oden, N. L. (1978a). Spatial autocorrelation in biology 1. Methodology. Biol J Linn Soc, 10: 199–237.
Sokal, R. S. and Oden, N. L. (1978b). Spatial autocorrelation in biology 2. Some biological implications and four applications of evolutionary and ecological interest. Biol J Linn Soc, 10: 229–249.
Sokal, R. S. and Wartenberg, D. E. (1983). A test of spatial autocorrelation analysis using an isolation-by-distance model. Genetics, 105: 219–237.
Sokal, R. S., Oden, N. L. and Thomson, B. A. (1997). A simulation study of microevolutionary inferences by spatial autocorrelation analysis. Biol J Linn Soc, 60: 73–93.
Waycott, M. (1998). Genetic variation, its assessment and implications to the conservation of seagrasses. Mol Ecol, 7: 793–800.
Wright, S. (1969). Evolution and Genetics of Populations vol. 2, the Theory of Gene Frequencies. University of Chicago Press, Chicago.
Acknowledgements
This project was funded by the European Union (contract no. ERBFMBICT961789) through a Marie-Curie fellowship to T.B.H.R. The Marine Botany Department, University of Kiel, kindly provided the necessary computer facilities.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Reusch, T., Hukriede, W., Stam, W. et al. Differentiating between clonal growth and limited gene flow using spatial autocorrelation of microsatellites. Heredity 83, 120–126 (1999). https://doi.org/10.1046/j.1365-2540.1999.00546.x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1046/j.1365-2540.1999.00546.x
Keywords
This article is cited by
-
The emergence of molecular profiling and omics techniques in seagrass biology; furthering our understanding of seagrasses
Functional & Integrative Genomics (2016)
-
Sex-specific pattern of spatial genetic structure in dioecious and clonal tree species, Populus alba L.
Tree Genetics & Genomes (2016)
-
SSR-based analysis of clonality, spatial genetic structure and introgression from the Lombardy poplar into a natural population of Populus nigra L. along the Loire River
Tree Genetics & Genomes (2011)
-
Asexual and sexual reproductive strategies in clonal plants
Frontiers of Biology in China (2007)
-
Microscale genetic differentiation in a sessile invertebrate with cloned larvae: investigating the role of polyembryony
Marine Biology (2007)