Abstract
The level of similarity between the DNA of Lolium multiflorum and three Festuca species, F. arundinacea, F. pratensis and F. glaucescens, was analysed on Southern blots, using DNA–DNA hybridization, and also on chromosomes using genomic in situ hybridization (GISH). It was demonstrated that the close relationship between L. multiflorum and the allohexaploid F. arundinacea arises principally from the affinity of L. multiflorum to one of the ancestral progenitors of F. arundinacea, namely F. pratensis. Using probes made from total genomic DNA of L. multiflorum, and also of L. perenne, particular regions of high homology, described here as ‘GISH bands’ were identified on F. pratensis chromosomes. These GISH bands, in combination with a specific probe for rDNA, provide us with new markers for the identification of chromosomes within the F. pratensis complement. Restriction analysis within the rDNA repeat unit revealed additional information on the close phylogeny of L. multiflorum and F. pratensis. The rDNA restriction patterns confirm that F. arundinacea originated as a hybrid between F. pratensis and F. glaucescens.
Similar content being viewed by others
Introduction
Species within the Lolium/Festuca complex are closely related, easily hybridized and exhibit promiscuous meiotic recombination (Thomas et al., 1994). The two genera are considered to have diverged from a common ancestor and have a basic chromosome number of x=7 (Malik & Thomas, 1966). Within the range of intergeneric Festulolium hybrids available, L. multiflorum (Lm) and diploid F. pratensis (Fp), or Lm and hexaploid F. arundinacea (Fa), show especially high compatibility (Buckner et al., 1961; Cremades & Bean, 1975) and provide a particularly valuable combination of complementary agronomic traits (Thomas & Humphreys, 1991). Fp and the tetraploid F. glaucescens (Fg) are the progenitors of the allohexaploid Fa (FpFpFgFgFg1Fg1) (Humphreys et al., 1995).
Some amphiploid cultivars have been established from crosses of Lm and Fp tetraploids (Breese & Lewis, 1984; Zwierzykowski et al., 1994), and these hybrids are valuable because they combine the good forage quality of Lolium with the high stress tolerance of Festuca. A problem of amphiploid breeding is the high level of homoeologous pairing between the different genomes, which leads to genetic instability and loss of hybridity in later generations.
To overcome this ‘genotypical deterioration’, an alternative strategy, based on introgression breeding, has been developed (Humphreys, 1989). This strategy has now been used successfully to transfer drought resistance from Fa into Lm using the pentaploid hybrid Lm (2n=4x=28)×Fa (2n= 6x=42) backcrossed to diploid Lm (2n=2x=14) as the recurrent parent (Humphreys & Thomas, 1993). GISH was used to follow the physical location of the introgressed segments, and it was demonstrated that these segments originated from the Fp genome of the Fa parent, and not from Fg (Humphreys & Pašakinskienė, 1996).
It is clear that to maximize the efficiency of such introgression programmes requires the fullest possible understanding of genomic relationships among these Lolium and Festuca species, and only then can determine the ideal partners for use in breeding programmes be determined. It is for this reason that GISH studies were undertaken initially. The first investigations were made with probes from genomic DNA of Fp in the hybrid cultivars and their derivatives from the cross of Lm×Fp. These demonstrated that the Fp and Lm genomes could be distinguished on the basis of their total genomic DNAs (Thomas et al., 1994). In a later study, probes from both Fp and Fg were used to discriminate and to determine the progenitors of hexaploid Fa (Humphreys et al., 1995). A Lm probe was used for the first time to investigate phenomena associated with chromosome elimination and somatic recombination in octoploid hybrids between Lm and Fa (Pašakinskienė et al., 1997).
In the present study, the novel use of a Lm probe to reveal differences between the Fp and Fg genomes of Fa is demonstrated. The Lm probe, as well as a probe made from L. perenne (Lp) are used as markers to produce GISH bands that identify some of the individual chromosomes of the Fp set. Also included is additional information on genome relationships using the Lm probe in Southern DNA–DNA hybridization, and the pTa71 probe for restriction analysis of the rDNA.
Materials and methods
In situ hybridization
Root-tip mitotic chromosomes of Fa genotype 870 (2n=42), originating from the Dutch cv. ’Barundi’, and of the hybrid between Lp×Fa 870 were prepared after pretreatment in ice-cold water for 24 h, followed by fixation in 1:3 acetic acid–ethanol. The roots were softened in a mixture of 20% pectinase and 2% cellulase, and then squashed in 45% acetic acid.
In situ hybridization was carried out at the Institute of Grassland and Environmental Research in Aberystwyth, according to the protocol described by Anamthawat-Jónsson & Heslop-Harrison (1996). To make the probes, total genomic DNAs of Lm,Lp and Fp were sonicated to give fragments of 5–10 kb and labelled with rhodomine-4-dUTP (Amersham), using the standard protocol for the nick translation system (Gibco BRL). A probe was also made from the rDNA clone pTa71 (Gerlach & Bedbrook, 1979) by labelling with fluorescein 12-dCTP. Blocking DNA of 200–500 bp fragments from Fa was prepared by autoclaving for 2 min. The probe hybridization mixture (40 μL per slide) contained 100 ng of labelled DNA, 4–6 μg of blocking DNA, 50% formamide in 2×SSC (0.3 M sodium chloride, 30 mM trisodium citrate), 10% dextran sulphate and 0.2% SDS (lauryl sulphate). The hybridization mixtures were denatured by boiling for 5 min, then placed on ice. The chromosome preparations on the slides were denatured using 70% formamide in 2×SSC at 70°C for 2 min, dehydrated in an ice-cold ethanol series (70%, 90% and 100% for 2 min each) and allowed to dry. Hybridization mixtures were then applied to the slides and incubated at 37°C overnight. Slides were washed with 20% formamide in 0.1×SSC at 42°C for 10 min and then rinsed three times in 2×SSC. For reprobing with Fp genomic DNA, the slide was soaked in 4×SSC with Tween 20, the coverslip removed, and then washed three times in 2×SSC. The routine procedure starting with denaturation in 70% formamide was then carried out as described above. DAPI-stained and antifade-mounted slides were studied with an epifluorescent microscope. Images were photographed on Fujichrome Sensia 400 slide film.
Southern hybridization
Experiments were carried out at the Icelandic Agricultural Research Institute in Keldnaholt. The plant material comprised Lm (the Dutch var. Bartissimo), Fa (the Polish var. Terros), Fp (the Lithuanian var. Dotnuva) and Fg (an accession from the Swiss Alps). DNA was extracted from leaf material using the method of Doyle & Doyle (1990). The nonradioactive chemiluminescent method (ECL; Amersham) was used for probe labelling, and hybridization and detection were carried out according to a modification of the manufacturer's instructions. Total genomic DNA from Lm and rDNA from the clone pTa71 were labelled by cross-linking to horseradish peroxidase with glutaraldehyde. Lm DNA was denatured by boiling for 5 min before labelling. The rDNA pTa71 clone was used as a probe (5 ng cm−2) on the Southern blots of total genomic DNAs (1 μg per lane) from Lm, Fp,Fg and Fa digested with EcoRI, DraI and BamHI. Lm DNA was used as a probe on the Southern blots of genomic DNAs from Lm (control), Fp, Fg and Fa digested with Bam HI. The nylon membranes were incubated for 30–60 min at 42°C in ECL hybridization buffer containing 6 M urea, with the addition of 0.1 M sodium chloride for the Lm probe or 0.5 M for the pTa71 probe. Before applying the Lm probe, the membrane was blocked with DNA from Fa (autoclaved for 2 min) at 42°C for 30 min. The ratio of the blocking DNA probe DNA was 125×. Hybridization was carried out at 42°C overnight. After post-hybridization washes at 86% stringency (6 M urea, 0.4% SDS in 0.5×SSC) for the pTa71 probe and 97% stringency (6 M urea, 0.4% SDS in 0.1×SSC) for the Lm total DNA probe, hybridization was visualized on film by enzyme-catalysed emission of light by oxidation of luminol to give the luminographs.
Results
In situ hybridization
GISH was carried out on Fa (FpFpFgFgFg1Fg1) and on the Lp×Fa hybrid. The results are shown in Fig. 1 for a representative single cell of Fa probed in three different ways: (i) total genomic DNA from Lm (Fig. 1a); (ii) combined probe of total Lm DNA+pTa71 (Fig. 1b); (iii) total genomic DNA of Fp (Fig. 1c); and for Lp×Fa probed with total Lp (Fig. 1d).
Metaphase chromosomes from a single root meristem cell of the allohexaploid Festuca arundinacea (Fa, 2n=6x=42), probed in various ways (a, b and c), and (d) from the tetraploid hybrid of Lolium perenne (Lp)×F. arundinacea (2n=4x=28). (a) The cell was probed with rhodamine-labelled total genomic DNA of L. multiflorum (Lm) and shows cross-hybridization to all 14 chromosomes of the F. pratensis (Fp) component of the Fa genome. A number of distinctive GISH bands are present on a number of chromosome pairs of Fp (e.g. the pair with the small arrows) and also on one pair from the F. glaucescens (Fg) set (large arrows) of Fa. (b) The same cell double-probed with the genomic DNA of Lm and the pTa71 rDNA probe (green). The combination of probes permits us to recognize the chromosome pairs labelled 1, 2, 3 and 4 of Fp. Two interstitial rDNA sites are visualized on Fp and three terminal sites on Fg. These sites correspond to the bands produced by the Lm genomic probe in (a). (c) The rhodamine-labelled probe made from Fp total genomic DNA establishes the identity of the Fp chromosomes. (d) Total genomic DNA of Lp labels the seven Lp chromosomes (red) in this hybrid cell and also gives an identical pattern of cross-hybridization and chromosome-specific GISH bands on the Fp chromosomes to that produced by the genomic DNA of Lm. The numbers in (d) correspond to those in (b), and the arrow identifies a terminal NOR site on one of the Fg chromosomes.
It is clear from Fig. 1a that the Lm probe cross-hybridizes with all 14 chromosomes of Fp, which are identified by the Fp probe shown in (Fig. 1c); and the way in which this hybridization occurs is a feature of particular interest. The hybridization pattern reveals a number of localized ‘GISH bands’ that serve as markers for individual Fp chromosomes. When these GISH bands are taken in conjunction with the pTa71 probe (Fig. 1b), the marking of chromosomes is even clearer and discriminates at least four out of seven of the Fp chromosome pairs. Pair 1 has an interstitial rDNA site; pair 2 is a metacentric with two GISH bands, symmetrically located in a median position in each arm; pair 3 shows one wide band located adjacent to the centromere, plus a smaller band in the other arm; and pair 4 has a single band at the centromere.
Apart from cross-hybridizing to all Fp chromosomes, the Lm genomic DNA acts as a sequence-specific probe and detects the interstitial pTa71 site in Fp and the terminal pTa71 sites in Fg (Fig. 1a). There are six rDNA loci (three pairs) in the Lm genome (Thomas et al., 1996), and the enrichment of this particular DNA fraction in the total genomic probe accounts for the detection of these sequence-specific sites in Fa. In general, the Lm probe shows limited affinity with Fg and, apart from the terminal rDNA sites, it only hybridizes to the centromere region of a single pair of chromosomes (Fig. 1a), large arrows). Evidently, there is much closer homology between the Lm and Fp genomes than between Lm and Fg.
The same kinds and the same pattern of GISH bands on Fp are also revealed by probing the Lp×Fa hybrid with total genomic DNA from Lp (Fig. 1d).
Southern hybridization
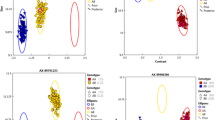
DNA from three Festuca species, Fa, Fp and Fg, digested with BamHI, show different levels of hybridization with total genomic DNA of Lm, corresponding to different levels of homology (Fig. 2). Fp clearly shows the highest level of homology with Lm, as indicated by a strong signal, whereas Fg shows very low hybridization. Fa is intermediate in intensity between Fp and Fg. In addition to these general patterns, there is a clearly identifiable common band shared by Lm, Fa and Fp of approximately 2.5 kb, which is not visible in Fg.
Genomic Southern hybridization using the ECL method. (a) Ethidium bromide-stained gels showing Bam HI digests of Lolium multiflorum (Lm),F. arundinacea (Fa),F. pratensis (Fp) and F. glaucescens (Fg), 1 μg per lane. (b) Luminograph showing hybridization of a genomic Lm probe to the same digests (a) of Lm, Fa, Fp and Fg; ECL detection 35—min. The DNA size maker is given in kb.
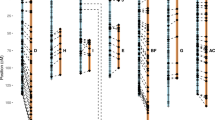
The results for the pTa71 probe are shown in the luminograph in.Fig. 3. The DNA of the four species was digested with three different restriction enzymes, EcoRI, DraI and BamHI. Because Fa is a natural hybrid between Fp and Fg, it would be expected to contain bands specific to its progenitor species. The BamHI digest is the most informative, and it demonstrates that the Fa bands derive from either Fp or Fg. It also shows that, for estimating relationships, the correspondence between dense and faint bands is as important as that between the dense bands. The difference in intensity of corresponding bands between Fg and Fa is caused by a difference in copy number. It is known from FISH (Thomas et al., 1997; this work) that Fg had lost three or four rDNA sites, whereas Fp retains them unchanged in Fa. A comparison of the similarities between Lm and Fp, and Lm and Fg, shows that Lm shares more bands with Fp than it does with Fg, in keeping with the relationships revealed by GISH and by the genomic Southern hybridizations.
Discussion
The application of GISH and Southern genomic hybridization gives strong visual evidence for high levels of homology between the genomes of Lm and Fp. Fp is a constituent genome of Fa, and this result clearly indicates that the Fp genome is the principal basis of the close relationship between Lm and Fa. The closeness between Lolium and Festuca in the section Bovinae, which includes Fp and Fa, has been reported widely (Lehvaslaiho et al., 1987; Xu & Sleper, 1994; Bulinska-Rodomska & Lester, 1998) and has led to calls to realign these species within a single genus (e.g. Darbyshire, 1993).
In this work, segmental regions on the Fp chromosomes showed an especially high affinity with a total genomic DNA Lm probe. These ‘GISH-banding’ markers offer a new dimension to chromosome discrimination in plants. What is the genetic significance of these bands? The way in which the total genomic DNA of Lm detects the rDNA in Fp suggests an explanation for their origin. Localized regions of the Fp genome appear to be represented as multiples in Lm, and this could provide a possible mechanism for genome evolution in the Festuca/Lolium complex. Another possibility is that some of them are repetitive sequences, common to Lm and Fp (Perez-Vincente et al., 1992), which are thought to be dispersed in Lm but to be present in Fp in a localized tandem repeat arrangement. However, other larger genomic rearrangements known to occur in the evolution of repetitive DNA sequences (Flavell, 1980) cannot be excluded.
The close structure between the Lm and Fp genomes explains the high levels of homoeologous meiotic pairing and recombination that takes place between them (Humphreys & Thorogood, 1993) but, notwithstanding their similarities, the two species are sufficiently divergent in their dispersed repeats to be discriminated readily by GISH. Previous evidence supporting a closer degree of homology between Lm and Fp than between Lm and Fg came from recombination studies (Humphreys & Ghesquière, 1994). The studies revealed the products of meiotic recombination involving all combinations of Lm, Fp and Fg in plants derived from Lm×Fa hybrids. However, in backcross populations between Lm×(Lm×Fa), the frequency of recombinants involving Lm×Fp was twice that found between the Lm and Fg chromosomes. It now seems possible that regions of particularly high homology, represented here as the GISH bands, could actually provide the physical basis for the high levels of recombination that occur in the hybrids between Lm and Fp species.
The rDNA gene cluster has been used widely in phylogenetics (Schlötterer, 1998), and the restriction analysis carried out here further extends our knowledge of these particular regions and reveals the close evolutionary links between the rDNAs of Lm and Fp. This result agrees with earlier work by Charmet et al. (1997), using the internal transcribed spacer (ITS), which also indicated these close relationships, as well as showing that Lolium is of more recent origin than Festuca. It is known, however, that rapid sequence changes are a characteristic feature of rDNA sites in plants (Flavell, 1986; Rogers & Bendich, 1987). Such changes are generally attributed to the intergenic spacer (IGS) region, as was clearly demonstrated in Hordeum species (Molnar et al., 1989). Furthermore, the rDNA loci can also change their map location (Dubcovsky & Dvorák, 1995) or may be lost under selection pressure resulting from adverse conditions (Fukui et al., 1994). Nevertheless, our restriction analysis shows that the Fa progenitors, Fp and Fg, have maintained their rDNA sequences in a highly conserved form, despite the fact that a number of the Fg original sites are no longer represented in Fa (Thomas et al., 1997). This kind of analysis could be useful for studying the origins of other allopolyploids, such as F. mairei (2n=4x=28) and F. atlantigena (2n=8x=56).
There are practical implications that follow from our understanding of these species relationships. It has been shown, for example, that drought-tolerant genotypes among the recombinants from Lm×Fa backcrosses carry segments of Fp introgressed into the Lm genome (Humphreys & Pašakinskienė, 1996). The molecular basis of the close homology between Lm and Fp, which is demonstrated by this work, indicates that it can be expected to produce valuable and viable new germplasm through introgression breeding.
Change history
19 January 2022
A Correction to this paper has been published: https://doi.org/10.1038/s41437-021-00488-9
References
Anamthawat-Jónsson, K. and Heslop-Harrison, J. S. (1996) Establishing relationships between closely related species using genomic DNA as a probe. In: Clapp, J. P. (eds) Methods in Molecular Biology, 50: 209–225. Humana Press, Totowa, NJ.
Breese, E. L. and Lewis, E. J. (1984). Breeding versatile hybrid grasses. Span, 27: 3–5.
Buckner, R. C., Hill, H. D. Burns, B. S. (1961). Some characteristics of perennial and annual ryegrass×tall fescue hybrids and of the amphyploid progenies of annual ryegrass×tall fescue. Crop Sci, 1: 75 80.
Bulinska-Rodomska, Z. and Lester, R. N. (1998). Intergeneric relationships of Lolium, Festuca Vulpia (Poaceae) and their phylogeny. Plant Syst Evol, 159: 217–227.
Charmet, G., Ravel, C. and Balfourier, F. (1997). Phylogenetic analysis in the Lolium–Festuca complex using molecular markers and ITS rDNA. Theor Appl Genet, 94: 1038–1046.
Cremades, P. Bean, E. W. (1975). Inflorescence development and seed production in Lolium spp. × Festuca pratensis tetraploid hybrids. J Agric Sci, 85: 301–307.
Darbyshire, S. J. (1993). Realignment of Festuca subgenus Schedonorus with genus Lolium (Poaceae). Novon, 3: 239–243.
Dubcovsky, J. and Dvorák, J. (1995). Ribosomal RNA multigene loci: nomads of the Triticeae genomes. Genetics, 140: 1367–1377.
Doyle, J. J. and Doyle, J. L. (1990). Isolation of plant DNA from fresh tissue. Focus, 12: 13–15.
Flavell, R. B. (1980). The molecular characterization of plant chromosomal DNA sequences. Annu Rev Plant Physiol, 3I: 569–596.
Flavell, R. B. (1986). The structure and control of expression of ribosomal RNA genes. Oxford Surveys Plant Mol Biol, 3: 251–274.
Fukui, K., Ohmido, N. and Khush, G. S. (1994). Variability in rDNA loci in the genus Oryza detected through fluorescence in situ hybridization. Theor Appl Genet, 8: 839–899.
Gerlach, W. L. and Bedbrook, J. R. (1979). Cloning and characterization of ribosomal RNA genes from wheat and barley. Nucl Acids Res, 7: 1869–1885.
Humphreys, M. W. (1989). The controlled introgression of Festuca arundinacea genes into Lolium multiflorum. Euphytic, 42: 105–116.
Humphreys, M. W. and Ghesquière, M. (1994). Assessing success in gene transfer between Lolium multiflorum and Festuca arundinacea. Euphytica, 77: 283–289.
Humphreys, M. W. and Pašakinskienė, I. (1996). Chromosome painting to locate genes for drought resistance transferred from Festuca arundinacea into Lolium multiflorum. Heredity, 77: 530–534.
Humphreys, M. W. and Thomas, H. (1993). Improved drought resistance in introgression lines derived from Lolium multiflorum×Festuca arundinacea hybrids. Plant Breed, 111: 155–161.
Humphreys, M. W. and Thorogood, D. (1993). Disturbed Mendelian segregation at isozyme marker loci in backcrosses of Lolium multiflorum×Festuca pratensis hybrids to Lolium multiflorum. Euphytica, 66: 11–18.
Humphreys, M. W., Thomas, H. M., Morgan, W. G., Meredith, M. R., Harper, J. A., Thomas, H. et al. (1995). Discriminating the ancestral progenitors of hexaploid Festuca arundinacea sing genomic in situ hybridization. Heredity, 75: 171–174.
Lehvaslaiho, H., Saura, A. and Lokki, J. (1987). Chloroplast DNA variation in the grass tribe Festucacea. Theor Appl Genet, 74: 298–302.
Malik, C. P. and Thomas, P. T. (1966). Karyotypic studies in some Lolium Festuca species. Caryologia, 19: 167–196.
Molnar, S. J., Gupta, P. K., Fedak, G. and Wheatcroft, R. (1989). Ribosomal DNA repeat unit polymorphism in 25 Hordeum species. Theor Appl Genet, 78: 387–392.
Pašakinskienė, I., Anamthawat-Jónsson, K., Humphreys, M. W. and Jones, R. N. (1997). Novel diploids following chromosome elimination and somatic recombination in Lolium multiflorum×Festuca arundinacea hybrids. Heredity, 78: 464–469.
Perez-Vincente, R., Petris, L., Osusky, M., Potrykus, I., Spangenberg, G. (1992). Molecular and cytogenetic characterization of repetitive DNA sequences from Lolium and Festuca: applications in the analysis of Festulolium hybrids. Theor Appl Genet, 84: 145–154.
Rogers, S. O. and Bendich, A. J. (1987). Ribosomal RNA genes in plants: variability in copy number and in the intergenic spacer. Plant Mol Biol, 9: 509–520.
Schlötterer, C. (1998) Ribosomal DNA probes and primers. In: Karp, A., Isaac, G. I. and Ingram, D. S. (eds), Molecular Tools for Screening Biodiversity, pp. 267–276 Chapman & Hall, London.
Thomas, H. and Humphreys, M. O. (1991). Progress and potential of inter-specific hybrids of Lolium and Festuca. Agric Sci, 117: 1–8.
Thomas, H. M., Morgan, W. G., Meredith, M. R., Humphreys, M. W., Thomas, H. and Legget, J. M. (1994). Identification of parental and recombined chromosomes in derivatives of Lolium multiflorum and Festuca pratensis by genomic in situ hybridization. Theor Appl Genet, 88: 909–913.
Thomas, H. M., Harper, J. A., Meredith, M. R., Morgan, W. G., Thomas, I. D., Timmis, E. and King, I. P. (1996). Comparison of ribosomal DNA sites in Lolium species by fluorescence in situ hybridization. Chrom Res, 4: 486–490.
Thomas, H. M., Harper, J. A., Meredith, M. R., Morgan, W. G. and King, I. P. (1997). Physical mapping of ribosomal DNA sites in Festuca arundinacea and related species by in situ hybridization. Genome, 40: 406–410.
Xu, W. W. and Sleper, D. A. (1994). Phylogeny of tall fescue related species using RFLPs. Theor Appl Genet, 88: 685–690.
Zwierzykowski, Z., Joks, W. and Nagonowska, B. (1994) Potential of tetraploid Festulolium (Festuca pratensis×L. multiflorum)In: Rognli, O. A., Solberg, E. and Schjelderup, I. (eds) Breeding Fodder Crops for Marginal Conditions, pp. 299–300. Kluwer, Dordrecht.
Acknowledgements
This work was supported by a Royal Society/NATO fellowship and a fellowship provided by The Ministry of Culture and Education in Iceland. I.P. wishes to thank Zina Adriulaitiene, John Harper and Ian Sant for their valuable technical assistance, and Zbigniew Zwierzykowski and Lukas Wolters for providing some seeds.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Pas̆akinskienė, I., Anamthawat-Jónsson, K., Humphreys, M. et al. New molecular evidence on genome relationships and chromosome identification in fescue (Festuca) and ryegrass (Lolium). Heredity 81, 659–665 (1998). https://doi.org/10.1046/j.1365-2540.1998.00446.x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1046/j.1365-2540.1998.00446.x
Keywords
This article is cited by
-
The current status of breeding research in Lolium genus
Journal of Crop Science and Biotechnology (2023)
-
Phylogenetic analysis of Festuca–Lolium complex using SRAP markers
Genetic Resources and Crop Evolution (2016)
-
Comparative transcriptome analysis within the Lolium/Festuca species complex reveals high sequence conservation
BMC Genomics (2015)
-
Molecular characterisation and interpretation of genetic diversity within globally distributed germplasm collections of tall fescue (Festuca arundinacea Schreb.) and meadow fescue (F. pratensis Huds.)
Theoretical and Applied Genetics (2012)
-
Evolutionary history of tall fescue morphotypes inferred from molecular phylogenetics of the Lolium-Festucaspecies complex
BMC Evolutionary Biology (2010)