Abstract
When Giemsa-stained metaphases of the grass carp Ctenopharyngodon idellus were digested with trypsin and then restained with basic Giemsa staining solution, chromosomes showed highly reproducible structural bands. The banding pattern was chromosome–specific, and corresponding chromosomes in different metaphases were identified unequivocally. A complete banding pattern of the fish with 217 bands was constructed. These results strongly indicate that good quality structural banding patterns can be obtained from metaphase chromosomes in fish.
Similar content being viewed by others
Introduction
Structural banding techniques for chromosomes are of fundamental importance to cytogenetic and evolutionary studies. Since the introduction of banding techniques for mammalian chromosomes in the early 1970s, knowledge of mammalian evolutionary genetics, medical genetics and gene mapping has vastly expanded. In contrast, the application of these chromosomal banding techniques to poikilothermic vertebrates, including fish, seems very difficult. Most of the successful chromosome banding reports in published data are restricted either to C-bands or to replication bands (Zenzes & Voiculescu, 1975; Kligerman & Bloom, 1977; Delany & Bloom, 1984; Galetti & Foresti, 1986; Mayr et al., 1987; Almeida Toledo et al., 1988; Giles et al., 1988; Hellmer et al., 1991). Distinct structural G-, R- and Q-banding patterns have only been described for the European eel Anguilla anguilla (Wiberg, 1983; Medranc et al., 1988). The difficulty in obtaining good-quality G-, R- and Q-banding patterns from the chromosomes of fish seems to be related to their chromosome structure. All available data indicate that there is a strong relationship between genome compartmentalization by base composition and Q-, R- and G-banding on chromosomes (Schweizer, 1976; Comings, 1978; Cuny et al., 1981; Furst et al., 1981; Holmquist et al., 1982; Calza et al., 1984; Goldman et al., 1984; Bernardi et al., 1985; Schmid & Guttenbach, 1988). Unlike warm-blooded mammals and birds, whose genomes can be divided into GC-rich and AT-rich compartments, cold-blooded vertebrates including fish either lack or have little compartmentalization of their genomes by base composition (Thiery et al., 1976; Hudson et al., 1980; Pizon et al., 1984; Medranc et al., 1988). These results indicate that, in order to obtain distinct multiple structural bands from the chromosomes of fish, new techniques, differing from the Q-, G- and R-banding of the chromosomes of warm-blooded vertebrates, should be used.
It is well known that, in both homeothermic vertebrates and poikilothermic vertebrates, meiotic pachytene bivalents show bead-like thickenings, the so-called chromomeres, which appear at irregular intervals along their length. In somatic cells, chromomeres are sometimes visible along interphase chromosomes. At metaphase, the chromosome is tightly coiled, and the chromomeres are no longer visible. Studies of chromomere maps in mammals have, however, revealed a very close correspondence between the dark G-bands of the mitotic prometaphase chromosomes and the chromomeres of pachytene bivalents (Ford et al., 1968; Burdick et al., 1974; Luciani et al., 1975; De Torres & Abrisqueta, 1978; Jagiello & Fang, 1982; Fang & Jagiello, 1991). I have discovered that good-quality chromomere maps can be induced in fish bivalents in pachytene spermatocytes (C. Luo, unpubl. results). These investigations suggest that it is possible to develop a multiple structural banding technique on mitotic metaphase chromosomes in fish. Here, a compound structural banding technique for fish metaphase chromosomes that produces satisfactory structural banding patterns is presented.
Materials and methods
Animals
Two male and four female specimens of grass carp Ctenopharyngodon idellus were obtained from a local fish farmer. The diploid chromosome number is 2n=48 (Ojima et al., 1972; Liu, 1980). The specimens were held in an aerated aquarium at room temperature for 2–72 h before each experiment. Sixteen hours before killing, fish were injected with 5 μg of PHA per g of fish (stock solution, 0.5% PHA in 0.9% NaCl). Four hours before killing, fish were injected with 5 μg of PHA per g again and 5 μg of colchicine per g of fish (stock solution, 0.5% colchicine in 0.9% NaCl).
Chromosome preparation
Chromosomes were prepared from kidney tissue directly after killing. Weighed kidney tissue was removed and then minced in a clean Petri dish in citric acid solution (2% citric acid in distilled water). After washing with citric acid solution three times, the sample was transferred to a reaction tube filled with the same solution and homogenized with a Pasteur pipette. The cell suspension was removed into another reaction tube and centrifuged at 69 g for 10 min. The supernatant was discarded and the pellet resuspended in hypotonic solution (0.075 M KCl). Hypotonic treatment continued at 25°C for 40 min. One millilitre of fresh fixative (3:1 methanol–acetic acid) was then carefully added to the hypotonic solution, and the sample was centrifuged at 69 g for 10 min. The pellet was gently resuspended in fresh fixative for 30 min. Cells were centrifuged and resuspended twice more. Two drops of suspension were placed on a clean glass slide that was submerged in ice-cold distilled water until use. Slides were then immediately dried over a flame.
Chromosome banding
Giemsa stock solution was prepared routinely by dissolving 0.5 g of Giemsa powder stain (Shanghai Third Regent Factory) in 33 mL of glycerine in a warm water bath (at 56°C for 90 min) and then filtering after mixing with 33 mL of methyl alcohol.
Basic staining solution (Giemsa stock solution was freshly diluted to a 5% staining solution with 0.01 M sodium phosphate dibasic solution, pH 8.5) and neutral staining solution (Giemsa stock solution was freshly diluted to a 5% staining solution with 0.01 M sodium phosphate buffer, pH 6.8) were used. Slides were stained with neutral staining solution for 30 min, then digested with 0.1% trypsin solution for 15 s at room temperature and restained with basic staining solution for 10 min.
Analysis of banding patterns
Complete metaphases of satisfactory quality and some late prophases were photographed with an Olympus BH-2 photomicroscope. Five complete metaphases from each of the six fish were used for detailed banding pattern analysis. Karyotypes were made of each mid-metaphase and then compared with each other to identify corresponding chromosomes.
Results
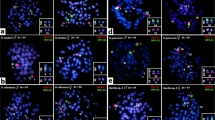
When slides were stained with neutral staining solution for 30 min first, then digested with trypsin and restained with basic staining solution for about 10 min, late prophase, early metaphase, middle metaphase and late metaphase chromosomes all showed defined banding patterns (Fig. 1). However, very poor banding was obtained with metaphase chromosomes that were digested with trypsin before being Giemsa stained. At late prophase, the centromeres of the chromosomes could not be identified unequivocally, and the chromosomes could not easily be spread well. At late metaphase, the chromosomes were too short and the bands became fewer in number, so that only the early and middle metaphase chromosomes were suitable for further analysis of the banding pattern.
Comparison of corresponding chromosomes, from different metaphases in Fig. 2, revealed that their banding patterns and numbers of bands were highly reproducible and very specific. Each individual chromosome was identified unequivocally by its banding pattern and number of bands. No difference in banding patterns was observed between the male and female specimens.
The complete metaphase banding pattern of grass carp was constructed (Fig. 2). The metaphase chromosomes of the fish were classified into two types according to the positions of their centromeres. The metacentric group includes nine pairs of chromosomes. The submetacentric group includes all the other 15 pairs. Table 1 shows a regional analysis of band number. The total number of bands, including dark and light, was 217. Because the chromosomes of grass carp are short, regional analysis of band number was classified by short and long arms.
Discussion
The field of fish cytogenetics is still poorly developed. This is largely because of the difficulty in obtaining good-quality multiple structural banding patterns from the chromosomes of fish (Wiberg 1983; Medranc et al., 1988; Schmid & Guttenbach, 1988; Bernardi, 1989; Hellmer et al., 1991). To overcome this difficulty, a new compound structural banding technique, different from the G-banding used on chromosomes of warm-blooded vertebrates, was used in this study. When metaphase chromosomes of fish were stained with neutral Giemsa staining solution first, then digested with trypsin and restained with basic staining solution, every middle metaphase chromosome showed highly reproducible structural bands. The banding pattern was chromosome-specific, and corresponding chromosomes in different middle metaphase cells could be identified unequivocally. These results indicate strongly that good-quality structural banding patterns can be obtained from middle metaphase chromosomes in fish. Indeed, similar results in other fish have been obtained with the same banding procedure (C. Luo, unpubl. obs.).
It should be stressed that, in this study, only when proteolytic enzyme digestion was performed after the metaphase was Giemsa stained could good quality structural banding patterns be obtained. This observation means that the structure of the fish chromosome for banding is very sensitive to trypsin; the Giemsa stain binding to the chromosome can selectively protect some of the structure of the chromosome from digestion in some way. The detailed process of protection is not yet clear and needs further study.
Compared with the G-banding patterns of warm-blooded animals, the dark bands shown in this study more closely resemble bead-like thickenings. Thus, it is reasonable to suggest that the present compound banding method is not mainly dependent on the compositional compartmentalization of the genome but the structure that is created by chromomeres concentrating at certain regions. Nevertheless, the banding pattern cannot be simply classified into a chromomere map, because the banding pattern of satisfactory quality was obtained only after proteolytic enzyme digestion and basic Giemsa staining solution restaining. It may be appropriate for the chromosome bands produced in this study to be named ‘chromomere-dependent G-bands’.
References
Almeida Toledo, L. F., Viegas-Pequinot, E., Foresti, F., Toledo Filho, S. A. and Dutrillaux, B. (1988). BrdU replication patterns demonstrating chromosome homoeologies in two fish species of the genus Eigenmannia. Cytogenet Cell Genet, 48: 117–120.
Bernardi, G. (1989). The isochore organization of the human genome. Ann Rev Genet, 23: 637–661.
Bernardi, G., Olofsson, B., Filipski, J., Zerial, M., Salinas, J., Cuny, G. et al. (1985). The mosaic genome of warm-blooded vertebrates. Science, 228: 953–958.
Burdick, A. B., Libbus, B. L., Athanassiou, V. and Falek, A. (1974). Mammalian male pachytene chromosome interconnections and karyotypes. Can J Genet Cytol, 16: 481–489.
Calza, R. E., Eckhardt, L. A., Delgiudice, T. and Schildkraut, C. L. (1984). Changes in gene position are accompanied by a change in time of replication. Cell, 36: 689–696.
Comings, D. E. (1978). Mechanisms of chromosome banding and implications for chromosome structure. Ann Rev Genet, 12: 25–46.
Cuny, G., Soriano, P., Macaya, G. and Bernardi, G. (1981). The major components of the mouse and human genomes: preparation, basic properties, and compositional heterogeneity. Eur J Biochem, 111: 227–233.
Delany, M. E. and Bloom, S. E. (1984). Replication banding patters in the chromosomes of the rainbow trout. J Hered, 75: 431–434.
De Torres, M. L. and Abrisqueta, J. A. (1978). Study of human male meiosis. II. Q banding in pachytene bivalents. Hum Genet, 42: 283–289.
Fang, J. S. and Jagiello, G. M. (1991). The chromomere map of the pachytene spermatocyte of the Turkish hamster (Mesocricetus brandti). Genome, 34: 626–630.
Ford, L., Cacheiro, N., Norby, D. and Heller, C. G. (1968). Identification of normal human pachytene chromosomes. I. The normal human karyotype. Nucleus, 11: 83–95.
Furst, A., Brown, E. H., Braunstein, J. D. and Schildkraut, C. L. (1981). α-Globin sequences are located in a region of early replication DNA in murine erythroleukemia cells. Proc Natl Acad Sci USA, 78: 1023–1027.
Galetti, P. M. Jr and Foresti, F. (1986). Evolution of the ZZ/ZW system in Leporinus (Pisces, Anastomidae). Cytogenet Cell Genet, 43: 434–446.
Giles, V., Thode, G. and Alvarez, M. C. (1988). Early replication bands in two Scorpion fishes, Scopaena porcus and S. notata (Order Scorpaneiformes). Cytogenet Cell Genet, 47: 80–83.
Goldman, M. A., Holmquist, G. P., Gray, M. C., Caston, L. A. and Nay, A. (1984). Replication timing of genes and middle repetitive sequences. Science, 224: 686–692.
Hellmer, A., Voiculescu, I. and Schempp, W. (1991). Replication banding studies in two cyprinid fishes. Chromosoma, 100: 524–531.
Holmquist, G., Gray, M., Porter, T. and Jordan, J. (1982). Characterization of Giemsa dark- and light-band DNA. Cell, 31: 121–129.
Hudson, A. P., Cuny, G., Cortadas, J., Haschemeyer, A. E. V. and Bernardi, G. (1980). An analysis of fish genomes by density gradient centrifugation. Eur J Biochem, 112: 203–210.
Jagiello, G. M. and Fang, J. S. (1982). Complete autosomal chromomere maps of human early and mid/late pachytene spermatocytes. Am J Hum Genet, 34: 112–124.
Kligerman, A. D. and Bloom, S. E. (1977). Rapid chromosome preparations from solid tissues of fishes. J Fish Res Board Can, 34: 266–269.
Liu, L. Y. (1980). On karyotypes of the grass carp, Ctenopharyngodon idellus. Acta Zool Sinica, 26: 126–131.
Luciani, J. M., Morazzani, M.-R. and Stahl, A. (1975). Identification of pachytene bivalents in human male meiosis using G-banding technique. Chromosoma, 52: 275–282.
Mayr, B., Kalat, M., Ráb, P. and Lambrou, M. (1987). Band karyotypes and specific types of heterochromatins in several species of European percid fishes (Percidea, Pisces). Genetica, 75: 199–205.
Medranc, L., Bernardi, G., Couturier, J., Dutrillaux, B. and Bernardi, G. (1988). Chromosome banding and genome compartmentalization in fishes. Chromosoma, 96: 178–183.
Ojima, Y., Hayashi, M. and Veno, K. (1972). Cytogenetic studies in lower vertebrates. X. Karyotypes and DNA studies in 15 species of Japanese Cyprinidae. Jap J Genet, 47: 431–440.
Pizon, V., Cuny, G. and Bernardi, G. (1984). Nucleotide sequence organisation in the very small genome of a tetradontid fish. Eur J Biochem, 140: 25–30.
Schmid, M. and Guttenbach, M. (1988). Evolutionary diversity of reverse (R) fluorescent chromosome bands in vertebrates. Chromosoma, 97: 101–114.
Schweizer, D. (1976). Reverse fluorescent chromosome banding with chromomycin and DAPI. Chromosoma, 58: 307–324.
Thiery, J. P., Macaya, G. and Bernardi, G. (1976). An analysis of eukaryotic genomes by density gradient centrifugation. J Mol Biol, 108: 219–235.
Wiberg, U. H. (1983). Sex determination in the European eel (Anguilla anguilla L.). Cytogenet Cell Genet, 36: 589–598.
Zenzes, M. T. and Voiculescu, I. (1975). C-Banding patterns in Salmo trutta a species of tetraploid origin. Genetica,, 45: 531–536.
Acknowledgements
I thank Z. Z. Yao for his help with the experiment.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Luo, C. Multiple chromosomal banding in grass carp, Ctenopharyngodon idellus. Heredity 81, 481–485 (1998). https://doi.org/10.1046/j.1365-2540.1998.00323.x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1046/j.1365-2540.1998.00323.x