Abstract
Ecological genomics is the study of adaptation of natural populations to their environment, and therefore seeks to link organism and population level processes through an understanding of genome organization and function. The planktonic microcrustacean Daphnia, which has long been an important system for ecology, is now being used as a genomic model as well. Here we review recent progress in selected areas of Daphnia genomics research. Production of parthenogenetic male offspring occurs through environmental cues, which clearly involves endocrine regulation and has also been studied as a toxicological response to juvenoid hormone analog insecticides. Recent progress has uncovered a putative juvenoid cis-response element, which together with microarray analysis will stimulate further research into nuclear hormone receptors and their associated transcriptional regulatory networks. Ecotoxicological studies indicate that mRNA profiling is a sensitive and specific research tool with promising applications in environmental monitoring and for uncovering conserved cellular processes. Rapid progress is expected to continue in these and other areas, as genomic tools for Daphnia become widely available to investigators.
Similar content being viewed by others
Introduction
A meaningful integration of ecology and genomics promises important new insights into the process by which populations respond and adapt to their environment. This research begins by identifying the genetic basis for ecologically relevant traits. Then, explicit and quantifiable associations are made between the heritable molecular variation within populations and individual fitness across varying ecological conditions (Gibson, 2002; Feder and Mitchell-Olds, 2003). In practice, two approaches are taken to achieve this level of integration; each presents a different set of advantages and challenges. First, ecological and evolutionary studies are pursued using traditional genetic model species that have well developed genomic tools, such as Drosophila, Caenorhabditis, Arabidopsis and Mus (cf. Landry et al., 2006; Shimizu and Purugganan, 2006). Although these species offer a staggering amount of molecular biological knowledge, many have ill-defined population boundaries and ecologies. This situation is starting to change owing to increased ecological study of these model organisms. Second, molecular genetic and genomic studies are initiated using organisms that have well-understood ecologies and evolutionary histories (Crawford, 2001; Strauss and Martin, 2004; Lai et al., 2006), of which the water flea, Daphnia, is one. Here, we outline the progress in the use of Daphnia as a model in evolutionary and ecological functional genomics.
Several attributes make Daphnia an especially tractable system for ecological genomics. In addition to their well-understood ecology and their sentinel roles within freshwater ecosystems, these microcrustaceans are easily collected in large numbers and reared in the lab with a 5- to 10-day generation time. They have a relatively small and sequenced genome (ca. 200 Mb) and perhaps most interesting, they reproduce by cyclical parthenogenesis. Females typically reproduce asexually, but environmental conditions can induce parthenogenetic production of males (genetically identical to their mothers) and haploid gametes that are fertilized and enter a state of extended metabolic dormancy called diapause. This life cycle allows the creation of inbred or outbred lines for mapping studies, and provides tremendous power for dissecting genetically based differences (among clones) from experimental or developmental noise by measuring replicates within a clone.
The ecology and evolution of Daphnia is well explored with respect to habitat differentiation (Fryer, 1991; Wellborn et al., 1996; Colbourne et al., 1997), but progress has been less in characterizing molecular mechanisms responsible for traits conferring a fitness advantage in defined environments. For instance, diurnal and seasonal variations in predation, parasitism, food availability and temperature interact in complex ways to produce phenotypes such as investment in sexual reproduction (Cáceres and Tessier, 2004). Although production of sexual eggs and males is expected to be coupled at some level as an adaptation to make diapausing embryos, mechanisms of this induction by environmental cues are not well understood. An important first step towards such an investigation is to examine the genetics of environmental sex determination. Similarly, Daphnia are often in temperature- and oxygen-stratified habitats, which has important ecological implications for predator avoidance and access to alternative grazing resources (Williamson et al., 1996). Acclimation to hypoxia via hemoglobin (Hb) synthesis has been a subject of extensive study over the last decades. Here, we discuss new discoveries enabled by genomic approaches that provide a mechanistic linkage between two well-studied phenomena, male production and Hb induction in Daphnia.
A consortium of investigators is progressing rapidly in creating tools for Daphnia genomics. First, the genome sequencing and assembly of a D. pulex genome by the Department of Energy's Joint Genome Institute is complete, and its annotation has begun. The annotation goals of identifying genes and their structures are being guided by a growing number of cDNA libraries and sequences for both D. pulex (Colbourne et al., submitted) and D. magna (Watanabe et al., 2005). Second, microsatellite DNA markers are deployed for population surveys (Fox, 2004) and quantitative trait locus (QTL) mapping experiments (Colbourne et al., 2004). Many of these markers form the basis of a genetic map (Cristescu et al., 2006) that is being integrated along with large insert BAC libraries to identify chromosomal segments under high and low rates of recombination. Third, both cDNA and oligonucleotide microarrays are being used for gene expression analysis in D. magna and D. pulex. Finally, bioinformatic efforts have resulted in the creation of wFleabase (Colbourne et al., 2005), an online repository of genetic, molecular and genomic data for Daphnia, while laboratory projects to create cell lines and virally induced transgenic lines are also ongoing (Robinson et al., 2006).
These resources are opening the way for the systematic investigation of the genes and pathways underlying ecologically relevant traits. Presentations at a recent meeting of the Daphnia Genomics Consortium summarized progress in the areas of gametogenesis, embryonic patterning, parasitology, toxicology and molecular evolution (http://conferences.cgb.indiana.edu/daphnia2006/). To date, much of the data obtained about molecular mechanisms in the area of ecological genomics stem from physiological rather than evolutionary perspectives, which is reflected in our choice of literature to review. Specifically, we focus on progress in understanding sex determination and environmental stress responses.
Male production pathway as an entry into ecological genomics
The ecological basis of sex determination in Daphnia has long been appreciated (Banta and Brown, 1939) and genetic variation for this trait is substantial (Yampolsky, 1992). Environmental factors such as temperature, photoperiod and crowding stimulate male production by parthenogenesis (Hobæk and Larsson, 1990) and, although varying investments into sexual reproduction have important ecological and evolutionary consequences (Tessier and Cáceres, 2004), molecular studies have only begun. Male production is a promising research avenue for several reasons. First, it provides an opportunity to study the molecular basis of environmental sex determination. Second, it is a basis for broader exploration of invertebrate endocrine function and disruption in general, which has lagged far behind our understanding of vertebrate systems (Oetken et al., 2004). Third, as a classic example of phenotypic plasticity and epigenetic regulation, it offers the possibility of uncovering molecular mechanisms responsible for these ecologically and evolutionarily central mechanisms. Fourth, evidence indicates important couplings between juvenile hormone (JH) pathways and ecdysone pathways in Daphnia (Mu and LeBlanc, 2004), which are likely to be useful for genetic and genomic analyses of these pathways across the Ecdysozoa.
Olmstead and LeBlanc (2002) were first to demonstrate production of male broods of D. magna in response to the sesquiterpenoid hormone methyl farnesoate (MF), the unepoxidated form of insect JH III. Discovery of MF in crustaceans prompted extensive efforts to characterize the functions of this hormone, which largely appears to mirror the effects of JH in insects (Laufer and Biggers, 2001). Other JH analogs (JHAs) in addition to MF have been shown to induce male production in Daphnia. The insecticides pyriproxyfen, fenoxycarb and methoprene cause male broods (Tatarazako et al., 2003), with orders of magnitude greater potency than MF for pyriproxyfen and fenoxycarb. Interestingly, all of these compounds also induce Hb production in Daphnia (Rider et al., 2005). Therefore, male production and Hb induction appear to be coupled at some level by JHA signaling (see below).
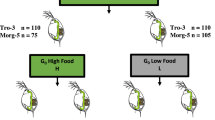
Studies with MF demonstrate that D. magna and D. pulex produce male broods in a dose-dependent manner, with concentrations above 300 nM resulting in all male broods (Olmstead and LeBlanc, 2002; personal observation). Furthermore, the sensitive period for MF induction of male development is 1–2 days before egg laying during the vitellogenic period of oogenesis. We have studied the transcriptional effects on genes from developmentally staged females exposed to MF, by using a microarray platform described elsewhere (Colbourne et al., submitted). Briefly, females from a single mother were kept in a common garden until their first broods were released. Then, 20 individuals were randomly selected for a 3-day exposure to either 400 nM MF (Echelon Biosciences, Salt Lake City, UT, USA, changed daily) or to methanol carrier (0.004%). Three replicates of each condition were used for RNA isolation, cDNA labeling and hybridization, and analysis according to standard protocols (http://dgrc.cgb.indiana.edu/microarrays/support/protocols.html).
Compared to controls, MF-treated animals showed higher steady-state levels of 39 unique transcripts and lower levels of 16 transcripts (Table 1). Twenty-two differentially regulated genes shared significant sequence homologies with proteins archived in the NCBI non-redundant database. In particular, Hb and neuronal acetylcholine receptor transcripts were elevated under MF treatment, in addition to arginine kinase, amylase, cytochrome c oxidase and cytochrome b, several cuticle proteins, actin and a putative ribosomal biogenesis regulatory protein. Control animals had higher levels of glucosamine-6-phosphate deaminase, glucose-6-phosphatase, several proteases, an actin de-polymerizing enzyme (profilin), a putative receptor of unknown function and a homolog of the Rheb G-protein-coupled receptor. Another transcript with lower levels during treatment had putative homology to an ecdysteroid-regulated transcript from Manduca Sexta (GenBank accession AAB08704.1). Several other genes differed in response to MF, but whose putative homologs in arthropods (such as the Drosophila melanogaster genes CG6770 and CG31997) have no known function. Subsequent study of these genes may reveal whether they have conserved roles in JH response. Several dozen genes with no sequence similarity to entries in the non-redundant protein database showed differential expression as well.
Taken together, results from this microarray experiment are consistent with current data for MF activity in crustaceans and also offer new avenues for exploration (Figure 1). First, higher steady-state levels of Hb are consistent with transcriptional control of this protein, an area of active investigation explored further below. Second, changes in polysaccharide metabolism-related genes, cuticle genes and cytoskeletal protein genes imply a coordinated shift in carbohydrate usage away from ATP production and towards chitin metabolism. Physiological and biochemical studies would be useful to uncover such systemic changes in metabolism in response to JHA treatment. Furthermore, alteration of chitin metabolism underscores connections between MF and ecdysis due to genetic crosstalk with ecdysone pathways (Mu and LeBlanc, 2004). Third, involvement of the rheb receptor in response to MF is, to our knowledge, a novel observation linking this protein to a crustacean hormonal response. In D. melanogaster, Rheb participates in cell growth and proliferation via the S/G1 mitotic transition, in addition to starvation response (Patel et al., 2003). These biological functions in flies are consistent with a role for Rheb in Daphnia related to molting. Fourth, an intriguing and under-explored connection between secretion of JHA and cholinergic regulation is highlighted by increased acetylcholine receptor mRNA levels under MF treatment. Research into molecular mechanisms of cholinergic hormonal control is only beginning to be investigated in arthropods (Kuo, 2002). Genomic tools available in Daphnia may help to inform research in suitable biochemical and physiological crustacean models of both rheb and cholinergic pathway interactions with JHA. Finally, genes of unknown function that are expressed constitutively in response to MF may be members of the MF pathway, and could be interesting targets of subsequent studies such as time course, dose–response and exposure to other JHA.
Schematic depicting the methyl farnesoate pathway in male production and hemoglobin induction. For further discussion, see Laufer and Biggers (2001) and Mu and LeBlanc (2004). Arrows (induction or production) and blocked lines (repression) may be direct or indirect effects (possibly both, indicated by dashed lines). Sequence and expression data have been obtained from cDNAs encoding farnesoic acid methyltransferase, two CHH peptides and homologs of D. melanogaster sex determination genes sex lethal (Sxl) and transformer-2 (Tra-2) (data not shown).
The MF pathway is an early example of genomic studies investigating the mechanism for environmental sex determination in natural settings. Current work in our laboratories includes additional microarray experiments using both wild-type and naturally occurring isolates that never produce males and are thus unresponsive to MF. Mapping panels from crosses between male and non-male producers will help identify QTL for this trait, which could subsequently be used to study ecological correlates of genetic polymorphisms.
Genomics of environmental stress responses
Research in Daphnia has long focused on the environmental stresses they encounter and mechanisms they use to counter or mitigate them. Prominent among these areas are toxicology, hypoxia, temperature, parasites, predators and ultraviolet radiation. Responses to these challenges have been documented at the level of physiology, genetics, maternal effects, morphology, behavior and life history. Adequately addressing these areas is beyond the scope of this review; our aim instead is to highlight recent progress in uncovering molecular mechanisms underlying these changes.
Acute hypoxia in Daphnia leads to compensation via increased heart rate or ventilation rate (Pirow et al., 1999), but chronic hypoxia leads to induction of Hb synthesis. This synthesis is subunit-specific (Kimura et al., 1999), with changes over time occurring at subunit-specific rates (Zeis et al., 2003). Levels of mRNA for subunits hb2 and hb3 are rapidly and highly induced during hypoxia, while hb1 is unaffected (Zeis et al., 2003). These results, plus the identification of putative binding sites for the mammalian hypoxia-inducible factor 1 (HIF-1; Kimura et al., 1999), led to a detailed investigation of Hb gene promoters. Induction of Hb gene expression during hypoxia is dependent on the binding of HIF to particular hypoxia response elements (HRE) in promoter regions of the four known D. magna Hb genes (Gorr et al., 2004). Three HREs upstream of the globin-2 gene were found using heterologous transfection of HIF-expressing human and D. melanogaster cells, showing specific binding of HIF complexes at two sites in vitro and binding of an unknown constitutive transcription factor at the third site (Gorr et al., 2004). A recent report (Gorr et al., 2006) has identified a putative juvenoid response element (JRE) in the promoter of hb2 in D. magna that binds an activated factor in response to JHA. This development is exciting for several reasons. Despite decades of intense interest and research into mechanisms of JHA action in arthropods, a nuclear hormone receptor for JHA has not been identified with certainty. The JRE is a powerful tool to establish the identity of the JHA receptor, which will undoubtedly stimulate molecular genetic research into pathways of JHA activity. These findings also link mechanisms for the ecologically relevant traits of male production and Hb induction for the first time.
The recent discovery of two additional members of the D. magna globin family using Southern blotting (Nunes et al., 2005) indicates that many more discoveries will emerge from functional studies of these molecules. Studies of Daphnia populations that are locally adapted to oxygen-rich or oxygen-starved habitats may help elucidate the ecological implications of lower mass-specific oxygen consumption rates in hypoxia-acclimated animals (Seidl et al., 2005) for community structure and trophic interactions. Furthermore, increased O2 delivery due to higher Hb content incurs physiological and fitness costs via increased protein synthesis and predation (Pirow et al., 2001), providing an opportunity to link physiological ecology with molecular mechanisms such as HIF binding.
In toxicogenomics, research into endocrine disrupting chemicals such as the JHA compounds discussed above continues to be a high priority (Oetken et al., 2004). In this regard, Daphnia will be useful for elucidating gene expression patterns and associated ecological changes in response to these challenges. The difficulty and potential of combining a genomics approach with environmental monitoring has been a subject of intense interest (for example, Lettieri, 2006). Some conceptual issues are as follows: (1) How specific are mechanisms of toxicity or clearance, as revealed by the genes and pathways responding to toxicants? (2) How sensitive and useful is expression profiling at uncovering those genes or pathways? (3) Are traditional toxicological approaches (for example, end points, LC50s, dose–response curves) optimal for genomic assays? (4) Are genomic responses to toxicants indicative of changes within a population and ultimately the state of its ecosystem? Data from groups performing toxicological genomics in Daphnia, although preliminary, bear directly on some of these questions. Future studies using more complete microarray platforms with tens of thousands instead of thousands of probes will provide interesting insight into variation among toxicological conditions. Yet, the trends described here are universal enough to provide a meaningful picture.
First, while ‘stress response’ genes or pathways tend to be induced consistently across physiological challenges as diverse as metal toxicity, ordinance (explosive) contamination or nanoparticle exposure, each challenge also provides a unique signature. These observations imply that certain types of molecular response, such as induction of a particular class of chaperones, may be diagnostic of organisms under stress. Also, condition-specific responses are likely, depending on experimental protocols (for example, exposure times and dosages) and the life stages of the animals. Second, low (sub-lethal and environmentally relevant) chronic exposures generally can induce measurable changes in gene expression. Low-dose exposures, along with exposure to simultaneous challenges, are often used as experimental tools for discerning potential mechanisms of toxicity (Lettieri, 2006). Therefore, transcriptional profiling should be useful, in combination with data from other sources, to uncover how toxins work. Beyond that, how genetic background of particular clones might affect outcomes (see Lopes et al., 2006), the importance of trans-generational (maternal or grand-maternal) effects, and how sequence, gene expression and/or physiological response vary are unanswered questions.
A recent study of the effect of the fungicide propiconazole on D. magna (Soetaert et al., 2006) illustrates the challenges and potential of environmental monitoring using microarrays. Propiconazole resembles other chemicals studied for toxicity and their effects on development (Kast-Hutcheson et al., 2001), such as diethylstilbestrol (Baldwin et al., 1995) and nonylphenol (Shurin and Dodson, 1997), which kill or deform embryos, often at concentrations similar to those in the environment. Transcriptional profiling of propiconazole treatment of 4- and 8-day-old animals using microarrays constructed from age-specific cDNA libraries established differential expression of over 10% of the arrayed genes (Soetaert et al., 2006). Gene expression varied with both time and dose, which underscores the difficulty of choosing appropriate end points for such studies. However, results are encouraging for a research community poised to apply microarrays to discover the condition dependency of responses to toxicants. For example, transcripts related to development and reproduction were differentially expressed, such as an eightfold decrease in transcript levels of the primary yolk protein vitellogenin. Also, some transcripts either had homologs of unknown function in other organisms or appeared unique to Daphnia. Profiling gene expression across a range of environmentally relevant conditions should elucidate cellular and organism level processes that are vital within natural settings, but that are not typically elicited in laboratory studies of traditional model organisms.
It would be interesting to extend the above observations by profiling offspring from mothers given a propiconazole exposure, which affects embryonic development in Daphnia (Kast-Hutcheson et al., 2001). The quality of the maternal environment influences traits such as size at birth and number of offspring (Sakwinska, 2004), strain-specific immunity to bacterial parasites (Little et al., 2003) and energy allocation to offspring (LaMontagne and McCauley, 2001). Interestingly, mothers reproducing in poor environments produced offspring more than twice as resistant to bacterial infection as mothers in favorable environments (Mitchell and Read, 2005). These findings underscore an important requirement for systematic approaches to quantify sources and consequences of both genetic and environmental variability in ecological studies (Forbes, 1998).
Maternal effects have been experimentally addressed during exposure to predator kairomones, some effects of which occur across generations. The formation of defensive structures like helmets and neckteeth in response to kairomones is a classic example of cyclomorphosis, and depends upon maternal environment to an even greater extent than in the late embryonic environment (Agrawal et al., 1999). A study of heat-shock protein (HSP) induction and the actin and tubulin cytoskeleton in predator-exposed Daphnia (Pijanowska and Kloc, 2004) demonstrated particular maternal effects from kairomone exposure. Upon exposure to the invertebrate predator Chaoborus or to fish kairomones, HSP40, HSP60 and HSP70 all decreased in expression. Using anti-phosphohistone 3 as a control also showed that α-tubulin and actin protein levels decreased in exposed animals, and that tubulin expression decreased in F1 progeny of exposed animals compared to F1 from control animals. Interestingly, Cu and Zn both reduced neckteeth induction in Daphnia neonates exposed to predator kairomones (Hunter and Pyle, 2004), indicating interference by metals in the cyclomorphosis signaling pathway. Employing combinations of different environmental stressors has considerable promise for dissecting signaling pathways and molecular responses to these challenges.
A powerful tool for microevolutionary studies is the hatching of dormant diapause embryos from sediment core samples of ponds and lakes, dubbed ‘resurrection ecology’ (Kerfoot and Weider, 2004). Implications of migration through time, or the effects of overlapping generations on population dynamics, are an active research area (for example, Cáceres and Tessier, 2004). In addition, hatching dormant propagules may compare ancestral populations and their descendents, for example before and after introduction of a novel predator or toxicant, to measure genetic differences related to the environmental change (Weider et al., 1997; Hairston et al., 1999). For instance, a Daphnia population exposed to variable and well-documented fish predation over 30 years was examined for changes in an adaptive quantitative trait (phototactic behavior, related to diel vertical migration and thus predator avoidance) and in neutral genetic markers (Cousyn et al., 2001). Results demonstrated increased behavioral plasticity in response to predator kairomones, and order of magnitude higher genetic differentiation for the trait than for neutral markers, consistent with a strong role for selection as the driver of rapid, directional changes. Another example demonstrated evolution of resistance to toxic food sources (cyanobacteria) through a reduction in phenotypic plasticity over time (Hairston et al., 2001). In a similar way, combining molecular biology and genomics with paleolimnology may test hypotheses about adaptation in populations exposed to severe metal contamination (Pollard et al., 2003). Monitoring changes in allele frequencies within naive and post-exposed populations is possible even with unhatched or gravely degraded embryos (Limburg and Weider, 2002); the greatest limitation, until now, was determining the sequence identity of candidate genes that have possibly evolved structurally in response to the documented changes in the environment.
Understanding genome structure in light of the stress of ecological competition (for example, for limited carbon or phosphorus) is a primary motivation for research into the functional consequences of rDNA length and copy number, termed biological stoichiometry (Weider et al., 2005). Interest in rDNA per se stems from mechanistic links between copy number and intergenic spacer (IGS) length, and their effects on organismal growth rate and phosphate requirements (Weider et al., 2005). The growth rate hypothesis (GRH) posits that N:P and C:P ratios reflect allocation of P to ribosomal RNA, with higher rates demanding more P for rRNA production and protein synthesis (Elser et al., 2000). Predictions based on the GRH have been supported by numerous studies (Weider et al., 2005), and environmental P concentrations are frequently low enough to limit growth of Daphnia (Anderson and Hessen, 2005). It is not clear whether stressful conditions due to poor food quality (low P content) are sufficient to drive the evolution of rDNA repeat length or copy number. However, evidence from laboratory experiments indicates that this may be the case. For example, by selecting for divergent weight-specific fecundity in D. pulex, Gorokhova et al. (2002) were able to demonstrate significant changes in age of first reproduction, size of first clutch, RNA and phosphorus content and IGS length after only five generations.
Conclusion
The growing genomics toolbox for Daphnia is providing opportunities to extend laboratory research from the molecular basis of morphological, physiological and behavioral differences to questions addressing the degree and importance of genetic variation within natural populations. Although Daphnia genomics research is still nascent, understanding the function of key genetic networks modulating traits with clear ecological relevance is advancing. For instance, microarray experiments and sequence analysis are uncovering members of these networks with characterized functions, plus genes that are apparently conserved across arthropods, but have no known function. Other genes are presumably unique in Daphnia. Indeed, at the recent Daphnia Genomics Consortium meeting, we presented data suggesting that at least 57% of over 12 600 cDNA sequences for D. pulex showed no similarity to protein sequences archived within the NCBI non-redundant database.
The structure and function of a genome is a product of the genome's environment, with interactions occurring at multiple scales. Current work – on genes linked within asexual genomes, on genes locked within a co-evolutionary arms race against parasites, and on multiple gene families whose copy numbers may be tied to ecological processes – is paving the way for genome-wide searches for signatures of selection on genomic elements. For example, the attributes of species are important in the macro-evolutionary processes acting on genome composition. These signatures range from adaptive amino-acid substitutions in genes associated with differences between humans and chimpanzees (Clark et al., 2003) to the distribution of elements such as introns and transposable elements as a function of long-term effective population size differences among some major divisions of life (Lynch and Conery, 2003). In a complementary way, research on populations of Daphnia will provide an important micro-evolutionary perspective. This view comes from studying genomic variation at local geographic scales and during recent demographic history, such as the latest demonstration that obligately asexual Daphnia lineages suffer increased deleterious mutations compared to sexual lineages (Paland and Lynch, 2006), which has clear evolutionary implications.
Accession codes
References
Agrawal AA, Larorsch C, Tollrian R (1999). Transgenerational induction of defences in animals and plants. Nature 401: 60–63.
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990). Basic local alignment search tool. J Mol Biol 215: 403–410.
Anderson TR, Hessen DO (2005). Threshold elemental ratios for carbon versus phosphorus limitation in Daphnia. Freshwater Biol 50: 2063–2075.
Baldwin WS, Milam DL, LeBlanc GA (1995). Physiological and biochemical perturbation in Daphnia magna following exposure to the model environmental estrogen diethylstilbestrol. Environ Toxicol Chem 14: 945–952.
Banta AM, Brown LA (1939). Control of male and sexual egg production. In: Banta AM (ed). Studies on the Physiology, Genetics, and Evolution of some Cladocera. Carnegie Institute: Washington, DC, USA. pp 106–129.
Cáceres CE, Tessier AM (2004). Incidence of diapause varies among populations of Daphnia pulicaria. Oecologia 141: 425–431.
Clark AG, Glanowski S, Nielsen R, Thomas PD, Kejariwal An, Todd MA et al. (2003). Inferring non-neutral evolution from human-chimp-mouse orthologous gene trios. Science 302: 1960–1963.
Colbourne JK, Hebert PDN, Taylor DJ (1997). Evolutionary origins of phenotypic diversity in Daphnia. In: Givnish TJ, Sytsma KJ (eds). Molecular Evolution and Adaptive Radiation. Cambridge University Press: Cambridge, UK. pp 163–188.
Colbourne JK, Robison B, Bogart K, Lynch M (2004). Five hundred and twenty eight microsatellite markers for ecological genomic investigations using Daphnia. Mol Ecol Notes 4: 485–490.
Colbourne JK, Singan VR, Gilbert DG (2005). wFleabase: the Daphnia genome database. BMC Bioinformatics 6: 45.
Cousyn C, De Meester L, Colbourne JK, Brendonck L, Vershuren D, Volckaert F (2001). Rapid, local adaptation of zooplankton behavior to changes in predation pressure in the absence of neutral genetic changes. Proc Nat Acad Sci USA 98: 6256–6260.
Crawford DL (2001). Functional genomics does not have to be limited to a few select organisms. Genome Biol 2: interactions 1001.1–1001.2.
Cristescu MEA, Colbourne JK, Radivojac J, Lynch M (2006). A microsatellite-based genetic linkage map of the waterflea, Daphnia pulex: on the prospect of crustacean genomics. Genomics 88: 415–430.
Elser JJ, Sterner RW, Gorokhova E, Fagan WF, Markow TA, Cotner JB et al. (2000). Biological stoichiometry from genes to ecosystems. Ecol Lett 3: 540–550.
Feder ME, Mitchell-Olds T (2003). Evolutionary and ecological functional genomics. Nat Rev Genet 4: 649–655.
Forbes VE (1998). Sources and implications of variability in sensitivity to chemicals for ecotoxicological risk assessment. Arch Toxicol Suppl 20: 407–418.
Fox JA (2004). New microsatellite markers for Daphnia galeata mendotae. Mol Ecol Notes 4: 544–546.
Fryer G (1991). Functional morphology and the adaptive radiation of the Daphniidae (Branchiopoda: Anomopoda). Philos Trans R Soc London B 331: 1–99.
Gibson G (2002). Microarrays in ecology and evolution: a preview. Mol Ecol 11: 17–24.
Gorokhova E, Dowling TE, Weider LJ, Crease TJ, Elser JJ (2002). Functional and ecological significance of rDNA intergenic spacer variation in a clonal organism under divergent selection for production rate. Proc R Soc London B 269: 2373–2379.
Gorr TA, Cahn JD, Yamagata H, Bunn HF (2004). Hypoxia-induced synthesis of hemoglobin in the crustacean Daphnia magna is hypoxia-inducible factor-dependent. J Biol Chem 279: 36038–36047.
Gorr TA, Rider CV, Wang HY, Olmstead AW, LeBlanc GA (2006). A candidate juvenoid hormone receptor cis-element in the Daphnia magna hb2 hemoglobin gene promoter. Mol Cell Endocrinol 247: 91–102.
Hairston Jr NG, Holtmeier CL, Lampert W, Weider LJ, Post DM, Fischer JM et al. (2001). Natural selection for grazer resistance to toxic cyanobacteria: evolution of phenotypic plasticity? Evolution 55: 2203–2214.
Hairston Jr NG, Lampert W, Cáceres CE, Holtmeier CL, Weider LJ, Gaedke MU et al. (1999). Rapid evolution revealed by dormant eggs. Nature 401: 446.
Hobæk A, Larsson P (1990). Sex determination in Daphnia magna. Ecology 71: 2255–2268.
Hunter K, Pyle G (2004). Morphological response of Daphnia pulex to Chaoborus americanus kairomone in the presence and absence of metals. Envrion Toxicol Chem 23: 1311–1316.
Kast-Hutcheson K, Rider CE, LeBlanc GA (2001). The fungicide propiconazole interferes with embryonic development of the crustacean Daphnia magna. Environ Toxicol Chem 20: 502–509.
Kerfoot WC, Weider LJ (2004). Experimental paleoecology (resurrection ecology): chasing Van Valen's red queen hypothesis. Limnol Oceanogr 49: 1300–1316.
Kimura S, Tokishita S, Ohta T, Kobayashi M, Yamagata H (1999). Heterogeneity and differential expression under hypoxia of two-domain hemoglobin chains in the water flea, Daphnia magna. J Biol Chem 274: 10649–10653.
Kuo R (2002). Cholinergic regulation of the corpora allata in adult male Loreyi leafworm Mythimna loreyi. Arch Ins Biochem Physiol 49: 215–224.
Lai Z, Gross BL, Zou Y, Andrews J, Rieseberg L (2006). Microarray analysis reveal differential gene expression in hybrid sunflower species. Mol Ecol 15: 1213–1227.
LaMontagne JM, McCauley E (2001). Maternal effects in Daphnia: what mothers tell their offspring and do they listen. Ecol Lett 4: 64–71.
Landry CR, Townsend JP, Hartl DL, Cavalieri D (2006). Ecological and evolutionary genomics of Sacharomyces cerevisiae. Mol Ecol 15: 575–591.
Laufer H, Biggers WJ (2001). Unifying concepts learned from methyl farnesoate for invertebrate reproduction and post-embryonic development. Amer Zool 41: 442–457.
Lettieri T (2006). Recent applications of DNA microarray technology to toxicology and ecotoxicology. Environ Health Persp 114: 4–9.
Limburg PA, Weider LJ (2002). ‘Ancient’ DNA in a microcrustacean resting egg bank can serve as a palaeolimnological database. Proc R Soc London B 269: 281–287.
Little TJ, O’Connor B, Colegrave N, Watt K, Read AF (2003). Maternal transfer of strain-specific immunity in an invertebrate. Curr Biol 18: 489–492.
Lopes I, Baird DJ, Ribeiro R (2006). Genetic adaptation to metal stress by natural populations of Daphnia longispina. Ecotoxicol Environ Safe 63: 275–285.
Lynch M, Conery JS (2003). The origins of genome complexity. Science 302: 1401–1404.
Mitchell SE, Read AF (2005). Poor maternal environment enhances offspring disease resistance in an invertebrate. Proc Biol Sci 272: 2601–2607.
Mu X, LeBlanc GA (2004). Cross-communication between signaling pathways: juvenoid hormones modulate ecdysteroid activity in a crustacean. J Exp Zool 301A: 793–801.
Nunes F, Spiering D, Wolf M, Wendler A, Pirow R, Paul RJ (2005). Sequencing of hemoglobin gene 4 (dmhb4) and Southern blot analysis provide evidence of more than four members of the Daphnia magna globin family. Biosci Biotechnol Biochem 69: 1193–1197.
Oetken M, Bachmann J, Schulte-Oehlmann U, Oehlmann J (2004). Evidence for endocrine disruption in invertebrates. Int Rev Cytol 236: 1–44.
Olmstead AW, LeBlanc GA (2002). Juvenoid hormone methyl farnesoate is a sex determinant in the crustacean Daphnia magna. J Exp Zool 293: 736–739.
Paland S, Lynch M (2006). Transitions to asexuality result in excess amino acid substitutions. Science 311: 990–992.
Patel PH, Thapar N, Guo L, Martinez M, Maris J, Gau C-L et al. (2003). Drosophila Rheb GTPase is required for cell cycle progression and cell growth. J Cell Sci 116: 3601–3610.
Pijanowska J, Kloc M (2004). Daphnia response to predation threat involves heat-shock proteins and the actin and tubulin cytoskeleton. Genesis 38: 81–86.
Pirow R, Bäumer C, Paul RJ (2001). Benefits of hemoglobin in the cladoceran crustacean Daphnia magna. J Exp Biol 204: 3425–3441.
Pirow R, Wollinger F, Paul RJ (1999). Importance of the feeding current for oxygen uptake in the water flea Daphnia magna. J Exp Biol 202: 553–562.
Pollard HG, Colbourne JK, Keller W (2003). Reconstruction of centuries-old Daphnia communities in a lake recovering from acidification and metal contamination. Ambio 32: 214–218.
Rider CV, Gorr TA, Olmstead AW, Wasilak BA, LeBlanc GA (2005). Stress signaling: coregulation of hemoglobin and male sex determination through a terpenoid signaling pathway in a crustacean. J Exp Biol 208: 15–23.
Robinson CD, Lourido A, Whelan SP, Dudycha JL, Lynch M, Isern S (2006). Viral transgenesis of embryonic cell cultures from the freshwater microcrustacean Daphnia. J Exp Biol 305A: 62–67.
Sakwinska O (2004). Persistent maternal identity effects on life history traits in Daphnia. Oecologia 138: 379–386.
Seidl MD, Paul RJ, Pirow R (2005). Effects of hypoxia acclimation on morpho-physiological traits over three generations of Daphnia magna. J Exp Biol 208: 2165–2175.
Shimizu KK, Purugganan MD (2006). Evolutionary and ecological genomics of Arabidopsis. Plant Physiol 138: 578–584.
Shurin JB, Dodson SI (1997). Sublethal toxic effects of cyanobacteria and nonylphenol on environmental sex determination and development in Daphnia. Environ Toxicol Chem 16: 1269–1276.
Soetaert A, Moens LN, van der Ven K, van Leemput K, Naudts B, Blust R et al. (2006). Molecular impact of propiconazole on Daphnia magna using a reproduction-related cDNA array. Comp Biochem Physiol C 142: 66–76.
Strauss SH, Martin FM (2004). Poplar genomics comes of age. New Phytol 164: 1–4.
Tatarazako N, Oda S, Watanabe H, Morita M, Iguchi T (2003). Juvenile hormone agonists affect the occurrence of male Daphnia. Chemosphere 53: 827–833.
Tessier AJ, Cáceres CE (2004). Differentiation in sex investment by clones and populations of Daphnia. Ecol Lett 7: 695–703.
Watanabe H, Tatarazako N, Oda S, Nishide H, Uchiyama I, Morita M (2005). Analysis of expressed sequence tags of the water flea Daphnia magna. Genome 48: 606–609.
Weider LJ, Elser JJ, Crease TJ, Mateos M, Cotner JB (2005). The functional significance of ribosomal (r)DNA variation: impacts on the evolutionary ecology of organisms. Annu Rev Ecol Evol Syst 36: 219–242.
Weider LJ, Lampert W, Wessels M, Colbourne JK, Limburg P (1997). Long-term genetic shifts in microcrustacean egg bank associated with anthropogenic changes in Lake Constance ecosystem. Proc R Soc London B 264: 1613–1618.
Wellborn GA, Skelly DK, Werner EE (1996). Mechanisms creating community structure across a freshwater habitat gradient. Annu Rev Ecol Syst 27: 337–363.
Williamson CE, Sanders RW, Moeller RE, Stutzman PL (1996). Utilization of subsurface food resources for zooplankton reproduction: implications for diel vertical migration theory. Limnol Oceanogr 41: 224–233.
Yampolsky LY (1992). Genetic cariation in the sexual reproduction rate within a population of a cyclic parthenogen, Daphnia magna. Evolution 46: 833–837.
Zeis B, Becher B, Goldmann T, Clark R, Vollmer E, Bölke B et al. (2003). Differential haemoglobin gene expression in the crustacean Daphnia magna exposed to different oxygen partial pressures. Biol Chem 384: 1133–1145.
Acknowledgements
We apologize to those whose work could not be cited because of space limitations, and thank the editors for an opportunity to contribute to this special volume on Ecological Genomics. Our work is supported in part by National Science Foundation grants (EF-0328516, DEB-0221837) and by the Center for Genomics and Bioinformatics via the METACyt Initiative and the Indiana Genomics Initiative under the Lilly endowment. This research is also made possible by the Daphnia Genomics Consortium.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Eads, B., Andrews, J. & Colbourne, J. Ecological genomics in Daphnia: stress responses and environmental sex determination. Heredity 100, 184–190 (2008). https://doi.org/10.1038/sj.hdy.6800999
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.hdy.6800999
Keywords
This article is cited by
-
Transcriptome analysis of different life-history stages and screening of male-biased genes in Daphnia sinensis
BMC Genomics (2022)
-
Combined effects of the pesticide spinetoram and the cyanobacterium Microcystis on the water flea Daphnia pulex
Environmental Science and Pollution Research (2022)
-
Transcriptional responses of Daphnia magna exposed to Akaki river water
Environmental Monitoring and Assessment (2022)
-
Differential gene transcription across the life cycle in Daphnia magna using a new all genome custom-made microarray
BMC Genomics (2018)
-
A high-density genetic map reveals variation in recombination rate across the genome of Daphnia magna
BMC Genetics (2016)