Abstract
Major genetic breaks between the Indian and Pacific oceans have been reported for marine fishes and invertebrates. The genetic structure and history of the Indian scad mackerel, Decapterus russelli, in the Indo-Malay archipelago were investigated using the cytochrome b gene sequence as mitochondrial marker and two length-polymorphic introns as nuclear markers. The existence of two major mitochondrial lineages separated by 2.2% average nucleotide divergence, and their heterogeneous geographical distributions, were confirmed. This indicated past geographic isolation, possibly caused by a Pleistocene drop in sea level. The presence, in sympatry, of the two mitochondrial lineages was thought to result from secondary contact. A recent population bottleneck and subsequent rapid population expansion were indicated by low genetic diversities and strongly negative Tajima's D-values. This evidence supports the hypothesis that the habitat available to D. russelli in the Pleistocene was restricted. Taking into account both mitochondrial and nuclear-DNA data, three geographically distinct populations were identified: one sampled in the Makassar Strait and Sulawesi Sea, one in the Arafura Sea and the third from the entire western region of the Indo-Malay archipelago. Considering the high hydrological connectivity of this region of the Indo-Pacific and the species pelagic life-history, the population structure may be maintained by homing behaviour and, perhaps, the association of spawning with retention zones.
Similar content being viewed by others
Introduction
Many marine organisms have large population sizes, high fecundity and high dispersal capabilities as planktonic larvae and free-swimming adults. Limited genetic differentiation between geographic populations would therefore be predicted, because of the high levels of gene flow facilitated by such traits. Similarly, allopatric speciation would be expected only when the populations have long been separated by geographical barriers (eg continents) or hydrological barriers (eg oceanographic fronts, divergences of water masses, salinity or temperature clines). Philopatric behaviour, gamete incompatibility, differences in spawning time or location and habitat selection could also be factors involved in inducing, maintaining or enforcing genetic differentiation.
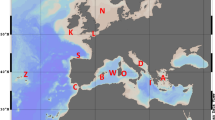
The Central Indo-West Pacific region is both geographically and hydrologically complex. This region is also reputed to have the highest marine biodiversity in the world (Mora et al, 2003; and references therein). Its shorelines have undergone important changes in the Pleistocene (Hewitt, 2000; Voris, 2000) with the shallow seas of the Sunda shelf (Java Sea) and Sahul shelf (Arafura Sea) left above present sea level, and the emergence of land barriers isolating the South China Sea from the Indian Ocean at its southern limit and from the Sulu Sea eastbound (Figure 1). The Sulu Sea was itself isolated from the Sulawesi Sea (Figure 1). Thus, repeated fluctuations of sea levels in this topographically complex region could have favoured the geographic isolation of several inland seas during the ice ages, leading to the genetic differentiation of the populations they harboured. This is thought to be the cause of a phylogeographic break between Indian and Pacific populations of several marine species, and also a factor contributing to the greater species diversity reported for the Indo-Malay region (Hewitt, 2000). Geographic discontinuities have been reported for several invertebrate species distributed widely in the Indo-West Pacific (Williams and Benzie, 1998; Barber et al, 2000, 2002; Benzie et al, 2002; Gopurenko and Hughes, 2002). While no genetic differences were evident for goatfish (Mulloidichthys vanicolensis) populations across the Indo-West Pacific (Stepien et al, 1994), two other surveys have reported geographic disjunction between Indian and West Pacific mitochondrial lineages and populations of inshore fishes, which was thought to be reminiscent of Pleistocene isolation. A large phylogeographic discontinuity has been reported for barramundi (Lates calcarifer) on either side of the Torres Strait (Chenoweth et al, 1998), which formed a land barrier connecting Australia and New Guinea during the Pleistocene (Voris, 2000). Admixture of haplotypes from two distinct mitochondrial lineages was interpreted as evidence for recent secondary introgression in barramundi (Chenoweth et al, 1998). The phylogeographic structure of false clown anemonefish (Amphiprion ocellaris), a coastal sedentary fish with short pelagic larval stage, was also explained by sea level changes during the Pleistocene, rather than by contemporary geography (Nelson et al, 2000). Significant differences in Cytochrome b haplotype frequencies were found between A. ocellaris populations from the western edge of the Sunda Shelf (West coast of Sumatra, Malacca Strait) and those from the rest of the Indo-Malay archipelago, including South China Sea, Sunda Strait, Bali Strait, Sulu Sea and Sulawesi Sea. Gene flow in A. ocellaris was inferred to be highest between regions connected by the permanent current that flows through the Makassar Strait from the Sulawesi Sea to the Bali Strait (Indonesian throughflow: Fieux et al, 1994). Pronounced geographic structure was also inferred from restriction-fragment length polymorphisms of the mitochondrial control region among populations of snapper (Pristipomoides multidens), sampled from the seas between northern Australia, Indonesia and West Papua (Ovenden et al, 2004). This was attributed to constraints to dispersal at all life stages.
Decapterus russelli. Sampling locations (open circles) in the Indo-Malay archipelago. Abbreviations for samples, sampling dates and sample sizes as following. K: Kelang, November 1998, N=12; C: Carita, October 1997, N=42; L: Labuan, May 1995 and October 1997, N=79; T: Tambelan, April 1995, N=52; P: Pekalongan, April 1995, N=67; Kk: Kota Kinabalu, October 1997, N=37; S: Sandakan, October 1997, N=66; M: Makassar, September 1998, N=22; Tt: Toli-Toli, May 1995, N=30; A: Arafura Sea, April 1998, N=51. MS, Makassar Strait; SS, Sunda Strait. Bold arrows: dominant surface currents; dotted arrows: seasonal surface currents (after Wyrtki, 1961; Anonymous, 1978). Inset: land areas (shaded) in the Indo-Malay region at the apogee of Pleistocene glaciations, inferred from Present bathymetric charts and using as coastline −120 m below Present sea level (see Voris, 2000).
In contrast with the foregoing examples of inshore, sedentary fishes and invertebrates, little genetic heterogeneity has been reported for pelagic fishes from the Indo-Pacific. In spite of extensive sampling, no significant genetic differences were found between Indian and Pacific-Ocean populations of bigeye tuna (Thunnus obesus) (Alvarado Bremer et al, 1998; Chow et al, 2000), or Indo-Pacific sailfish (Istiophorus platypterus) (Graves and McDowell, 1995). Although a preliminary survey hinted at the possible genetic distinctness of round scad mackerel (Decapterus macrosoma) from the Sunda Strait vs populations of the Sunda Shelf (Arnaud et al, 1999), no genetic differences were evident at either mitochondrial or nuclear loci among round scad mackerel populations at the scale of the Indo-Malay archipelago (Borsa, 2003). The spatial heterogeneity reported for yellowfin tuna (Thunnus albacares) (Ward et al, 1997) is a noticeable exception to the general pattern of panmixia in pelagic fishes in the Indo-Pacific.
Here we address the case of a highly mobile, pelagic fish of the scad mackerel group that exhibits geographic structure across the Indo-Malay archipelago. The Indian scad mackerel, D. russelli (Carangidae), is abundant and widely distributed in the Indo-West Pacific and is a major fishery resource in South-East Asia (Mansor and Abdullah, 1995; Smith-Vaniz, 1999). However, knowledge of the biology, ecology, distribution and stock structure of this and all other tropical Carangidae is still preliminary. First-hand information on the spawning areas and the reproductive periods of D. russelli is scarce (Delsman, 1926; De Jong, 1940). Fishery statistics show large fluctuations in the catches of D. russelli depending on the season (Sadhotomo, 1998). Salinity variation in relation with the monsoon cycle would be an important factor in the distribution and the migration of Indian scad mackerel in the Indo-Malay archipelago (Hardenberg, 1937; Potier and Boely, 1990). From the temporal patterns in fishery catches, Hardenberg (1937) suggested that at least two populations of scad mackerel seasonally enter the Java Sea. One population may originate from the seas east to the Java Sea and reach the central-western part of the Java Sea during the East monsoon; the other stock may originate in the Indian Ocean and reach the western part of the Java Sea, through the Sunda Strait, during the West monsoon. Hardenberg (1937) also suggested that a third population occupying the southern part of the South China Sea reaches the northwestern part of the Java Sea during the West monsoon.
In a preliminary survey based on the sequence polymorphism of the cytochrome b gene, Perrin and Borsa (2001) distinguished two putative lineages in D. russelli mitochondria in the Indo-Malay archipelago, which were separated by 2.2% average nucleotide divergence. These two mitochondrial lineages had a very heterogeneous geographic distribution and co-occurred at high frequencies in the Sulawesi Sea only. The presence in some samples of these two distinct lineages could be explained by either recent secondary contact between formerly isolated populations, or admixture of cryptic species. Mitochondrial-DNA haplotype frequency data and sequence phylogeny provided insufficient information to distinguish among the potential scenarios of genetic differentiation or cryptic speciation in D. russelli.
In the present study, we increased the precision of the mitochondrial phylogeography of this species by expanding the study area and the total sample size, and the population genetic structure of D. russelli in the Indo-Malay archipelago was investigated using novel nuclear markers. The question we addressed was that of the origin and maintenance of geographic differences in this highly vagile species.
Materials and methods
DNA samples
In total, 458 Indian scad mackerel (D. russelli (Rüppell 1830)) were sampled in 1995–1998 from small-sized purse seiners operating on a daily basis in the vicinity of eight landing places in the Indo-Malay archipelago (Kelang, Carita, Labuan, Pekalongan, Kota Kinabalu, Sandakan, Toli-Toli and Makassar). The Tambelan sample, from around the Tambelan Islands in the Indonesian part of the South China Sea was collected from the fish-hold of a medium-sized purse seiner upon calling in Pontianak (Kalimatan Barat, Indonesia). Another sample, caught by trawling in the Arafura Sea off West Papua, was purchased frozen at the fish market in Ambon, South Maluku. Figure 1 reports the sites and dates where Indian scad mackerel were sampled in Indonesia, Malaysia and West Papua (see Table 1 for sample sizes). Individual length was available for one sample (Toli-Toli). Length-frequency data allowed us to define size classes and to test the null hypothesis of genetic homogeneity across cohorts.
For each individual, a piece of muscle or a pectoral fin was stored in 95% ethanol. DNA was extracted from ca 5 mg dehydrated fish tissue placed in 500 μl extraction buffer (25 mM EDTA; 0.05 M Tris HCl pH 8; 1% SDS) with 12 U proteinase K overnight at 55°C, followed by a phenol–chloroform–isoamylalcohol extraction cycle (Sambrook et al, 1989). The DNA pellet was resuspended in 100 μl deionized water and frozen at −20°C for subsequent use as template DNA in polymerase chain reactions (PCR).
Cytochrome b gene: PCR/SSCP and sequencing
A 355-base pair (bp) fragment of the mitochondrial (mt) DNA cytochrome b gene was amplified by PCR using universal primers PAT (5′>GCC CCT CAG AAT GAT ATT TGT CCT C 3′>) and CTB (5′> CCA ACA TTT CAG CAT GAT GAA 3′>) modified from, respectively, primers CB1-L and CB2-H of Palumbi et al (1991). PCR amplifications were carried out in 25 μl reaction mixture containing ∼50 ng template DNA, 2.5 mM MgCl2, 0.48 mM dNTPs, 0.12 μM each primer and 0.38 U Taq DNA polymerase (Promega, Madison WI, USA) in 1 × specific Taq buffer (from 10 × buffer supplied by Promega). PCR conditions were 40 s at 94°C, followed by 35 cycles of denaturing at 94°C for 20 s, hybridization at 45°C for 12 s and polymerization at 72°C for 12 s with a final elongation step at 72°C for 3 min, in a Crocodile III thermocycler (Appligène, Strasbourg). Sequence polymorphism was detected by single-strand conformation polymorphism (SSCP) analysis (Lessa and Applebaum, 1993). This technique allows the detection of ∼90% sequences for fragments of size between 300 to 450 bp when these differ from one another by as little as one nucleotide change (Hayashi, 1991; Lessa and Applebaum, 1993). For SSCP, 3 μl PCR product was added to 6 μl denaturing buffer containing 1 vol 6 × loading buffer (0.25% xylene cyanol, 0.25% bromophenol blue, 30% glycerol, 0.5 M EDTA) and 5 vol 95% formamide, 10 mM NaOH and 20 mM EDTA, and was denaturated at 95°C for 5 min. The single-strand DNAs were run at 18°C for 15 h 30 min at 2 W in 1 × nondenaturing MDE polyacrylamide gel (FMC Corporation, Rockland ME, USA) in a vertical electrophoresis tank (Hoefer, San Francisco CA, USA). DNA bands were visualised by silver-nitrate staining (Wray et al, 1981). All haplotypes that were detected by SSCP (Figure 2A) were sequenced using the Thermo Sequenase Radiolabeled Terminator kit of Amersham Life Science (Cleveland OH, USA) after the purification of the PCR products (1 U exonuclease I and 1 U alkaline phosphatase (Amersham) added to 5 μl PCR product, incubated at 37°C for 15 min, and the enzymes inactivated at 80°C for 15 min). The DNA fragments produced by the sequencing reaction were separated by vertical electrophoresis in 6% denaturing polyacrylamide gel (6% 29:1 acrylamide/bis-acrylamide (BioRad Laboratories, Hercules, Canada) in 0.5 × TBE buffer). Our nomenclature for SSCP phenotypes followed that of Perrin and Borsa (2001).
D. russelli. (a) Single-stranded DNA conformation polymorphism at the Cytochrome b locus. Composite drawing of all 19 SSCP phenotypes found in the total sample. (b) Length polymorphism at locus Aldolase B-Intron 1, revealed by electrophoresis on denaturing polyacrylamide gel. Six of the seven alleles thus detected in the total sample are shown. Lane 1: inferred genotype was r2/r2; lanes 2, 8: r0/r0; lanes 3, 4, 5: r0/r−3; lanes 6, 7: r0/r−1; lane 9: r0.5/r0; lane 10: r1/r1. (c) Intron-length polymorphism at a Gonadotropin Releasing Hormone locus (GnRH 3-2), revealed by electrophoretic differences on denaturing polyacrylamide gel. Nine out of the 10 alleles detected in the total sample by using this technique are shown. Lane 1: inferred genotype was 45/50; lane 2: 50/50; lane 3: 40/50; lane 4: 45/100; lane 5: 60/90; lane 6: 35/50; lane 7: 45/50; lane 8: 45/110; lane 9: 30/50.
Nuclear-DNA markers
Two introns showing length polymorphism in preliminary tests were used as nuclear genetic markers in D. russelli. The first intron of the gene encoding aldolase B (Aldolase B-1) was amplified using PCR primers 1.1F (5′> GCT CCA GGA AAG GGA ATC CTG GC 3′>) and 1.2R (5′>CTC GTG GAA GAA GAT GAT CCC GCC 3′>) designed in the flanking exons by Lemaire (2001). Just before the PCR, primer 1.2R was radioactively labelled for 30 min at 37°C in a mixture of 2 μM primer, 1 U T4 polynucleotide-kinase (Eurogentec, Liège, Belgium) and 1.7 μM (1.85 MBq) [γ-33P]ATP (Isotopchim, Ganagobie, France). PCR amplifications were carried out in 10 μl reaction mixture containing 4 μl DNA solution, 2.5 mM MgCl2, 0.3 mM dNTP, 0.03 μM radioactively labelled primer, 0.4 μM primer 1.1F, 0.5 U Taq DNA polymerase (Promega) in 1 × Taq buffer. The PCR was run in a Crocodile III thermocycler (Appligène, Strasbourg, France). PCR programme parameters were 3 min at 95°C, followed by 35 cycles of denaturing at 94°C for 12 s, hybridization at 52°C for 12 s and polymerization at 72°C for 20 s, followed by a final elongation step at 72°C for 3 min. Length polymorphism was detected after allowing the heat-denaturated PCR products to migrate vertically for 3 h 30 min at 50 W in 6% denaturing polyacrylamide gel (Figure 2B). The gel was transferred onto a sheet of Whatman paper, vacuum-dried for 1 h at 80°C, and autoradiographed against X-Omat film (Eastman-Kodak, Rochester NY, USA).
The second intron of the gonadotropin-releasing hormone gene (GnRH3) was amplified using PCR primers GnRH2F (5′> AGA AGT GTG GGA GAG CTA GAG GC 3′>) and GnRH2R (5′> AGA GAC ACC ACT TCT CCT GTA CCC 3′>) designed by Hassan et al (2002) from a sequence of the Cichlidae Astatotilapia burtoni (GenBank AF076963). PCR amplifications were carried out in 10 μl reaction mixture containing 1 μl DNA solution, 1.25 mM MgCl2, 0.296 mM dNTP, 0.3 μM of primer GnRH2F labelled with 6-FAM fluorescent dye (Sigma Genosys, London, UK; carboxylfluorescein; maximum absorbance at 494 nm, maximum emission at 525 nm), 0.3 μM of the other primer (GnRH2R), 0.25 U Taq polymerase (Promega) in 1 × Taq buffer. The PCR programme used was the same as for amplifying Aldolase B Intron 1. After 5 min denaturing at 95°C in 2 vol denaturing buffer, the PCR products were run for 6 h at 40 W in 8% denaturing polyacrylamide gel. Fluorescent DNA bands were revealed under blue laser light (488 nm) in an FMBIO II gel scanner (Hitachi Instruments, San José CA, USA) equipped with an emission filter specific to fluorescein (505±20 nm). Intron-length polymorphism was thus detected at intron locus GnRH 3-2 (Figure 2c).
Data analysis
A neighbour-joining phylogeny of D. russelli mitochondria was constructed using Kimura's two-parameter model (MEGA2; Kumar et al, 2001). Parsimony analysis (MP) was also conducted, using MEGA2. Homologous Cytochrome b sequences of D. kurroides, D. macrosoma and D. tabl from Ambon, South Maluku; D. macarellus from Tobelo, Maluku and Selar crumenophthalmus from Biak, West Papua were used as outgroups. Support values for internal nodes of the tree were estimated using the bootstrap resampling procedure (Felsenstein, 1985). A total of 1000 pseudoreplicates were carried out for the NJ and MP runs.
A minimum parsimony network was produced to visualize the intraspecific phylogeny of D. russelli mtDNA haplotypes using the software TCS v. 1.13 (Clement et al, 2000). TCS estimates genealogies by implementing the statistical parsimony method described in Templeton et al (1992).
The gene diversities (h), the nucleotide diversities (π) and their standard deviations were estimated using ARLEQUIN (Schneider et al, 2000). The average nucleotide divergence between mitochondrial haplogroups A and M was estimated with Kimura's two-parameter model using MEGA2. ARLEQUIN was also used to test for departures from the equilibrium between mutation and genetic drift in the mtDNA sequence data, with Tajima's D-test (Tajima, 1989). For a selectively neutral marker, negative D-values indicate excesses of low-frequency haplotypes, relative to mutation/drift equilibrium, with one or very few alleles at high frequency and rare alleles that derive from the latter by very few mutations. This is generally ascribed to rapid population expansion following a severe reduction in effective size (bottleneck). Conversely, positive values point to secondary contact among previously differentiated lineages.
Correspondence analysis (CA) (Benzécri, 1982) was performed on the matrix of allelic frequencies (AFC procedure in BIOMECO; Lebreton et al, 1990). This amounts to estimating the degree of genetic differentiation between populations because the eigenvalues of each axis of the CA are analogous to partial Fst (Guinand, 1996).
Wright's Fis and global- and pairwise Fst were estimated, respectively, using Weir and Cockerham's (1984) multiallelic f̂ and θ̂ estimators (procedure FSTATS of GENETIX (Belkhir et al, 2000)). The estimator θ̂ can be used for both haploid and diploid data and is independent of the sampling strategy (Weir and Cockerham, 1984). It is widely used, thus allowing meaningful comparisons with other surveys of genetic differentiation in natural populations. The null hypotheses f=0 and θ=0 were tested using the permutation test procedures implemented in GENETIX, where pseudo-distributions of f̂ and θ̂ were obtained by random permutations of alleles and genotypes, respectively, and used as the expected distributions under the null hypothesis. The probability of occurrence of a parameter value larger than or equal to the observed value was estimated as P=(n+1)/(N+1), where n is the number of pseudo-values larger (or lower for negative f̂) than or equal to the observed value and N is the number of random permutations (Sokal and Rohlf, 1995).
Pairwise linkage disequilibria were tested using GENETIX (Belkhir et al, 2000). The average correlation coefficient between alleles at two loci (Rij) was estimated according to Garnier-Géré and Dillmann (1992). The null hypothesis (Rij=0) was tested by permutations of genotypes at a locus.
Results
Cytochrome b sequences
A total of 19 different SSCP phenotypes at the Cytochrome b locus were detected in the total sample (Figure 2A), including three novel SSCP phenotypes that were not present in the samples formerly surveyed by Perrin and Borsa (2001). One of the SSCP phenotypes (h) proved after sequencing to consist of two haplotypes (h1, h2) differing by one nucleotide substitution (Perrin and Borsa, 2001). Phenotype A was the most common in all samples (frequency >0.82) except Toli-Toli and Makassar, where the most common SSCP phenotype was M (frequency >0.63). Phenotype M was absent from all the other samples, but some other haplotypes (h1, h2, i) of Perrin and Borsa's (2001) haplogroup M were present at low frequencies in samples Carita, Labuan and Pekalongan. The frequencies of all SSCP phenotypes except A and M were lower than 0.04 in a sample, and lower than 0.01 in the total sample.
Among the 307 bp sequenced for each of the 20 Cytochrome b haplotypes represented by the above 19 SSCP phenotypes, 20 sites were variable, 11 of which were phylogenetically informative. Sequences have been deposited in GENBANK (http://www.ncbi.nlm.nih.gov) (accession numbers nos. AF307494-AF307510 and AF515757-AF515759). Of the 20, 18 nucleotide substitutions in the total sequence data set occurred at third-nucleotide positions.
Both NJ and MP trees of partial nucleotide sequences of the Cytochrome b gene (data not shown) featured the two phylogenetically distinct mitochondrial lineages, haplogroup A and haplogroup M, formerly identified by Perrin and Borsa (2001). The average nucleotidic distance between the two haplogroups was d±SE=2.2±0.7%. Haplogroups A and M were supported by bootstrap values of 72 and 56–64%, respectively. This is statistically weak, but the geographic distribution of haplotypes also pointed to the delineation between haplotypes A and M, which were the most frequent haplotypes of haplogroups A and M, respectively. While the inclusion of haplotype M into haplogroup M together with haplotypes h1, h2 and i was not strongly supported, the clade formed by the three latter haplotypes was more substantiated (bootstrap score=78–79%). The three novel haplotypes detected in the samples (ii, bb′ and d) clustered with haplogroup-A haplotypes.
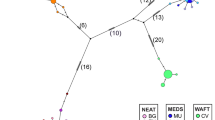
The haplotype network (TCS diagram) derived from cytochrome b gene partial sequences is presented in Figure 3. Three of these rarer haplotypes (h1, h2 and i) previously assigned to haplogroup M actually appeared to form a distinct cluster, separated from A by four mutational steps. Haplotype M was separated from A by five mutational steps and from the h1/h2/i cluster by three or four mutational steps only. The reticulations in the haplotype network, including the apparently intermediate position of M, may be explained by homoplasy, resulting from multiple substitutions at a given nucleotide site, or by recombination (see Hoarau et al (2002) for such an example in flounder, Platichthys flesus). The average nucleotidic distance between haplogroup A and haplotype M (now singled out) was 2.2%.
D. russelli. Statistical parsimony network of mtDNA haplotypes. Dot circles designate haplotypes whose frequency was <0.01 in the total sample. Small squares designate putative mutationally intermediate haplotypes. Nucleotide sequences deposited in GENBANK under accession nos AF307494-AF307510 and AF515757-AF515759.
Samples Toli-Toli and Makassar showed high gene (ĥ≥0.40) and nucleotide (π̂≥0.007) diversities at the Cytochrome b locus (Table 1). Both samples possessed the M haplotype at high frequency, together with a substantial proportion of clade-A haplotypes (Table 1). All the other samples had ĥ≤0.32 and π̂≤0.002 (Table 1) and the Arafura Sea sample was totally monomorphic for haplotype A (Table 1).
Tajima's D-values were strongly negative for all samples except Makassar and Toli-Toli, for which they were strongly positive (Table 1).
Intron-length polymorphism
The size of allele r0 at the Aldolase B-1 locus was 265 bp. All the other alleles differed from allele r0 by a few base pairs. Allele r0 was the most common (frequency >0.75) in all samples except the Arafura Sea sample, which was dominated by allele r1 (frequency=0.67). Five other alleles were detected at locus Aldolase B-1, with frequencies ≤0.06 (Figure 2b; Table 1).
Allele sizes (including primers) at the GnRH 3-2 locus were centred around 300 pb, estimated by linear interpolation against the CXR 60–400 bp ladder (Promega, Madison WI, USA). Owing to the difficulty in distinguishing some alleles differing from one another by tiny size differences, these were grouped as compound alleles. Compound alleles 30, 40, 90 and 110 each included two or three alleles (see footnote to Table 1) that were distinct on some gels, but not on other gels. The three alleles dominant in frequency were compound alleles 40 (0.54> frequency per sample >0.16), 50 (0.54> frequency >0.27) and 110 (0.33> frequency >0.09). The four other alleles that were detected, including compound allele 90 had frequencies ≤0.15 (Table 1).
Population genetic structure
Fixation index values (f̂) estimated from each sample at the Aldolase B-1 and GnRH 3-2 loci were not significantly different from zero (0.04<P<0.83; two-tailed test). Therefore, the hypothesis of panmixia was not rejected for any of the populations sampled.
The θ̂-values on the total set of samples significantly departed from 0 at all three loci (Cytochrome b: θ̂=0.370, P<0.001; Aldolase B-1: θ̂=0.165, P<0.001; GnRH 3-2: θ̂=0.016, P<0.01; 1000 random permutations using GENETIX), thereby demonstrating strong geographic structure of D. russelli populations across the Indo-Malay archipelago. The null hypothesis of homogeneity could not be rejected at any one locus (Cytochrome b: θ̂=−0.006; Aldolase B-1: θ̂=−0.005; GnRH 3-2: θ̂=−0.001; 1000 random permutations using GENETIX) for the group of samples (that is, Carita, Labuan, Tambelan, Pekalongan and Kinabalu) selected for testing Hardenberg's hypothesis of two or three stocks in the periphery of the Java Sea (see Introduction).
There were some discrepancies among loci regarding the geographic patterns of genetic differentiation, as indicated by population-pairwise θ̂-estimates at each locus (Table 2). At the Cytochrome b locus, all significant pairwise θ̂-estimates involved either population Makassar or Toli-Toli vs any other population in the Indo-Malay archipelago. At the Aldolase B-1 locus, the Arafura Sea population distinguished itself from all the other populations. At the GnRH 3-2 locus, significant genetic differences were found between the Arafura Sea population and a homogeneous group constituted by populations Sandakan, Makassar and Toli-Toli, while each of these two entities was not significantly distinct from the other populations in the Indo-Malay archipelago. Slight, albeit nonsignificant, genetic differences were also observed at locus GnRH3-2 between the Sandakan/Makassar/Toli-Toli group and the western populations.
Considering each locus separately, the patterns of genetic differentiation were meaningful from a geographical perspective (Figure 1). However, the geographic barrier uncovered by genetic analysis at the Cytochrome b locus was not evident when considering the Aldolase B-1 locus, and vice versa, while at locus GnRH 3-2 a geographic barrier was evident only between those populations that were singled out at either of the two other loci. The synthetic picture emerging from the present multiple-locus analysis (see Figure 4 presenting the results of CA) was that of three genetically differentiated geographic populations, being the Arafura Sea population, the Sulawesi Sea/Makassar Strait population and the rest of the Indo-Malay archipelago.
Correspondence analysis (CA) of D. russelli samples, where each sample was defined by its allele frequencies at loci Cytochrome b, Aldolase B-1, and GnRH 3-2. Projection of samples on CA's princial plane (axis 1 and axis 2). Inertia of each axis (that is, the percentage of total genetic variation that is explained by it; see Benzécri, 1982) is given in brackets. Ellipses delineate groups of samples that did not exhibit significant pairwise θ̂ between them at any one locus (see Table 1). Abbreviations for samples as in legend to Figure 1.
The distribution of size classes in the Toli-Toli sample was bimodal (data not shown), designating two separate size-cohorts. No allele-frequency differences were evident between the two size-cohorts at loci Cytochrome b (χ2[1]=1.167; P=0.28) and Aldolase B-1 (χ2[2]=0.359; P=0.836), while significant heterogeneity between cohorts was detected at locus GnRH3-2 (χ2[4]=10.413; P=0.034).
No pairwise genotypic or nucleo-cytoplasmic disequilibria between loci was evident, as indicated by the generally low Rij values per sample, none of which was significant under the null hypothesis of linkage equilibrium (Table 3).
Discussion
A single population or two sympatric species?
Three differentiated mitochondrial lineages were present in Indian scad mackerel in the Indo-Malay archipelago. Former haplogroup M of Perrin and Borsa (2001) is now recognized to consist of two separate lineages. One lineage comprises haplotypes h1, h2 and i formerly identified by Perrin and Borsa (2001), whereas haplotype M should now be singled out as a phylogenetically separate lineage. The geographic distribution of the three lineages was heterogeneous, with the western populations (those represented by samples Kelang, Carita, Labuan, Tambelan, Pekalongan, Kinabalu, and Sandakan) harbouring both the lineage represented by haplogroup A and that represented by haplotypes h1, h2 and i, and the easternmost population (Arafura) possessing haplotype A only. Only the geographically intermediate population sampled in Makassar Strait/Sulawesi Sea harboured haplotype M, at high frequency, together with haplotype A. Following these observations, we hypothesize that the admixture in the latter population of haplotype-A and haplotype-M individuals may reflect the occurrence of two cryptic species. Mitochondrial phylogenies have, in some instances, been used for elucidating systematic relationships among closely related species in fishes (eg Miya and Nishida, 1997; Rocha-Olivares et al, 1999; Agnèse and Teugels, 2001; Colborn et al, 2001). The co-occurrence in some samples of separate mitochondrial lineages may indeed be indicative of reproductive isolation in marine fishes (eg Hoarau and Borsa, 2000; Borsa and Quignard, 2001; Pfeiler et al, 2002). Here, the alternative, null hypothesis that Toli-Toli and Makassar samples were drawn from a single panmictic population could not be rejected because genotypic frequencies at two nuclear loci did not depart from those expected under panmixia. Also, there was no evidence of nucleo-cytoplasmic disequilibrium in these two samples. In other words, there was no indication that A and M individuals corresponded to different species. Therefore, the co-occurrence in the Sulawesi Sea/Makassar Strait region of these two distinct haplotypes indicates past geographic isolation that has been followed by secondary contact.
The present results emphasize the utility of combining information from recombinant (nuclear) markers and mtDNA sequences for systematic research.
Population structure
Surface currents in the western part of the Indo-Malay archipelago seasonally vary according to typical monsoon cycles (Figure 1; Wyrtki, 1961; Anonymous, 1978). During the wet monsoon (November to March), currents flow towards the East with speeds of ca. 1 to 2 knots. During the dry monsoon (May–September), the circulation is completely reversed with currents flowing towards the West (0.5–1 knots). During the intermonsoon (April–October), the winds and the currents are weak and variable (Anonymous, 1978). The monsoon cycle induces shifts in water circulation and changes in salinity, which coincide with the seasonal migration of adults and influence the dispersion of the larvae of pelagic fishes (Hardenberg, 1937). According to Hardenberg (1937), two or perhaps three stocks of ‘layang’ (a term used by this author to designate both D. russelli and round scad mackerel, D. macrosoma) are present in the periphery of the Java Sea (see Introduction), which follow different migration routes and timings in relation with monsoons. However, the high connectivity of the seas in the Indo-Malay archipelago may favour high levels of gene flow between populations. Genetic data so far have failed to confirm Hardenberg's (1937) designation of several stocks in the periphery of the Java Sea in both D. macrosoma (Borsa, 2003) and D. russelli (present results). At the scale of the entire Indo-Malay archipelago, however, at least three distinct populations of D. russelli were identified by the present study: (i) Arafura Sea, (ii) Sulawesi Sea and Makassar Strait and (iii) the rest of the Indo-Malay archipelago. Sound fishery management commands that these three geographically delineated, genetically differentiated populations be considered as separate management units.
That no correlation was observed between size class and either mitochondrial haplogroup or genotype at locus Aldolase B-1 in sample Toli-Toli, but a significant correlation was observed with genotype at locus GnRH 3-2, suggests, perhaps, age-related selection at the latter locus or at a closely linked locus. Alternatively, this result could indicate subtle, random genetic differences between cohorts (see eg Ruzzante et al, 1996), warranting more insight into the fine-scale genetic and demographic structures of Indian scad mackerel. Another hypothesis is that the two cohorts may be drawn from genetically differentiated populations that mix outside the reproductive season (hereafter referred to as the ‘admixture hypothesis’). In this case, larger sample sizes for each cohort would be necessary to provide evidence of genetic differences at locus GnRH 3-2 and perhaps at the two other loci. However, the fact that samples Toli-Toli and Makassar did not exhibit any genetic differences at all three loci implies that, under the admixture hypothesis, the very same proportions of individuals from different cohorts were present in the two samples, which is unlikely.
Concordant evolutionary patterns point to Pleistocene vicariance
Assuming that the average rate of nucleotide change of the cytochrome b gene in marine fishes lies between the extremes of mammals and sharks, that is, respectively, 2.5 and 1.0% per million year (Irwin et al, 1991; Martin et al, 1992), the ca. 2.2% Cytochrome b divergence between lineages A and M in Decapterus russelli would correspond to between approximately 1.1 and 0.4 million year of allopatric isolation. This indicates that the split between haplogroup A and haplotype M dates to the mid-Pleistocene. The geographic distribution of haplotypes suggests that the differences may result from vicariance. Through the Pleistocene, the Sunda and Sahul Shelves were repeatedly exposed above sea level, constituting land barriers between the northeastern Indian Ocean and the western Pacific Ocean. The Java Sea and the Arafura Sea were completely above sea level, the South China Sea was completely isolated from the Indian Ocean in its South and from the Sulu Sea in its East, and the Makassar Strait was much narrower than Present (Voris, 2000), presumably restricting the oceanic flow between the Sulawesi Sea and the rest of the Indo-Malay archipelago. The pattern of Makassar Strait/Sulawesi Sea populations being isolated by Pleistocene sea levels has been suggested for a mantis shrimp (Barber et al, 2000, 2002). Nucleotidic distances between Indian and Pacific populations in each of two monophyletic groups of nominal species of butterflyfishes (Mc Millan and Palumbi, 1995) were of the same order as between D. russelli's haplogroup A and haplotype M. Two haplogroups separated by ca. 2.3% nucleotide divergence at the Cytochrome b locus were also detected in round scad mackerel (D. macrosoma) (Borsa, 2003). A similarly shallow phylogeographic break was reported in false clownfish on either side of the Malay Peninsula (Nelson et al, 2000) and between barramundi populations on either side of Torres Strait (Chenoweth et al, 1998). This apparently recurrent feature of the mitochondrial phylogenies of coastal fishes in the Indo-Malay region lends support to the hypothesis that changes in sea level associated with Pleistocene glaciations in higher latitudes provoked the geographic isolation of populations in the Indo-Malay region. Genetic divergence between geographically isolated population might have reached a level associated with possible reproductive isolation in coastal, sedentary species such as butterflyfishes (Mc Millan and Palumbi, 1995), but migratory, pelagic fishes such as D. russelli might have experienced shorter durations of geographic isolation. This briefer isolation could have prevented total reproductive isolation, hence secondary contact would have been possible when the sea level rose again.
Population bottlenecks
Bottlenecks followed by demographic expansions leave a genetic signature in extant populations in the form of a departure from the equilibrium between mutation and genetic drift, whereby rare alleles differ by only a few mutations from a single or a few prevalent alleles (Grant and Bowen, 1998). The general aspect of the mitochondrial haplotype network in D. russelli was consistent with the above hypothesis. Tajima's test was applicable for all samples except Arafura, which exhibits sample monomorphism. Strong, negative D-values indicate population expansion in all populations except Makassar and Toli-Toli. The highly significant, positive D-values inferred for the two latter populations points to another form of disequilibrium between mutation and drift. This was caused by the occurrence in these samples of two differentiated mitochondrial lineages, which may stem from recent secondary contact between populations that were isolated for a long period.
The apparent discrepancy observed between nuclear and mitochondrial markers is precisely what is expected from the scenario of a vicariance event separating two populations for a long time, followed by secondary contact. The discrepancy arises because of the markers' different modes of transmission and evolutionary dynamics. Since mtDNA is uniparentally inherited, its effective population size is four-fold smaller than that of nuclear genes (Palumbi and Baker, 1994; FitzSimmons et al, 1997). MtDNA is therefore more sensitive to demographic events such as reduction in population size and geographic isolation. Mitochondrial genetic differences between the Makassar Strait/Sulawesi Sea population and the Arafura Sea population were two to 10 times higher than those at nuclear loci, roughly consistent with the expected four-fold ratio derived from their respective effective population sizes. Additional explanations are necessary to account for the higher discrepancy among genetic markers observed between the Makassar Strait/Sulawesi Sea population and the western populations, such as sex-biased gene flow (eg male dispersal/female philopatry; Mossman and Waser, 1999) or selection at nuclear loci linked to the marker loci.
Low haplotype and nucleotide diversities point to small effective population sizes, due to either variance in reproductive success, cycles of demographic expansion and regression or recent bottlenecks or foundation events (Grant and Bowen, 1998). When compared to the other predominantly haplotype-A populations, the Arafura Sea population was remarkably monomorphic. To explain the presence of the single haplotype A in Arafura Sea D. russelli, a recent bottleneck or founding event should have involved no more than a few females, and the Arafura Sea population should be considered as geographically isolated from the rest of the Indo-Malay archipelago. The data from loci Aldolase B-1 and GnRH 3-2 indeed concur with the founder population hypothesis. As expected under this hypothesis, the effective number of alleles at locus GnRH 3-2 was lowest in the Arafura Sea sample. At Aldolase B-1, the alleles sampled in the Arafura Sea were the three most frequent ones in the rest of the Indo-Malay archipelago, even though the effective number of alleles was higher (Table 1), as a presumed consequence of shifts in allelic frequencies due to genetic drift. As expected from a scenario of geographic isolation, allelic frequencies at Aldolase B-1 in the Arafura Sea were radically different from those in the rest of the Indo-Malay archipelago.
At the apogee of Pleistocene glaciations, when most continental seas were above sea level, the habitat of coastal pelagic fishes was considerably reduced, implying much lower effective population sizes than at present. The rise in sea level and the concomitant expansion of potentially favourable habitat may have allowed demographic expansion and the reclaiming of shallow seas such as the Java Sea and the Arafura Sea.
Present barriers to gene flow
Both nuclear and mitochondrial markers showed sharp geographic structure in D. russelli in the Indo-Malay archipelago. How might allele-frequency differences between populations of this pelagic species have been maintained to the Present? D. russelli has a high dispersal potential, both by juveniles and adults which are known to undertake massive seasonal migrations (Hardenberg, 1937), and by larvae (Delsman, 1926). Eggs and larvae are transported passively by the strong marine currents that flow through the archipelago (Figure 1). The putative secondary contact between different populations following the increase of the sea level should have induced a rehomogenization of allele frequencies, which did not happen. We therefore have to admit that D. russelli populations still exchange very few effective migrants despite the present, apparent continuity of the pelagic habitat. This suggests either reproductive homing, or the choice by adults of spawning areas that also constitute retention zones for eggs and larvae, or both.
It would seem sensible to assume that the extreme migratory abilities of some pelagic fishes favour wide-scale geographic homogeneity in allele frequencies. The observation of sharp geographic differences in Indian scad mackerel (the present results) and some other highly mobile, pelagic species (eg Kotoulas et al, 1995; Ward et al, 1997) shows that migratory abilities may also be associated with increased potential for homing, hence for reproductive isolation.
Accession codes
References
Agnèse J-F, Teugels GG (2001). The Bathyclarias-Clarias species flock. A new model to understand rapid speciation in African Great Lakes. C R Acad Sci Paris, Sér III 324: 683–688.
Alvarado Bremer JR, Stequert B, Robertson NW, Ely B (1998). Genetic evidence for inter-oceanic subdivision of bigeye tuna (Thunnus obesus) populations. Mar Biol 132: 547–557.
Anonymous (1978). Grand archipel d'Asie (partie méridionale): Indonésie. Instructions Nautiques K6, Service Hydrographique et Océanographique de la Marine, Paris.
Arnaud S, Bonhomme F, Borsa P (1999). Mitochondrial DNA analysis of the genetic relationships among populations of scad mackerel (Decapterus macarellus, D. macrosoma and D. russelli) in South-East Asia. Mar Biol 13: 699–707.
Barber PH, Palumbi SR, Erdmann MV, Moosa MK (2000). Biogeography. A marine Wallace's line? Nature 406: 692–693.
Barber PH, Palumbi SR, Erdmann MV, Moosa MK (2002). Sharp genetic breaks among populations of Haptosquilla pulchella (Stomatopoda) indicate limits to larval transport: patterns, causes, and consequences. Mol Ecol 11: 659–674.
Belkhir K, Borsa P, Raufaste N, Chikhi L, Bonhomme F (2000). Genetix version 4.02, logiciel sous WindowsTM pour la génétique des populations. Université Montpellier 2: Montpellier, France.
Benzécri J-P (1982). L'analyse des données. 2. L'analyse des correspondances. Dunod: Paris.
Benzie JAH, Ballment E, Forbes AT, Demetriades NT, Sugama K, Haryanti et al (2002). Mitochondrial DNA variation in Indo-Pacific populations of the giant tiger prawn, Penaeus monodon. Mol Ecol 11: 2553–2569.
Borsa P (2003). Genetic structure of round scad mackerel Decapterus macrosoma (Carangidae) in the Indo-Malay archipelago. Mar Biol 142: 575–581.
Borsa P, Quignard J-P (2001). Systematics of the Atlantic-Mediterranean soles Pegusa impar, P. lascaris, Solea aegyptiaca, S. senegalensis, and S. solea (Pleuronectiformes: Soleidae). Can J Zool 79: 2297–2302.
Chenoweth SF, Hughes JM, Keenan CP, Lavery S (1998). When oceans meet: a teleost shows secondary intergradation at an Indian-Pacific interface. Proc R Soc Lond B 265: 415–420.
Chow S, Okamoto H, Miyabe N, Hiramatsu K, Barut N (2000). Genetic divergence between Atlantic and Indo-Pacific stocks of bigeye tuna (Thunnus obesus) and admixture around South Africa. Mol Ecol 9: 221–227.
Clement M, Posada D, Crandall KA (2000). TCS: a computer program to estimate gene genealogies. Mol Ecol 9: 1657–1659.
Colborn J, Crabtree RE, Shaklee JB, Pfeiler E, Bowen BW (2001). The evolutionary enigma of bonefishes (Albula spp.): cryptic species and ancient separations in a globally distributed shorefish. Evolution 55: 807–820.
Crow JF, Kimura M (1970). An Introduction to Population Genetics Theory. Alpha: Edina.
De Jong JK (1940). A preliminary investigation of the spawning habits of some fishes of the Java Sea. Treubia 17: 307–330.
Delsman HC (1926). Fish eggs and larvae from the Java sea.5. Caranx kurra, macrosoma, and crumenophthalmus. Treubia 8: 199–211.
Felsenstein J (1985). Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39: 783–791.
Fieux M, Andrié C, Delécluse P, Ilahude AG, Kartavtseff A, Mantisi F et al (1994). Measurements within the Pacific–Indian oceans throughflow region. Deep-Sea Res 41: 1091–1130.
FitzSimmons NN, Moritz C, Limpus CJ, Pope L, Prince R (1997). Geographic structure of mitochondrial and nuclear gene polymorphisms in Australian green turtle populations and male-biased gene flow. Genetics 147: 1843–1854.
Garnier-Géré P, Dillmann C (1992). A computer program for testing pairwise linkage desequilibria in subdivided populations. J Hered 83: 239.
Gopurenko D, Hughes JM (2002). Regional patterns of genetic structure among Australian populations of the mud crab, Scylla serrata (Crustacea: Decapoda): evidence from mitochondrial DNA. Mar Freshw Res 53: 849–857.
Grant WS, Bowen BW (1998). Shallow population histories in deep evolutionary lineages of marine fishes: insights from sardines and anchovies and lessons for conservation. J Hered 89: 415–426.
Graves JE, McDowell JR (1995). Inter-ocean genetic divergence of istiophorid billfishes. Mar Biol 122: 193–203.
Guinand B (1996). Use of a multivariate model using allele frequency distributions to analyse patterns of genetic differentiation among populations. Biol J Linn Soc 58: 173–195.
Hardenberg JDF (1937). Preliminary report on a migration of fish in the Java sea. Treubia 16: 295–300.
Hassan M, Lemaire C, Fauvelot C, Bonhomme F (2002). Seventeen new exon-primed intron-crossing polymerase chain reaction amplifiable introns in fish. Mol Ecol Notes 2: 334–340.
Hayashi K (1991). PCR-SSCP: a simple and sensitive method for detection of mutations in the genomic DNA. PCR Meth Appl 1: 34–38.
Hewitt G (2000). The genetic legacy of the Quaternary ice ages. Nature 405: 907–913.
Hoarau G, Borsa P (2000). Extensive gene flow within sibling species in the deep-sea fish Beryx splendens. C R Acad Sci Sér III 323: 315–325.
Hoarau G, Holla S, Lescasse R, Stam WT, Olsen JL (2002). Heteroplasmy and evidence for recombination in the mitochondrial control region of the flatfish Platichthys flesus. Mol Biol Evol 19: 2261–2264.
Irwin DM, Kocher TD, Wilson AC (1991). Evolution of the cytochrome b gene of mammals. J Mol Evol 32: 128–144.
Kotoulas G, Magoulas A, Tsimenides N, Zouros E (1995). Marked mitochondrial DNA differences between Mediterranean and Atlantic populations of the swordfish, Xiphias gladius. Mol Ecol 4: 473–481.
Kumar S, Tamura K, Jakobsen IB, Nei M (2001). Mega2: Molecular Evolutionary Genetics Analysis Software. Arizona State University: Tempe.
Lebreton J-D, Roux M, Banco G, Bacou A-M (1990). Biomeco (biométrie-écologie). Logiciel d'écologie statistique pour PC et compatibles. Centre National de la Recherche Scientifique: Montpellier.
Lemaire C (2001). Flux génique et adaptations locales: étude de la divergence génétique en milieu marin chez Dicentrarchus labrax (Linné, 1758). Thèse de Doctorat, Institut National Agronomique Paris-Grignon, Paris.
Lessa EP, Applebaum G (1993). Screening techniques for detecting allelic variation in DNA sequences. Mol Ecol 2: 119–129.
Mansor MI, Abdullah S (1995). Growth and mortality of Indian mackerel (Rastrelliger kanagurta) and slender scad (Decapterus russelli) off the East Coast of Peninsular Malaysia. Sci Mar 59: 533–547.
Martin AP, Naylor GJP, Palumbi SR (1992). Rates of mitochondrial evolution in sharks are slow compared with mammals. Nature 357: 153–155.
Mc Millan WO, Palumbi SR (1995). Concordant evolutionary patterns among Indo-West Pacific butterflyfishes. Proc R Soc Lond B 260: 229–236.
Miya M, Nishida M (1997). Speciation in the open ocean. Nature 389: 803–804.
Mora C, Chittaro PM, Sale PF, Kritzer JP, Ludsin SA (2003). Patterns and processes in reef fish diversity. Nature 421: 933–936.
Mossman CA, Waser PM (1999). Genetic detection of sex-biased dispersal. Mol Ecol 8: 1063–1067.
Nelson JS, Hoddell RJ, Chou LM, Chan WK, Phang VPE (2000). Phylogeographic structure of false clownfish, Amphiprion ocellaris, explained by sea level changes on the Sunda shelf. Mar Biol 137: 727–736.
Ovenden JR, Salini J, O'Connor S, Street R (2004). Pronounced genetic population structure in a potentially vagile fish species (Pristipomoides multidens, Teleostei, Perciformes, Lutjanidae) from the East Indies triangle. Mol Ecol 13: 1991–1999.
Palumbi SR, Baker CS (1994). Contrasting population structure from nuclear intron sequences and mtDNA of humpback whales. Mol Biol Evol 11: 426–435.
Palumbi S, Martin A, Romano S, McMillan WO, Stice L, Grabowski G (1991). The Simple Fool's Guide to PCR, Version 2.0. Department of Zoology and Kewalo Marine Laboratory, University of Hawaii: Honolulu.
Perrin C, Borsa P (2001). Mitochondrial DNA analysis of the geographic structure of Indian scad mackerel, Decapterus russelli (Carangidae) in the Indo-Malay archipelago. J Fish Biol 59: 1421–1426.
Pfeiler E, Colborn J, Douglas MR, Douglas ME (2002). Systematic status of bonefishes (Albula spp. from the eastern Pacific Ocean inferred from analyses of allozymes and mitochondrial DNA. Env Biol Fishes 63: 151–159.
Potier M, Boely T (1990). Influence de paramètres de l'environnement sur la pêche à la senne tournante et coulissante en mer de Java. Aquat Living Resour 3: 193–205.
Rocha-Olivares A, Rosenblatt RH, Vetter RD (1999). Cryptic species of rockfishes (Sebastes: Scorpaenidae) in the Southern Hemisphere inferred from mitochondrial lineages. J Hered 90: 409–411.
Ruzzante DE, Taggart CT, Cook D (1996). Spatial and temporal variation in the genetic composition of a larval cod (Gadus morhua) aggregation: cohort contribution and genetic stability. Can J Fish Aquat Sci 53: 2695–2705.
Sadhotomo B (1998). Bioécologie des principales espèces pélagiques exploitées en mer de Java. Thèse de Doctorat, Université Montpellier 2, Montpellier.
Sambrook J, Fritsch EF, Maniatis T (1989). Molecular Cloning: a Laboratory Manual. Cold Spring Harbor Laboratory Press: Cold Spring Harbor.
Schneider S, Roessli D, Excoffier L (2000). Arlequin: a Software for Population Genetic Data Analysis. Genetics and Biometry Laboratory, University of Geneva: Geneva.
Smith-Vaniz WF (1999). Carangidae. In: Carpenter KE, Niem VH (eds) The Living Resources of the Western Central Pacific, Volume 4. Bony Fishes Part 2 (Mugilidae to Carangidae) FAO species Identification Guide for Fishery Purposes Food and Agriculture Organization of the United Nations, Rome. pp. 2659–2756.
Sokal RR, Rohlf FJ (1995). Biometry. The Principles and Practice of Statistics in Biological Research, 3rd edn. Freeman and Co.: New York.
Stepien CA, Randall JE, Rosenblatt RH (1994). Genetic and morphological divergence of a circumtropical complex of goatfishes: Mulloidichthys vanicolensis, M. dentatus, and M. martinicus. Pac Sci 48: 44–56.
Tajima F (1989). The effect of change in population size on DNA polymorphisms. Genetics 123: 437–460.
Templeton AR, Crandall KA, Sing CF (1992). A cladistic analysis of phenotypic associations with haplotypes inferred from restriction endonuclease mapping and DNA sequence data. III. Cladogram estimation. Genetics 132: 619–633.
Voris HK (2000). Maps of Pleistocene sea levels in Southeast Asia: shorelines, river systems and time durations. J Biogeogr 27: 1153–1167.
Ward RD, Elliott NG, Innes BH, Smolenski AJ, Grewe PM (1997). Global population structure of yellowfin tuna, Thunnus albacares, inferred from allozyme and mitochondrial DNA variation. Fishery Bull US 95: 566–575.
Weir BS, Cockerham CC (1984). Estimating F-statistics for the analysis of population structure. Evolution 38: 1358–1370.
Williams ST, Benzie JAH (1998). Evidence of a biogeographic break between populations of a high dispersal starfish: congruent regions within the Indo-West pacific defined by color morphs, mtDNA, and allozyme data. Evolution 52: 87–99.
Wray W, Boulikas T, Wray VP, Hancock R (1981). Silver staining of proteins in polyacrylamide gels. Anal Biochem 118: 197–203.
Wyrtki K (1961). Physical oceanography of the southeast Asian waters. Naga Report, Scripps Institute of Oceanography, University of California, Vol. 2. pp 1–95.
Acknowledgements
We are grateful to R Andamari, AS Cabanban, G Lesage, M Potier, Rumini, B Sadhotomo, V Sarpedonti, Suwarso and Suprapto for helping in the collection of samples; to AS Cabanban and J-M Ecoutin for hospitality in Malaysia and in Indonesia; to F Bonhomme, B Crouau-Roy and J-D Durand for comments on earlier versions of this paper; to R Andamari, AS Cabanban, J-R Durand, J-M Ecoutin, M Fatuchri Sukadi, P Levang, J Panfili, S Prakoso, J Roch and J Widodo for facilitating research and contacts in Indonesia, Malaysia, South Maluku and West Papua; to M Tortelier for her help in drawing Figure 1; to two anonymous referees for helpful comments and corrections. Finally, we express our gratitude to F Bonhomme for invaluable support at Station Méditerranéenne de l'Environnement Littoral and Laboratoire Génome et Populations. Part of the present work was undertaken as research operations of Pelfish and Genetikan, two French-Indonesian (IRD-CRIFI) cooperative projects on the assessment of pelagic fish resources in the Indonesian seas. Funded by Réseau Diversité Marine, IRD/DRV, IFREMER URM 16 and CNRS UMR 5000.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Rohfritsch, A., Borsa, P. Genetic structure of Indian scad mackerel Decapterus russelli: Pleistocene vicariance and secondary contact in the Central Indo-West Pacific Seas. Heredity 95, 315–326 (2005). https://doi.org/10.1038/sj.hdy.6800727
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.hdy.6800727
Keywords
This article is cited by
-
Genetic analyses reveal panmixia in Indian waters and population subdivision across Indian Ocean and Indo-Malay Archipelago for Decapterus russelli
Scientific Reports (2023)
-
Comparative mitogenomics and phylogenetics of the family Carangidae with special emphasis on the mitogenome of the Indian Scad Decapterus russelli
Scientific Reports (2022)
-
Genetic diversity, population structure and historical demography of the two-spined yellowtail stargazer (Uranoscopus cognatus)
Scientific Reports (2021)
-
Complex phylogeographic patterns in the intertidal goby Chaenogobius annularis around Kyushu Island as a boundary zone of three different seas
Ichthyological Research (2021)
-
Connectivity and population structure of albacore tuna across southeast Atlantic and southwest Indian Oceans inferred from multidisciplinary methodology
Scientific Reports (2020)