Abstract
The CpG island methylator phenotype (CIMP+) in colorectal cancer (CRC) is defined as concomitant and frequent hypermethylation of CpG islands within gene promoter regions. We previously demonstrated that CIMP+ was associated with elevated concentrations of folate intermediates in tumour tissues. In the present study, we investigated whether CIMP+ was associated with a specific mRNA expression pattern for folate- and nucleotide-metabolising enzymes. An exploratory study was conducted on 114 CRC samples from Australia. mRNA levels for 17 genes involved in folate and nucleotide metabolism were measured by real-time RT-PCR. CIMP+ was determined by real-time methylation-specific PCR and compared to mRNA expression. Candidate genes showing association with CIMP+ were further investigated in a replication cohort of 150 CRC samples from Japan. In the exploratory study, low expression of γ-glutamyl hydrolase (GGH) was strongly associated with CIMP+ and CIMP+-related clinicopathological and molecular features. Trends for inverse association between GGH expression and the concentration of folate intermediates were also observed. Analysis of the replication cohort confirmed that GGH expression was significantly lower in CIMP+ CRC. Promoter hypermethylation of GGH was observed in only 5.6% (1 out of 18) CIMP+ tumours and could not account for the low expression level of this gene. CIMP+ CRC is associated with low expression of GGH, suggesting involvement of the folate pathway in the development and/or progression of this phenotype. Further studies of folate metabolism in CIMP+ CRC may help to elucidate the aetiology of these tumours and to predict their response to anti-folates and 5-fluorouracil/leucovorin.
Similar content being viewed by others
Main
Cancer is a disease with genetic and epigenetic abnormalities. Aberrant CpG island methylation is a common epigenetic alteration in a variety of malignancies (Jones, 2002). De novo methylation of CpG islands in promoter regions is believed to contribute to tumorigenesis by causing transcriptional silencing of tumour suppressor genes. Colorectal cancer (CRC) is one of the malignancies in which epigenetic changes have been extensively analysed. Research on clinical samples has shown that a subgroup of CRC shows concurrent hypermethylation of a large number of CpG islands. These have been termed CIMP+, for CpG island methylator phenotype (Toyota et al, 1999). CIMP+ tumours occur more frequently in the proximal colon of older patients and are associated with the microsatellite instability (MSI+) phenotype, tumour-infiltrating lymphocytes (TILs) and mutations in the BRAF oncogene (Hawkins et al, 2002; van Rijnsoever et al, 2002; Samowitz et al, 2005). Quantitative DNA methylation analysis using real-time techniques indicates that approximately 17% of CRC are CIMP+ (Ogino et al, 2006; Weisenberger et al, 2006; Iacopetta et al, 2007). A panel of five CpG island markers was recently proposed to standardise the definition of CIMP+ status (Weisenberger et al, 2006).
Although the existence of a CIMP+ CRC subgroup is evident, the aetiology of this phenotype is not well understood. We previously reported that CIMP+ was associated with elevated concentrations of the folate intermediates CH2FH4 and FH4 in CRC tissues (Kawakami et al, 2003). This suggests that folate metabolism may be an important factor in determining the DNA methylation status of primary CRC. Folate plays a major role in cellular homeostasis as a donor of one-carbon units for DNA methylation, protein methylation and nucleotide synthesis. Increased dietary folate intake and serum levels of folate show correlations with increased global DNA methylation levels in epidemiological studies (Pufulete et al, 2005b), animal models (Sohn et al, 2003) and clinical intervention studies (Pufulete et al, 2005a). Associations between dietary folate intake (van Engeland et al, 2003) or genetic variants in folate-metabolising enzymes (Paz et al, 2002) and CpG island hypermethylation in CRC have also been reported, although other workers have found less evidence for this (Curtin et al, 2007; Slattery et al, 2007). These observations suggest that folate metabolism, at least in part, can influence CpG island methylation and may therefore be involved in the development of CIMP+ CRC, although firm evidence for this is still lacking.
Two key metabolic pathways for methyl donor/one-carbon transfer reactions are the synthesis of folate and nucleotides. In the present study, we hypothesised that a specific expression pattern for folate- and nucleotide-metabolising enzymes occurs in CIMP+ CRC. Our rationale was that a distinctive gene expression signature may be associated with the aberrant methyl group metabolism of CIMP+ tumours, as evidenced by the frequent CpG island hypermethylation. To test this hypothesis, the mRNA expression levels of 17 genes with important roles in folate and nucleotide metabolism were measured by real-time RT-PCR in two series of primary CRC in which the CIMP+ status was determined by methylation-specific real-time PCR.
Materials and methods
Samples
For exploratory analysis of gene expression levels, tumour samples from a consecutive series of 114 CRC patients undergoing elective surgery at the Colorectal Unit of the Royal Adelaide Hospital in Australia were used. These samples were snap-frozen in liquid nitrogen within 20–40 min of resection and stored at −70 °C. DNA was extracted using a QIAamp DNA mini kit (Qiagen, Hilden, Germany), according to the manufacturer's protocol. RNA was obtained from the corresponding formalin-fixed and paraffin-embedded (FFPE) tissues. FFPE tissue blocks were reviewed for quality and tumour content, and 5-μm-thick sections were obtained. Sections were mounted on uncoated glass slides, deparaffinised in xylene, hydrated and stained with nuclear fast red (American MasterTech Scientific Inc., Lodi, CA, USA). Tumour cells were isolated by laser capture microdissection (PALM Microsystem; Leica, Wetzlar, Germany), according to the standard procedures (Bonner et al, 1997). RNA isolation after dissection was performed according to a proprietary procedure (Response Genetics Inc., US patent no. 6248535). We have previously measured the concentrations of the folate intermediates CH2FH4 and FH4 (Kawakami et al, 2003) and analysed for BRAF V600E mutation (Iacopetta et al, 2006) in this tumour series. Approval of this project was obtained from the IMVS Human Research Ethics Committee.
For the validation tumour set, 150 primary CRC samples from patients undergoing surgical treatment at Kanazawa University Hospital in Japan were used. Tumour was dissected manually from FFPE archival tissue sections of 10 μm thickness. After deparaffinisation using xylene and ethanol, genomic DNA was isolated using a QIAamp DNA mini kit. RNA was obtained from the manually dissected FFPE samples using the same method as for the Australian CRC series. Approval of this project was obtained from the Ethics Committee of Kanazawa University School of Medicine.
Real-time RT-PCR and immunohistochemistry
Complementary DNA was prepared as described previously (Lord et al, 2000). Quantification of the genes of interest (Table 1) and an internal reference gene (ACTB) was conducted using a fluorescence-based real-time detection method (ABI PRISM 7700 Sequence Detection System (TaqMan); Perkin-Elmer Applied Biosystems, Foster City, CA, USA), as previously described (Gibson et al, 1996; Dziadziuszko et al, 2006). Gene expression values were expressed as ratios (differences between Ct values) between the gene of interest and an internal reference gene (ACTB). Primer and probe sequences used in this study are listed in Supplementary Table 1.
For the validation study with Japanese CRC samples, different primer sets for ECGF1, γ-glutamyl hydrolase (GGH), RRM2 and ACTB (Supplementary Table 1) were used with SYBR Premix Ex Taq (TaKaRa Bio, Otsu, Japan) and following the protocol provided by the manufacturer using ABI PRISM 7700 Sequence Detection System. The quantity of mRNA was expressed as the ratio of the expression level between each test mRNA and ACTB mRNA.
Protein expression of GGH in tumour tissues was examined by immunohistochemistry for selected samples from the Japanese CRC cohort. The avidin–biotin–peroxidase complex method with chicken polyclonal antibody (IgY) to human GGH (diluted 1 : 100; GenWay Biotech, San Diego, CA, USA) and biotinylated rabbit anti-chicken IgY (diluted 1 : 200; Open Biosystems, Huntsville, AL, USA) was used following microwave antigen retrieval of paraffin sections, as described previously (Ougolkov et al, 2002).
Methylation analysis
Promoter methylation was evaluated for the CIMP panel of markers comprising CACNA1G, IGF2, NEUROG1, RUNX3 and SOCS1, where PMR (percentage methylated reference) values were derived using the ALU normalisation control reaction (Weisenberger et al, 2006). Simultaneous hypermethylation (PMR ⩾10) of 3 or more of these 5 markers was considered to represent CIMP+. Promoter hypermethylation of GGH was analysed as previously described (Cheng et al, 2006). Sperm DNA and fully methylated DNA by SssI methylase (NewEngland Biolabs, Ipswich, MA, USA) were used as unmethylated and methylated control samples, respectively.
Statistical analysis
Because mRNA expression levels did not show normal distribution, the results were expressed as median values (25th to 75th percentiles) in tables or boxplots. Non-parametric models were used for univariate analyses. The Mann–Whitney U-test was used to compare mRNA expression levels between two categorical variables. Correlations between mRNA expression and the concentration of folate intermediates were analysed by Spearman's rank test. A multivariate stepwise logistic regression approach was used to select genes whose mRNA expression was significantly related to CIMP status. All P-values shown are two tailed, with P<0.05 taken as significant.
Results
Associations between mRNA expression levels for folate- and nucleotide-metabolising enzymes and CIMP+ or CIMP+-related features
RT-PCR assays were conducted for 17 genes in 114 colorectal tumour samples from Australia. The assays were performed in triplicate for RT samples and in a single assay for non-RT controls, resulting in 1938 mRNA measurements by 7752 assays. The non-RT control reaction was positive in 31 measurements and the coefficient of variance was high among triplicate assays in four measurements. These were deemed as ‘no result’. In all, 1903 out of 1938 (98.2%) real-time RT-PCR measurements were successful using RNA derived from laser capture microdissected FFPE tumour tissues. CIMP+ was found in 18 out of 114 (15.8%) CRC samples.
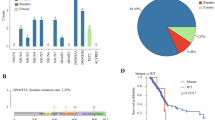
Cluster analysis did not reveal a distinctive mRNA expression profile associated with CIMP+ (data not shown). In univariate analysis (Mann–Whitney U-test), GGH expression was significantly lower in CIMP+ than CIMP− CRC, whereas the expression of DCK, DPYD, ECGF1, MTHFR and RRM2 was all higher in CIMP+ (Table 2). Multivariate analysis using a logistic regression model showed that GGH expression (odds ratio 0.70, 95% CI: 0.51–0.95, P=0.023) and RRM2 expression (odds ratio 1.25, 95% CI: 1.04–1.49, P=0.015) were associated with CIMP+ (P=0.008). Univariate analysis (Mann–Whitney U-test) showed that ECGF1 and GGH expressions were strongly associated with the CIMP+ features of proximal tumour site, TILs and BRAF mutation (Table 3 and Supplementary Table 2). γ-Glutamyl hydrolase expression was lower, whereas ECGF1 was higher in tumour with these CIMP+ features. The analyses showed that low expression of GGH was consistently associated with CIMP+ and CIMP+-related features (Figure 1). The high expression levels of RRM2 and ECGF1 also showed strong associations with CIMP+ and CIMP+-related features, respectively.
γ-Glutamyl hydrolase mRNA expression according to the CIMP status, tumour site, TILs and BRAF mutation status in an Australian CRC cohort was shown by boxplot. The level of GGH mRNA expression was significantly different between all dichotomised variables (Mann–Whitney U-test; CIMP, P=0.013; tumour site, P=0.021; TILs, P=0.001; BRAF mutation, P=0.002).
Finally, mRNA expression was compared to the concentrations of the folate intermediates CH2FH4 and FH4 in these CRC tissues (Table 4). None of the genes examined showed significant correlation with the concentrations of these intermediates, although high RFC1 expression was significantly correlated with low concentrations of FH4 (Spearman's ρ=−0.205, P=0.046). Trends for negative correlation between GGH expression and CH2FH4 and FH4 concentrations were observed (Spearman's ρ=−0.200, P=0.053 and Spearman's ρ=−0.180, P=0.083, respectively). The above exploratory analyses suggest that low GGH mRNA expression is a candidate CIMP+ molecular signature, possibly through its involvement in folate metabolism.
Validation of GGH downregulation in CIMP+ CRC
A validation study was conducted using 150 primary CRC samples from a Japanese cohort of patients. γ-Glutamyl hydrolase, ECGF1 and RRM2 were selected as candidates for further study because the expression of these genes was consistently associated with CIMP+ and/or CIMP+-related features in the Australian CRC series. Only 14 out of 150 (9.3%) of the Japanese CRC samples were found to be CIMP+ compared to 15.8% of the Australian tumours (P=0.11 in χ2 test). A random selection of CIMP− CRC (n=79) and all 14 CIMP+ CRC samples from the Japanese cohort were subjected to RT-PCR analysis of GGH, ECGF1 and RRM2 expression. The results confirmed the previous result that GGH mRNA expression was significantly lower in CIMP+ CRC samples from a separate tumour series (P=0.0012, Figure 2). No significant associations were observed between CIMP+ and either ECGF1 or RRM2 mRNA expression.
To further examine whether the mRNA level reflects GGH protein expression, selected paraffin tissues of Japanese CRC were immunostained using polyclonal antibody to human GGH. γ-Glutamyl hydrolase protein was not detectable or was weakly expressed in four samples with low mRNA levels (0.01, 0.06, 0.28 and 0.32), whereas much stronger expression was observed in five samples with high mRNA levels (3.25, 3.50, 3.73, 6.33 and 8.29). Representative cases are shown in Figure 3. The results indicate an association between levels of GGH mRNA and its protein expression.
Immunohistochemical staining of GGH in CRC tissues. γ-Glutamyl hydrolase protein was not detectable in tumour cells in case Nos. 5209 and 2426, in which the GGH mRNA levels were 0.01 and 0.06, respectively. Expression of GGH was observed in tumour cells in case Nos. 4040 and 616, in which the GGH mRNA levels were 3.73 and 8.29, respectively.
GGH promoter methylation is not a cause of GGH downregulation in CIMP+ CRC
A recent study in leukaemia found that hypermethylation of the GGH promoter was associated with silencing of GGH gene expression (Cheng et al, 2006). The above exploratory analyses showing that CIMP+ CRC samples have low GGH mRNA expression levels also raise this possibility. Methylation of the GGH promoter was therefore analysed in 18 CIMP+ tumours and in 20 randomly selected CIMP− tumours from the Australian CRC cohort. Only one CIMP+ tumour (5.6%) showed hypermethylation of the GGH promoter (Figure 4), indicating that it does not play a major role in downregulating the mRNA expression of this gene in CRC.
Methylation-specific PCR analysis of the GGH promoter. Promoter methylation of GGH was analysed using unmethylated DNA-specific primer sets (U) and methylated DNA-specific primer sets (M). Representative results using 18 samples of CIMP+ and 20 randomly selected samples of CIMP− tumours are shown. Only one sample, a CIMP+ tumour, showed GGH promoter hypermethylation. Sperm DNA and fully methylated sperm DNA produced with SssI methylase were used for unmethylated control (UC) and methylated control (MC), respectively. DNA from the colon cancer cell line WiDr was also used as a positive control.
Discussion
In this study, we explored the possibility that genes involved in folate and nucleotide metabolism have a distinct mRNA expression signature in CIMP+ CRC. Although no clear expression pattern was found for the 17 genes analysed, low GGH expression was observed in two independent series of CIMP+ CRC and could therefore play a role in the development of this phenotype. In the Australian CRC cohort, the mRNA expression levels for GGH and RRM2 were shown by univariate and multivariate analyses to be significantly associated with CIMP+ CRC. In addition, the mRNA expression of GGH and ECGF1 was associated with characteristic clinicopathological and molecular features of CIMP+, including proximal tumour site, TILs and BRAF mutation. Moreover, the concentrations of two folate intermediates, CH2FH4 and FH4, showed trends for association with GGH mRNA expression. As might be predicted from the function of GGH in hydrolysing glutamated folates and allowing escape from the cell (Figure 5), low GGH expression was associated with higher folate concentrations.
Simplified representation of folate transport and polyglutamylation reactions within the cell. RFC1 is ubiquitously expressed in epithelial cells and plays a role as a major transport system for folates. FOLR1 is anchored to cell membranes and transports folates via an endocytotic process. Intracellular monoglutamyl folate is converted to the polyglutamate form by FPGS, whereas the polyglutamate chains are removed by GGH.
Overall, the results of the exploratory study on the Australian CRC cohort provided evidence that low expression of GGH mRNA was associated with CIMP+ and with CIMP+-related features. This led us to conduct a further study using an independent cohort of primary CRC from Japan and in which we confirmed the relationship between low GGH mRNA expression and CIMP+ status. The frequency of CIMP+ was lower among Japanese CRC (9.3%) than that among Australian CRC (15.8%). Although this difference did not reach statistical significance, it suggests that dietary, environmental and genetic differences between these two populations could influence the frequency of the CIMP+ subgroup as a proportion of total CRC. Nevertheless, low expression of GGH mRNA was a consistent finding in both CIMP+ cohorts. Although a recommended panel of markers was used here to define CIMP+ (Weisenberger et al, 2006), the GGH/CIMP+ association was also found using a different CpG island panel comprising of MLH1, p16 (INK4A), TIMP3 and p14 (ARF) (data not shown).
The present results suggest that low GGH mRNA expression may play a role in the development and/or progression of CIMP+ CRC. A possible explanation for this is the role played by GGH in regulating intracellular folate levels (Figure 5). Monoglutamyl folate is transported into mammalian cells mainly by FOLR1 and RFC1 (Matherly and Goldman, 2003). Intracellular monoglutamyl folate is converted to the polyglutamate form by FPGS (Qi et al, 1999), whereas the polyglutamate chains are removed by GGH (Schneider and Ryan, 2006). Polyglutamate forms of folate are more strongly retained within the cell and are a better substrate for intracellular folate-dependent enzymes than the monoglutamate form. Therefore, low GGH expression would be expected to lead to a higher concentration of polyglutamated folate because of better retention in the cell. In agreement with this, we observed trends for an inverse correlation between GGH expression and the concentrations of folate intermediates CH2FH4 and FH4 (Table 4). FOLR1 and FPGS mRNA expression were not associated with the concentrations of these folate intermediates; however, the increased expression of RFC1 was significantly correlated with low concentrations of FH4. These results suggest that GGH expression plays a role in regulating the intracellular folate level in CRC tissues, although other factors such as RFC1 expression are also likely to be involved.
We previously reported that frequent CpG promoter hypermethylation was associated with high folate levels in CRC (Kawakami et al, 2003). A recent study also found that the level of p16 (INK4A) promoter methylation in the normal colonic mucosa of older mice increased following folate supplementation (Keyes et al, 2007). Together, the above results suggest that low GGH expression may be linked to increased promoter methylation in CIMP+ tumours by causing elevation of the folate concentration. An alternate explanation involving transcriptional silencing of GGH by promoter methylation was excluded by the finding that only 5.6% of CIMP+ tumours showed GGH hypermethylation (Figure 4). It is unknown whether low GGH expression and its link with high tissue folate concentrations play a causal or even supportive role in the development of CIMP+ CRC. Further studies are required in which GGH expression, folate status and CpG island methylation are evaluated in normal colonic tissue as well as in the proposed precursor lesion for CIMP+, the so-called serrated adenoma or hyperplastic polyp (Jass, 2006). The mechanism(s) by which GGH expression is regulated in both normal and malignant colorectal tissue also warrants further investigation. Apart from two studies that found GGH expression was increased in CRC compared to adjacent normal colonic mucosa (Odin et al, 2003; Kidd et al, 2005), no other work has been published in this area.
In addition to possible implications for the aetiology of CIMP+ CRC, the current findings are also relevant for the response of CRC to inhibitors of dihydrofolate reductase and thymidylate synthase (TS), both of which are key enzymes in nucleotide synthesis. The growth inhibitory effect of anti-folates such as methotrexate (MTX) and raltitrexed depends upon the polyglutamylation state of these agents (Barnes et al, 1999). Methotrexate is transported into cells using the same mechanism as that for folates and is also better retained following polyglutamylation. High GGH activity has been associated with the resistance of tumour cell lines to MTX via shortening of polyglutamate chains and consequently a lower intracellular drug concentration and less inhibition of dihydrofolate reductase and TS (Rhee et al, 1993; Barnes et al, 1999). Raltitrexed, a specific inhibitor of TS, is also polyglutamylated and its antitumour activity correlates with the amount of polyglutamylated drug inside the cells (Takemura et al, 1996). The importance of polyglutamylation in the antitumour activity suggests that CIMP+ CRC might have higher sensitivity to these anti-folates because of low GGH expression in this subtype of CRC. Neither MTX nor raltitrexed is widely used in chemotherapy for CRC. However, these anti-folates might be of clinical use for tailored chemotherapy.
In contrast to above-mentioned anti-folates, 5-fluorouracil (5-FU) and leucovorin have been key drugs for the chemotherapy of CRC. 5-Fluorouracil is thought to exert its major cytotoxic activity by inhibiting TS. It does this by forming a stable ternary complex between 5,10-methylenetetrahydrofolate, TS and fluoro-dUMP, the metabolite of 5-FU (Longley et al, 2003). Leucovorin, also known as folinic acid, increases the activity of 5-FU by raising the intracellular levels of 5,10-methylenetetrahydrofolate and thereby prolonging the inhibition of TS. 5,10-Methylenetetrahydrofolate is also better retained following polyglutamylation (Radparvar et al, 1989), and this is critical for the antitumour activity of 5-FU even when this folate intermediate is present at relatively high concentrations (Romanini et al, 1991). Therefore, supplementation of 5-FU with leucovorin may be more effective in CIMP+ compared to CIMP− CRC because the low GGH levels would better allow the retention and modulatory action of 5,10-methylenetetrahydrofolate. An earlier study did indeed find that adjuvant treatment with 5-FU/leucovorin conferred more benefit to CIMP+ tumours in stage III CRC (van Rijnsoever et al, 2003). Two more recent studies reported that CIMP+ CRC was associated with poor survival in advanced CRC treated with 5-FU-based chemotherapy (Ogino et al, 2007; Shen et al, 2007). This may, however, be a reflection of the prognostic rather than predictive value of CIMP+. Moreover, the regimens used in the two studies of advanced CRC included not only 5-FU alone but also in combination with other chemotheraputic agents such as IFNα-2a and irinotecan. Further prospective studies are needed to test whether CIMP+ is a predictive marker for response to 5-FU/leucovorin in CRC. These may allow chemotherapy regimens to be tailored according to the CIMP+ status, leading to more effective cancer treatment.
The present study investigated 17 genes involved in folate and nucleotide metabolism. Low expression of GGH was one of the features associated with CIMP+ CRC; however, it was not a specific marker for this phenotype because many CIMP− tumours also showed low expression of this gene. The aberrant promoter methylation observed in CIMP+ CRC is likely to be a multistep phenomenon that involves many factors in addition to folate metabolism and could include, for example, the expression levels of methyltransferases and histone deacetylase. Some of these factors may be revealed by array-based transcriptome analysis of CIMP+ and CIMP− CRC tissues. Although no study to date has addressed this issue, a few reports have described the mRNA expression profile of the closely associated MSI+ phenotype in CRC (Banerjea et al, 2004; di Pietro et al, 2005; Lanza et al, 2007). Interestingly, each of these studies showed that GGH mRNA expression was lower in MSI+ compared to MSI− CRC. Because of the strong concordance between MSI+ and CIMP+ in population-based CRC cohorts (Hawkins et al, 2002; van Rijnsoever et al, 2002; Samowitz et al, 2005; Ogino et al, 2006; Weisenberger et al, 2006), the results from these independent, array-based studies confirm the current results obtained using RT-PCR and two separate CIMP+ CRC series. Together, the studies provide strong evidences of low GGH expression in MSI+ and/or CIMP+ CRC. The three array-based studies did not show consistent association between expression of the other genes analysed in current study with MSI status. However, two of them (Banerjea et al, 2004; di Pietro et al, 2005) demonstrated higher expression of TYMS in MSI+ compared to MSI− CRC. Our result on TYMS expression between CIMP+ and CIMP− CRC did not support this association (P=0.099, Table 2). TYMS might differently express in MSI+ compared to CIMP+ CRC, requiring further study to establish molecular difference between MSI+ and CIMP+ CRC.
In conclusion, the present study is the first to investigate the expression of genes involved in folate and nucleotide metabolism in relation to CIMP+ CRC. This tumour phenotype is associated with low expression of GGH, suggesting involvement of the folate pathway in its development and/or growth. Further studies of folate metabolism in CIMP+ CRC, premalignant precursors and normal colonic mucosa may help to elucidate the aetiology of these tumours. A better understanding of the role of folate metabolism in DNA methylation may also lead to tailored chemotherapy that employs anti-folates, 5-FU/leucovorin and the use of CIMP+ markers.
Change history
16 November 2011
This paper was modified 12 months after initial publication to switch to Creative Commons licence terms, as noted at publication
References
Banerjea A, Ahmed S, Hands RE, Huang F, Han X, Shaw PM, Feakins R, Bustin SA, Dorudi S (2004) Colorectal cancers with microsatellite instability display mRNA expression signatures characteristic of increased immunogenicity. Mol Cancer 3: 21
Barnes MJ, Estlin EJ, Taylor GA, Aherne GW, Hardcastle A, McGuire JJ, Calvete JA, Lunec J, Pearson AD, Newell DR (1999) Impact of polyglutamation on sensitivity to raltitrexed and methotrexate in relation to drug-induced inhibition of de novo thymidylate and purine biosynthesis in CCRF-CEM cell lines. Clin Cancer Res 5: 2548–2558
Bonner RF, Emmert-Buck M, Cole K, Pohida T, Chuaqui R, Goldstein S, Liotta LA (1997) Laser capture microdissection: molecular analysis of tissue. Science 278: 1481–1483
Cheng Q, Cheng C, Crews KR, Ribeiro RC, Pui CH, Relling MV, Evans WE (2006) Epigenetic regulation of human gamma-glutamyl hydrolase activity in acute lymphoblastic leukemia cells. Am J Hum Genet 79: 264–274
Curtin K, Slattery ML, Ulrich CM, Bigler J, Levin TR, Wolff RK, Albertsen H, Potter JD, Samowitz WS (2007) Genetic polymorphisms in one-carbon metabolism: associations with CpG island methylator phenotype (CIMP) in colon cancer and the modifying effects of diet. Carcinogenesis 28: 1672–1679
di Pietro M, Sabates Bellver J, Menigatti M, Bannwart F, Schnider A, Russell A, Truninger K, Jiricny J, Marra G (2005) Defective DNA mismatch repair determines a characteristic transcriptional profile in proximal colon cancers. Gastroenterology 129: 1047–1059
Dziadziuszko R, Witta SE, Cappuzzo F, Park S, Tanaka K, Danenberg PV, Baron AE, Crino L, Franklin WA, Bunn Jr PA, Varella-Garcia M, Danenberg KD, Hirsch FR (2006) Epidermal growth factor receptor messenger RNA expression, gene dosage, and gefitinib sensitivity in non-small cell lung cancer. Clin Cancer Res 12: 3078–3084
Gibson UE, Heid CA, Williams PM (1996) A novel method for real time quantitative RT-PCR. Genome Res 6: 995–1001
Hawkins N, Norrie M, Cheong K, Mokany E, Ku SL, Meagher A, O’Connor T, Ward R (2002) CpG island methylation in sporadic colorectal cancers and its relationship to microsatellite instability. Gastroenterology 122: 1376–1387
Iacopetta B, Grieu F, Li W, Ruszkiewicz A, Caruso M, Moore J, Watanabe G, Kawakami K (2006) APC gene methylation is inversely correlated with features of the CpG island methylator phenotype in colorectal cancer. Int J Cancer 119: 2272–2278
Iacopetta B, Grieu F, Phillips M, Ruszkiewicz A, Moore J, Minamoto T, Kawakami K (2007) Methylation levels of LINE-1 repeats and CpG island loci are inversely related in normal colonic mucosa. Cancer Sci 98: 1454–1460
Jass JR (2006) Colorectal cancer: a multipathway disease. Crit Rev Oncog 12: 273–287
Jones PA (2002) DNA methylation and cancer. Oncogene 21: 5358–5360
Kawakami K, Ruszkiewicz A, Bennett G, Moore J, Watanabe G, Iacopetta B (2003) The folate pool in colorectal cancers is associated with DNA hypermethylation and with a polymorphism in methylenetetrahydrofolate reductase. Clin Cancer Res 9: 5860–5865
Keyes MK, Jang H, Mason JB, Liu Z, Crott JW, Smith DE, Friso S, Choi SW (2007) Older age and dietary folate are determinants of genomic and p16-specific DNA methylation in mouse colon. J Nutr 137: 1713–1717
Kidd EA, Yu J, Li X, Shannon WD, Watson MA, McLeod HL (2005) Variance in the expression of 5-fluorouracil pathway genes in colorectal cancer. Clin Cancer Res 11: 2612–2619
Lanza G, Ferracin M, Gafa R, Veronese A, Spizzo R, Pichiorri F, Liu CG, Calin GA, Croce CM, Negrini M (2007) mRNA/microRNA gene expression profile in microsatellite unstable colorectal cancer. Mol Cancer 6: 54
Longley DB, Harkin DP, Johnston PG (2003) 5-Fluorouracil: mechanisms of action and clinical strategies. Nat Rev Cancer 3: 330–338
Lord RV, Salonga D, Danenberg KD, Peters JH, DeMeester TR, Park JM, Johansson J, Skinner KA, Chandrasoma P, DeMeester SR, Bremner CG, Tsai PI, Danenberg PV (2000) Telomerase reverse transcriptase expression is increased early in the Barrett's metaplasia, dysplasia, adenocarcinoma sequence. J Gastrointest Surg 4: 135–142
Matherly LH, Goldman DI (2003) Membrane transport of folates. Vitam Horm 66: 403–456
Odin E, Wettergren Y, Nilsson S, Willen R, Carlsson G, Spears CP, Larsson L, Gustavsson B (2003) Altered gene expression of folate enzymes in adjacent mucosa is associated with outcome of colorectal cancer patients. Clin Cancer Res 9: 6012–6019
Ogino S, Cantor M, Kawasaki T, Brahmandam M, Kirkner GJ, Weisenberger DJ, Campan M, Laird PW, Loda M, Fuchs CS (2006) CpG island methylator phenotype (CIMP) of colorectal cancer is best characterised by quantitative DNA methylation analysis and prospective cohort studies. Gut 55: 1000–1006
Ogino S, Meyerhardt JA, Kawasaki T, Clark JW, Ryan DP, Kulke MH, Enzinger PC, Wolpin BM, Loda M, Fuchs CS (2007) CpG island methylation, response to combination chemotherapy, and patient survival in advanced microsatellite stable colorectal carcinoma. Virchows Arch 450: 529–537
Ougolkov AV, Yamashita K, Mai M, Minamoto T (2002) Oncogenic β-catenin and MMP-7 (matrilysin) cosegregate in late-stage clinical colon cancer. Gastroenterology 122: 60–71
Paz MF, Avila S, Fraga MF, Pollan M, Capella G, Peinado MA, Sanchez-Cespedes M, Herman JG, Esteller M (2002) Germ-line variants in methyl-group metabolism genes and susceptibility to DNA methylation in normal tissues and human primary tumors. Cancer Res 62: 4519–4524
Pufulete M, Al-Ghnaniem R, Khushal A, Appleby P, Harris N, Gout S, Emery PW, Sanders TA (2005a) Effect of folic acid supplementation on genomic DNA methylation in patients with colorectal adenoma. Gut 54: 648–653
Pufulete M, Al-Ghnaniem R, Rennie JA, Appleby P, Harris N, Gout S, Emery PW, Sanders TA (2005b) Influence of folate status on genomic DNA methylation in colonic mucosa of subjects without colorectal adenoma or cancer. Br J Cancer 92: 838–842
Qi H, Atkinson I, Xiao S, Choi YJ, Tobimatsu T, Shane B (1999) Folylpoly-gamma-glutamate synthetase: generation of isozymes and the role in one carbon metabolism and antifolate cytotoxicity. Adv Enzyme Regul 39: 263–273
Radparvar S, Houghton PJ, Houghton JA (1989) Effect of polyglutamylation of 5,10-methylenetetrahydrofolate on the binding of 5-fluoro-2′-deoxyuridylate to thymidylate synthase purified from a human colon adenocarcinoma xenograft. Biochem Pharmacol 38: 335–342
Rhee MS, Wang Y, Nair MG, Galivan J (1993) Acquisition of resistance to antifolates caused by enhanced gamma-glutamyl hydrolase activity. Cancer Res 53: 2227–2230
Romanini A, Lin JT, Niedzwiecki D, Bunni M, Priest DG, Bertino JR (1991) Role of folylpolyglutamates in biochemical modulation of fluoropyrimidines by leucovorin. Cancer Res 51: 789–793
Samowitz WS, Albertsen H, Herrick J, Levin TR, Sweeney C, Murtaugh MA, Wolff RK, Slattery ML (2005) Evaluation of a large, population-based sample supports a CpG island methylator phenotype in colon cancer. Gastroenterology 129: 837–845
Schneider E, Ryan TJ (2006) Gamma-glutamyl hydrolase and drug resistance. Clin Chim Acta 374: 25–32
Shen L, Catalano PJ, Benson III AB, O’Dwyer P, Hamilton SR, Issa JP (2007) Association between DNA methylation and shortened survival in patients with advanced colorectal cancer treated with 5-fluorouracil based chemotherapy. Clin Cancer Res 13: 6093–6098
Slattery ML, Curtin K, Sweeney C, Levin TR, Potter J, Wolff RK, Albertsen H, Samowitz WS (2007) Diet and lifestyle factor associations with CpG island methylator phenotype and BRAF mutations in colon cancer. Int J Cancer 120: 656–663
Sohn KJ, Stempak JM, Reid S, Shirwadkar S, Mason JB, Kim YI (2003) The effect of dietary folate on genomic and p53-specific DNA methylation in rat colon. Carcinogenesis 24: 81–90
Takemura Y, Kobayashi H, Miyachi H, Gibson W, Kimbell R, Jackman AL (1996) Biological activity and intracellular metabolism of ZD1694 in human leukemia cell lines with different resistance mechanisms to antifolate drugs. Jpn J Cancer Res 87: 773–780
Toyota M, Ahuja N, Ohe-Toyota M, Herman JG, Baylin SB, Issa JP (1999) CpG island methylator phenotype in colorectal cancer. Proc Natl Acad Sci USA 96: 8681–8686
van Engeland M, Weijenberg MP, Roemen GM, Brink M, de Bruine AP, Goldbohm RA, van den Brandt PA, Baylin SB, de Goeij AF, Herman JG (2003) Effects of dietary folate and alcohol intake on promoter methylation in sporadic colorectal cancer: the Netherlands cohort study on diet and cancer. Cancer Res 63: 3133–3137
van Rijnsoever M, Elsaleh H, Joseph D, McCaul K, Iacopetta B (2003) CpG island methylator phenotype is an independent predictor of survival benefit from 5-fluorouracil in stage III colorectal cancer. Clin Cancer Res 9: 2898–2903
van Rijnsoever M, Grieu F, Elsaleh H, Joseph D, Iacopetta B (2002) Characterisation of colorectal cancers showing hypermethylation at multiple CpG islands. Gut 51: 797–802
Weisenberger DJ, Siegmund KD, Campan M, Young J, Long TI, Faasse MA, Kang GH, Widschwendter M, Weener D, Buchanan D, Koh H, Simms L, Barker M, Leggett B, Levine J, Kim M, French AJ, Thibodeau SN, Jass J, Haile R, Laird PW (2006) CpG island methylator phenotype underlies sporadic microsatellite instability and is tightly associated with BRAF mutation in colorectal cancer. Nat Genet 38: 787–793
Acknowledgements
This work was supported in part by Grants-in-Aid for Scientific Research from the Japan Society for the Promotion of Science and by Cancer & Bowel Research Trust from Australia.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on British Journal of Cancer website (http://www.nature.com/bjc)
Supplementary information
Rights and permissions
From twelve months after its original publication, this work is licensed under the Creative Commons Attribution-NonCommercial-Share Alike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Kawakami, K., Ooyama, A., Ruszkiewicz, A. et al. Low expression of γ-glutamyl hydrolase mRNA in primary colorectal cancer with the CpG island methylator phenotype. Br J Cancer 98, 1555–1561 (2008). https://doi.org/10.1038/sj.bjc.6604346
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.bjc.6604346
Keywords
This article is cited by
-
Relationship between folate concentration and expression of folate-associated genes in tissue and plasma after intraoperative administration of leucovorin in patients with colorectal cancer
Cancer Chemotherapy and Pharmacology (2018)
-
Colorectal Cancer: Why Does Side Matter?
Drugs (2018)
-
A single-arm phase II trial of combined chemotherapy with S-1, oral leucovorin, and bevacizumab in heavily pre-treated patients with metastatic colorectal cancer
BMC Cancer (2015)
-
γ-Glutamyl hydrolase modulation significantly influences global and gene-specific DNA methylation and gene expression in human colon and breast cancer cells
Genes & Nutrition (2015)
-
DNA methylation and expression of the folate transporter genes in colorectal cancer
Tumor Biology (2015)