Abstract
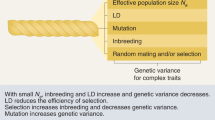
The variance components (VC) model has been popular for genetic analysis. It has received wide applications in a variety of genetic practices, and been extended to various forms for different settings. However, most of the existing VC models are, explicitly or implicitly, under the assumption of the Hardy–Weinberg and/or linkage equilibria, which is impractical in some realistic settings since more or less deviations from this assumption are common. We propose a new VC model that incorporates both these disequilibria, and includes the existing models as special cases. The corresponding variance components are computed for some commonly used relative pairs conditional on the observed marker identity-by-descent data. Parameters can be estimated by the traditional methods such as the maximum likelihood estimate. Simulation studies suggest that this extended model improves inference significantly over the existing models when deviations of these disequilibria are present.
Similar content being viewed by others
Introduction
The variance components(VC) models1, 2, 3, 4, 5, 6, 7, 8 has received much attention and wide applications in quantitative genetic trait studies, as this method requires few model assumptions. It has been extended to various forms for different data structures under different algorithms and model assumptions. Lange and Boehnke9 extended it to multivariate traits, Duggirala et al10 applied it to dichotomous traits, Amos et al11 studied the least squares algorithm of it. Andrade et al12 extended it to longitudinal pedigree data. This model and its variants have been used extensively in genetic linkage analysis. However, most of the existing VC models are, explicitly or implicitly, under the assumption of the Hardy–Weinberg and/ or linkage equilibria. These fundamental assumptions are sometimes not easy to justify, and in practice they are often more or less deviated. In linkage analysis the latter assumption may be inappropriate, since putative disease locus are usually in linkage disequilibrium(LD) with the flanking marker loci.13 Almasy et al14 proposed a combined linkage/disequilibrium analysis in which the LD are incorporated into the VC model. There are some VC models for combined linkage and association studies,15 a VC model incorporated with the two disequilibria is of practical meaning, and has not been in the literature. Here we consider such model in the settings of Hardy–Weinberg and/or LD, as an extension of the existing VC models. In our model the LD is parameterized via the trait-marker composite genotype, differently from that in Almasy et al14 in which the LD is parameterized via the trait-marker alleles. The correspondindg variance components are computed for some commonly used relative pairs conditional on the observed marker identity-by-descent (IBD) data. Parameters can be estimated by the traditional methods such as the maximum likelihood estimate (MLE) under the normal model assumption. This extended VC model is expected to have more accurate estimation of parameters, can be used for linkage and combined linkage and LD mapping (association study), using pedigree data, and have more power for such analysis.
The common VC model
We first describe the likelihood of the commonly used variance components model, for example as in Amos.5 Since the total likelihood is a product of likelihood over all the families under study, we only present the model for a given family for the sake of simplicity.
Let Yi be the trait value of the ith individual in the family.The VC model describing the trait value is

where μ is the overall mean, gi is the unobserved random major gene effect at the trait locus with alleles denoted by A and B, Gi is the unobserved polygenic effects,

where the ηj's are effects associated with the covariates xij's, and ei is the residual random error. The usual assumption is that gi, Gi and ei are uncorrelated and E(gi)=E(Gi)=E(ei)=0. Let p be the population proportion of allele A. Under the Hardy–Weinberg assumption one has E(gi)=a(2p−1)+2p(1−p)d=0. The covariance between individuals i and j is

where σa2=2p(1−p)[a−d(2p−1)]2 is the additive genetic variance due to the locus, σ2d=4p2(1−p)2d2 is the dominant genetic variance, Φij=Δ7ij/2+Δ8ij/4 is the kinship coefficient16 between individuals i and j, and Δ7ij, Δ8ij, Δ9ij are the condensed kinship coefficient,17 between individuals i and j. The Δkijs(k=1, …, 9) are the probabilities for the nine possible condensed IBD status as divided by Jacquard,17 in which Δ7ij, Δ8ij and Δ9ij are commonly used in practice. They are the population probabilities of sharing 2, 1 and 0 genes IBD for individuals (i, j), without regard to their particular genotypes, but only (i, j)'s kinship relationships, and under the Mendelian inheritance. Also, 2Φij is the expected proportion of gene IBD for individuals (i, j), at this locus.
For linkage analysis, usually IBD sharing data {πij} {πij=0, 1, 2} between a relative pair individual i and j, at marker locus is available, Amos5 proposed the following model for the conditional covariance

where θ is the recombination fraction between the trait and the marker loci. The values of f(θ, πij) and g(θ, πij) can be found.5 It is noted that g(θ, πij)=0 for most human relative pairs except full sibs and it's related to the possibility of sharing two allales IBD.
VC model with disequilibria
In this section we derive VC models with disequilibria in different settings, by incorporating these parameters into the covariances (2).
Hardy–Weinberg disequilibrium at trait locus
We first consider incorporating the Hardy–Weinberg disequilibrium at the trait locus into the VC model, without marker information Let Ak denote allele k at the trait locus (k=1, …, K), pk its proportion in the population, Pkl the corresponding proportion of the genotype AkAl. One way to deal with the deviation from the Hardy–Weinberg assumption is the use of the within population inbreeding cofficient18, 19 f at the trait locus, which is the odds that at any gene, both alleles of the gene pair were inherited from the same ancestor. Let I(·) be the indicator function. Given f we have

Here 0≤f≤1, and f=0 corresponds to Hardy–Weinberg equilibrium. Let p(kl)(km) be the conditional probability that two individuals have genotype (AkAl, AkAm) or (AlAk, AmAk) at the trait locus given that they share Ak IBD (Assuming random mating and phase known, these are the only cases they share Ak IBD. The possibilities for the cases AkAl, AmAk or (AlAk, AkAm) are negligible). Let Y be the trait value of a general individual and g be his/her genotype, and μkl=E(Y∣g=AkAl). Following Fisher1 and Lange,16 let αk's be the optimal additive genetic effects in the sense that they minimize the sum of squared residuals ΣkΣlδ2klpkl, where δkl=μkl−αk−αl. We show in Appendix A that

and

where γ7(f)=(1+(f/2))σ2a+(1−f)σ2d+fσ20, γ8(f)=((1+f)2/2)σ2a, σ2a=2Σkα2kpk, σ2d=ΣkΣlδ2klpkpl, σ20=Σkδ2kkpk is the part of variance explained by the optimal additive genetic effects, and αk=Σlμklpkl/[(1+f)pk] for all k.
Note that if f=0, (6) reduces to (2). The αk's and is δkl's are the optimal additive major gene effects and the residual effects.16
Linkage to marker
Now we consider the case with marker information available in addition to the trait locus data. Let πij(=0, 1, 2) to be the number of IBD allele sharing between individuals i and j at the marker locus, π′ij be the corresponding unobserved number at the trait locus, and θ be the recombination fraction between the two loci. Expressions for Cov(Yi, Yj∣πij=k) can be found by the formula

Usually, for each individual the IBD data πij is not directly available. However, their probabilities P(πij=k)(k=0, 1, 2) can be computed from the corresponding observed marker genotypes. So the covariances between individual pair (i, j) in a given family is

Covariance with Hardy–Weinberg disequilibrium at trait given marker IBD
In the previous section, we derived the variance components under Hardy–Weinberg equilibrium at the trait locus. Here we give these componenets with the linked marker information, that is, conditional on the trait-marker IBD data. In this case the variance components are

where Δ7ij(πij)=P(π′ij=2∣πij), Δ8ij(πij)=P(π′ij=1∣πij) and Δ9ij(πij)=P(π′ij=0∣πij) are the conditional IBD sharing at the trait locus given the IBD sharing at the marker locus, for individuals (i, j). The derivation is the same as that for Cov(Yi, Yj∣f) with Δ7ij and Δ8ij replaced by Δ7ij(πij) and Δ8ij(πij), whose values are obtained from the relationships

and the known values of P(π′ij=0, πij) as listed in the literatures cited before. Note here given π′ij=0, Yj and Yj are independent, and Cov(Yi, Yj∣f,π′ij=0)=0, thus we don't have the term for Δ9ij(πij).
Since in real data the set {πij} is unobservable, we only have the computed the set of probabilities {P(πij=k)}, thus the covariance is

Covariance with LD between trait and marker
In linkage analysis, LD between the trait locus and the genotype marker locus should be taken into consideration. In this section we compute the covariances between relative pairs when in addition to the case of LD is also present between the trait and marker loci. Let ak and akal denote the alleles and genotypes at the marker locus, qk and qkl be the corresponding population frequencies. Since the within-population inbreeding coefficient f is common for any locus in the genome of the given population, f describes the relationship between the marker genotype frequencies qks allele frequencies qkls, in the same way as it did between the pks and pkls at the trait locus. That is, we have

Let  be a general notation for the trait-marker composite genotype. We assume
be a general notation for the trait-marker composite genotype. We assume

It is easy to check that under (11), ΣkΣlp(kl, rs)=prs and ΣrΣsp(kl, rs)=pkl,, the probabilities of composite genotypes satisfy such consistent condition with its marginal probabilities. Here 0⩽ζ⩽1 is the LD parameter, and it should not be confused with the definition of LD that is used in some texts, such as in Weir20 or Almasy et al.14 Note that ζ=0 corresponds to linkage equilibrium. Also, ζ manifests the vertical connection between the trait and marker loci, while the recombination fraction describes the horizontal link between the alleles.
For a relative pair, let p(kl)(mn)∣π′ij=P(AkAl, AmAn∣π′ij) be the conditional probability that individual i has trait genotype AkAl and individual j has trait genotype AmAn given their IBD value π′ij at this locus, pklm∣π′ij=½P(AkAl, AkAm∣π′ij)+½P(AkAl, AmAk∣π′ij) be the probability when they also share one allele identical by state (IBS) at the trait; pkl∣π′ij=P(AkAl, AkAl∣π′ij) be the probability when they share both alleles IBS at the trait locus. We have (Appendix B)

where γ7(f, ζ, πij), γ8(f, ζ, πij) and γ9(f, ζ, πij) denote respectively Cov(gi, gj∣f, ζ, πij, π′ij=2), Cov(gi, gj∣f, ζ, πij, π′ij=1) and Cov(gi, gj∣f, ζ, πij, π′ij=0). Note that by conditioning on the IDB values at both the trait and marker loci, we cannot assert Cov(gi, gj∣f, ζ, πij, π′ij=0)=0 as we did for the previous section. We have γ7(f, ζ, πi,j)≡γ7(f),

and

where

which is also written as

Since the genetic covariance between the relative pair can be written as

by (12), when π′ij=2 or πij=0, the expression for genetic variance between a relative pair is the same regardless LD is present or not. In fact, from the derivation in Appendix B, this conclusion is true for any consistent composite genotype specification: under random mating and any consistent specification P(G) of the composite genotype, the IBD status (π′ij, πij) of a relative pair (i, j) contributes information of LD to their genetic variance at the trait locus only if π′ij≤1 and πij≥1.
Again in practice, given the estimated IBD probabilities, the covariance is computed as

Parameter estimation
Let β=(μ, η1, …, ηj)T be the parameters in the mean, α=(θ, f, ζ, σ2a, σ2d, σ2G, σ2e, σ20, σ21, σ22, σ23, σ11, σ12, σ13)T be the parameters in the covariance matrices, yk be the observations of all the members in the kth family, and μk=μk(β)=E(Yk)=Xkβ, where Xk is the covariate matrix for the kth family, and nk is the total number of individuals in this family. The commonly used model based estimation method is MLE, while the common model for quantitative trait is the normal distribution. Under these assumptions, the likelihood of the kth family is Lk(α, β∣Yk)=φ(Yk−μk∣Ωk), where φ(Y−μ∣Ω) is the density of the nk dimensional normal N(μ, Ω) distribution,  is the covariance matrix of the kth family, with
is the covariance matrix of the kth family, with

as specified in (12) in the most general case. The P(πij=r∣gij)'s can be obtained by some common IBD computation methods. The covariances can also take any of the more specific form (8), (6), (3) and (2) in the equilibrium case. Here we used (Yi, Yj) for (Yki, Ykj), the (i, j)th relative pair in the kth family. The total likelihood is thus L(α, β∣Y)=Πkk=1Lk(α, β∣Yk), and the log-likelihood, omitting the normalizing constant, is

The MLE is the parametric value (α̂, β̂) that maximizes (14), and it has many desired optimality properties.
Power
The power of the method can be easily estimated and will shown is dependent only on the parameters α in the covariance matrix. Let H0:α=α0 and H1:α=α1 (H0⊂H1 or α0 be part of α1) be the null and alternative hypothesis considered in the previous sections, dim(H1)−dim(H0)=k and f(·∣α, β) be the density of the model considered. Let α̂0 and α̂1 be the MLE of α under H0 and H1, respectively. Note our hypothesis only involves α, not the parameters β in the mean specification. Let

be the relative entropy (Kullback–Leibler divergence) between the two densities f(·∣α1, β1) and f(·∣α0, β0). It is known that D(α1∣∣α0)≥0 with equality hold if and only if α1=α0. Assuming homogenous familial structures for all the families, for give level γ>0, the asymptotic power qn for the likelihood ratio test of H0 vs H1, with a dataset of size (number of families) n, is (Appendix C)

where Vk is the χ2 random variable with k degrees of freedom and χ2k(1−γ) is its 1−γ upper quantile.
Given f(·∣·, ·), α1 and α0, D(α1∣∣α0) can be easily computed. In fact, since our model f(·∣·, ·) is multivariate normal, it is easy to see that

where d=dim(μ), Ω(·) is the Ωk's with the elements given in (12), in which the τij's take the theoretical mean values. To plot the power surface, we fix the parameter values at their MLE, except those for f and ζ. Then for a given γ>0 and a set of selected (f, ζ) values, we can compute qn=qn(γ, f, ζ) for different γ, f, ζ and n.
Application
Simulation study
Data of 10 000 sibpairs are simulated in our study. We give some detailed description of how the two levels of disequilibria are incorporated in the simulation process. It can be described in the following three steps.
Step 1
For each sibpair we simulate the their trait genotypes gis and the marker IBD probabilities πijs. Let Gi=(aras)/(AkAl) be the composite genotype of the trait and marker for the ith individual, with lower case letters aras for marker genotype. we simulate (Gi, Gj) for each sibpair, and πij is generated along. We first generate the composite genotypes Gf of the father and Gm of the mother by the probability given in (11) with ζ=0.1, and pkl and qrs are given (4) and (10) with f=0.12, p1=0.55, p2=0.45, q1=0.65 and q2=0.35. Although (Gf, Gm) are not part of the data to be used in the computation, they are needed to generate the sibs composite genotypes. Now given (Gf, Gm) we generate Gi, Gj and πij as below. Let Gf=(af1 af2)/(Af1 Af2), Gm=(am1 am2)/(Am1 Am2). During meiosis, if there is no recombination (with probability 1−θ, θ=0.25), Gf splits into two gametes (af1/Af1) and (af2/Af2). Then one of the gametes is selected with probability 0.5 to pass to the next generation. Here we only consider the recombination at the marker, since we want the IBD πij at the marker. The recombination at the trait is similar, and we omit it for simplicity, since this will not affect the probabilities of the Gis. Similarly, Gm will split into (am1/Am1) and (am2/Am1), or (am2/Am1) and (am1/Am2), and one of the gamets is selected with probability 0.5 to pass to the next generation. For example, if for the father, there is recombination during meiosis and (af1/Af1) is selected, and for the mother there is no recombination during meiosis and (am1/Am1) is selected, then Gi=(af1 am1)/(Af2 Am1) and gi=(Af2Am1). Repeat the above process to get, say, Gj=(af2 am1)/(Af1 Am1) and gj=(Af1Am1). Since at the marker locus, sibpair (i, j) has a composite genotype (af1am1, af2am2), we have πij=1, which comes from the common maternal allele am1.
Step 2
Simulate each pair's covariates. The mean μI of the ith individual is given by (1). Specifically, we take μ=23, gi=1 if individual i has genotype A1A1=0 if A1A2, and =−1 if A1A2. We take Gi∼N(0,σ2G) with σ2G=0.2. Two covariates are genetated, xi1 and xi2, stand for age (years) and sex index for the ith individual, xi2=1 for female and =0 for male. The coefficient for age is η1=0.2 and that for sex is η2=1.5. ei is the random error from N(0, 1) distribution.we always assume the first dib is younger with xi1∼U[10, 60], then for the second sib, with xj1=xi1+z with z∼U[1, 10]. For xi2, using the gender ratio from the real data, we sample z∼U(0, 1), if z≤0.54 let xi2=1 (female) otherwise 0 (male).
Step 3
Simulates the sibpair covariance matrices Ωij=Cov(Yi, Yj)=(ωij) and the final observed trait values. By (3.9), ω11=ω22=(1+f/2)σ2a+(1−f)σ2d+fσ20+σ2G+σ2e, σ2a=2Σkα2kpk, α2d=Σk,lδ2klpkpl, σ20=Σkδ2kkpk and pk is the population proportion of allele Ak, δkl=μkl−αk−αl, αk=Σlμklpkl/[(1+f)pk], pkl=(1−f)pkpl+fpkI(l=k) is the population proportion of genotype AkAl, and μkl=E(Y∣g=AkAl)=μ+gkl+ηlE(xi1)+η2E(xi2)=μ+gkl+40η1+0.54η2,  as is given in (3.9), and for sibpairs Φij=1/4. Δkij(πij) is defined after (8) and can be found in Wright,18 where they are implemented in terms of the recombination fraction θ. The marker IBD data πijs are generated above, the trait IBD π′ij are unknown, but only the conditional probability P(π′ij∣πij)s are used, which are easily derived.20 The γk(f, ζ, πij)s are defined after (12). The definition of σ1,2 involved p(kl)(km) which is given in the definition of the γk(f, ,ζ, πij)s. Now we have implemented Ωij and are ready to simulated the yis. We simulate the data pairwise. For a sibpair (yi, yj), denote Y=(yi, yj) and μ=(μi, μj). We sample Z∼N(0, I2), the two-dimensional standard normal distribution, and let
as is given in (3.9), and for sibpairs Φij=1/4. Δkij(πij) is defined after (8) and can be found in Wright,18 where they are implemented in terms of the recombination fraction θ. The marker IBD data πijs are generated above, the trait IBD π′ij are unknown, but only the conditional probability P(π′ij∣πij)s are used, which are easily derived.20 The γk(f, ζ, πij)s are defined after (12). The definition of σ1,2 involved p(kl)(km) which is given in the definition of the γk(f, ,ζ, πij)s. Now we have implemented Ωij and are ready to simulated the yis. We simulate the data pairwise. For a sibpair (yi, yj), denote Y=(yi, yj) and μ=(μi, μj). We sample Z∼N(0, I2), the two-dimensional standard normal distribution, and let  , and simulate such Y 10 000 times.
, and simulate such Y 10 000 times.
For γ8(f, ζ, πij) in te case πij=2, σ1, 1, σ1, 2 and σ1, 3 are not independently estimable, so in this case we write γ8(f, ,ζ, 2)=(1−ζ)γ8(f)=σ42, where σ42=−ζ(1+f)σ1, 1+2ζσ1, 2+ζ2σ1, 3 viewed as a single parameter to be estimated.
Table 1 displays the values of the real parameters of interest from the simulation, and their MLE estimates (estimated standard deviation in bracket) under H0: f=ζ=0.0 and H1: all parameters free, respectively.
The difference 2(log likelihood(H1)−log likelihood(H0))=20.9934, with a P-value of 0.000106 under a χ2 distribution with two degrees of freedom, that is, the evidence of rejecting H0 is very strong. This example shows that incorporating the disequilibria mechanism into the variance components model can improve the inference significantly when such disequilibria are present.
Real data application
We used the AADM data (African-American Diabetes mellitus) to illustrate the method. The data is from an international collaboration between West Africa and US investigators in mapping type II diabetes susceptibility genes in West African ancestral populations of African-Americans. Affected sib-pairs along with unaffected spouse controls were being enrolled. Eligible participants were invited to study clinics to obtain detailed epidemiological, familial and medical history information. For detailed description of the data, see Rotimi et al.21 For this data we computed the model parameter estimates using VC model (2), or under the hypothesis of equilibria, H0: f=ζ=0; and under the VC model with Hardy–Weinberg/LD (12), H1:f and ζ are free parameters, to fit the data. The response variable is BMI, the covariate is age. The results are shown in Table 2, where the estimated standard deviations are listed inside the brackets.
The −2 loglikelihood difference is 12.5076 with a P-value of 0.0058, which is highly significant. So the inference should be based on H1. We see a large Hardy–Weinberg disequilibrium at the triat locus, suggesting that the genetic background of the sample under study is not as simple as assumed by the existing VC model. The low recombination rate (0.0016) indicates that the trait and marker loci are tightly linked, and the LD between the trait and marker is non-negligible. The overall BMI of this sample is 23.58, and the age effect is 0.053, which are quite common for normal populations.
The power depends on all the parameters in the model, we highlight its dependence on (f, η) to study its relationship with these two parameters. Using (15) and the parameters above, the following Figure 1 shows the powers of the likelihood ratio test for H0 vs H1, for various combinations of f, ζ, and n.
Powers of the likelihood ratio test for H0 vs H1. (a)–(c) Power for the real data, with parameters set at the MLE values. (a) H0: f=0 vs H1: f≠0. Horizontal axis for f, vertical axis for power, n=8 000 000; (b) H0: ζ=0 vs H1: ζ≠0. Horizontal axis for ζ, vertical axis for power, n=675; (c) (the third on the first row): H0: (f, ζ)=(0, 0) vs H1: (f, ζ)≠(0, 0). n=675. (d)–(f) Power for the simulated data. The parameters used are α=(σ4, σ3, σ2, σg, σ0, σa, σd, θ)=3.08, 16.3, 43.5, 0.45, 21.8, 7.24, 20.8, 0.2468), sample size is n=250 for the three panels. (d): H0: f=0 vs H1: f≠0; (e) H0: ζ=0 vs H1: ζ≠0; (f) (the third on the second row): H0: (f, ζ)=(0, 0) vs H1: (f, ζ)≠(0, 0).
Since the LD depends on the unobservable trait genotype, its needs larger sample size to detect. For the real data, with the observations and the estimated parameter setting, it is easy to detect the HWE disequilibrium with reasonable sample size, while it is very difficult to detect the LD, or requires very large sample size to achieve high power. For the simulated data-parameter setting, the powers are high for the joint HWE disequilibrium, LD and the joint HWE and LD disequilibria.
The software for this extended VC model is written in SAS; the current version is for sibpair familial structure only, and is available upon request from the second author at gchen@genomecenter.howard.edu. The CPU time to compute the parameter estimates depends on the machine, data size, number of regressors, pedigree structure and starting values for the parameters etc. For the two examples above, with suitably chosen starting values, the CPU times for computing the MLEs are 27.24 and 27.33 s on our machine.
Discussion
We have generalized the VC model to the cases of the Hardy–Weinberg and LD or both, this gives more practical application of this popular model. In some practices, these disequilibria are not justified. In these cases, the existing VC model is clearly inadequate, and our generalized VC model might be beneficial in more estimates, and in enhancing the inference power of parameters of interest. Also this generalized model can be used in testing these disequilibria by forming the corresponding likelihood ratio statistic, along with the parameter estimates. Other inferences on one or both of the two disequilibria are sometimes also of direct interest, which are now available under this generalized VC model.
We computed the variance components for some common relative pairs. The cases of other relative pairs are similar and straightforward. We considered the parameter estimation in several ways and computed the IBD under some common cases.
Further extensions/modifications to implement more features will be similar, such as the multivariate traits,9 the multipoint VC, dichotomous trait, robust LOD score correction,7 the conditioning adjustment.21
References
Fisher RA : The correlation between relatives on the supposition of mendel inheritance. Trans Roy Soc Edinburgh 1918; 52: 399–433.
Harris DL : Genotypic covariances between inbred relatives. Genetics 1964; 50: 1319–1348.
Amos CI, Elston RC : Robust methods for the detection of genetic linkage for quantitative data from pedigrees. Genet Epidemiol 1989; 6: 306–349.
Goldgar DE : Multipoint analysis of human quantitative genetic variation. Am J Human Genet 1990; 47: 957–967.
Amos CI : Robust variance-components approach for assessing gene linkage in pedigrees. Am. J Human Genet 1994; 54: 535–543.
Almasy L, Blangero J : Multipoint quantitative-trait linkage analysis in general pedigrees. Am J Human Genet 1998; 62: 1198–1211.
Blangero J, Williams JT, Almasy L : Robust LOD score for variance component-based linkage analysis. Genetic Epidemiol 2000; 19 (Suppl 1): S8–S14.
Sham PC, Purcell S : Equivalence between Haseman–Elston and variance-components linkage analysis for sib pairs. Am J Human Genet 2001; 68: 1527–1532.
Lange K, Boehnke M : Extensions to pedigree analysis. IV covariance components models for multivariate traits. Am J Med Genet 1983; 14: 513–524.
Duggriala R, Williams JT, Williams-Blangero S, Blangero J : A variance component approach to dichotomous trait linkage analysis using a threshold model. Genet Epidemiol 1997; 14: 987–992.
Amos CI, Gu X, Chen J, Davis BR : Least squares estimation of variance components for linkage. Genetic Epidemiol 2000; 19 (Suppl 1): S1–S7.
Andrade M, Gueguen R, Visvikis S, Sass C, Siest G, Amos C : Extension of variance components approach to incorporate temporal trends and longitudinal pedigree data analysis. Genetic Epidemiol 2002; 22: 221–232.
Xiong M, Jin L : Combined linkage and linkage disequilibrium mapping for genome screens. Genetic Epidemiol 2000; 19: 211–234.
Almasy L, Williams J, Dyer T, Blangero J : Quantitative trait locus detection using combined linkage/disequilibrium analysis. Genetic Epidemiol 1999; 17 (Suppl. 1): S31–S36.
Fulker DW, Cherny SS, Sham PC, Hewitt JK : Combined linkage and association sib-pair analysis for quantitative traits. Am J Human Genet 1999; 64: 259–267.
Lange K : Mathematical and statistical methods for genetic analysis. Berlin: Springer-Verlag, 1997.
Jacquard A : The genetic structure of populations. New York: Springer-Verlag, 1974.
Wright S : The genetic structure of populations. Ann. Eugenics 1951; 15: 323–354.
Cockerham CC : Variance of gene frequencies. Evolution 1969; 23: 72–84.
Weir B : Genetic Data Analysis II. Sinauer Associates, Inc. Publishers: Sunderland, Massachusetts, 1996.
Rotimi CN, Dunston GM, Berg K : In search of susceptibility genes for type 2 diabetes in West Africa: The design and results of the first phase of the AADM study. Ann Epidemiol 2001; 11: 51–58.
Lange K : Central limit theorems for pedigrees. J Math Biol 1978; 6: 59–66.
Acknowledgements
We appreciate the suggestions/comments from the Editor and the two anonymous reviewers, which greatly improved the quality of this manuscript. The work was supported by the United States Public Service Grant No AG 16996 and the National Center for Research Resources Grant No 2G12RR003048 from the National Institutes of Health. The AADM study was supported by NIH Grants no. 3T37TW00041 from NCMHD and NHGRI. G Chen and Rotimi were also partly supported by the NIGMS/MBRS program.
Author information
Authors and Affiliations
Corresponding author
Appendices
Appendix A
We first derive (5). When fixed Ak, the events AkAl and AkAm are independent, so by (4) we have,

For (6), we use the method as in Lange16 (pp. 87–89). Since  and the αk's minimize the squared error
and the αk's minimize the squared error  , take derivative with respect to αk, we get
, take derivative with respect to αk, we get

Sum over k in (A.1) we have

Now (A.1) and (A.2) gives

that is,

Then we have

When i=j, Cov(Gi, Gj)=σG2, Cov(ei, ej)=σe2 and

By (A.1), the above is

Since ∑lpkl=∑lplk=pk, we have

and

so by (A.3) and the above three equations we have

If i≠j, Cov(eiej)=0, by the central limit theorem of Lange22 and assume no dominance, we have approximately Cov(Gi, Gj)=2ΦijσG2 and

By (A.1) and (A.2), the coefficient Δ9ij is zero. By the calculation for E(gi2), the first term above is

the second term is

From (5) it is easy to check that

so by (A.2) the coefficient of Δ8ij is

Since ∑k∑lαkαlpkl=fσa2/2, the above is

By (5) and (A.1), the middle term in the above is

By the same way, the last term in (A.6) is

so the coefficient of Δ8ij is

Now collecting terms we have

Appendix B
When i=j, πii′=2 which is noninformative about trait-marker relationship, so  , which has the same expression as in (8). When i≠j,
, which has the same expression as in (8). When i≠j,

We first derive the conditional probabilities  in (B.1). Since conditioning on the IBD status, those quantities are independent of relatedness of the pair, only depend on the relationships among the trait and marker alleles through f and ζ, in other words, given IBD status, different alleles in one configuration are independent with those in the other one. We have
in (B.1). Since conditioning on the IBD status, those quantities are independent of relatedness of the pair, only depend on the relationships among the trait and marker alleles through f and ζ, in other words, given IBD status, different alleles in one configuration are independent with those in the other one. We have

Now the two configurations share AkAl in common, if we fix it, the two configurations are independent each other, so we rewrite the above as

Similarly,



and

where p(kl, r)=∑sp(kl, rs). The same reason gives

so we have

Also

where

So


and

so

Also

and

where p(k, rs)=∑lp(kl, rs)=pk(qrs−ζprs+ζ(1−f)psI(r=k)+ζI(r=s=k)), so

Lastly

Now we compute the covariance (B.1) for different values of the π′ij's. If π′ij=0, by (B.2)–(B.4) and Appendix A, we have the same expression of (B.1) as in (8).
If π′ij=1, by (B.5)–(B.7), the coefficient of Δ7ij(πij) and Δ8ij(πij) in (B.1) are the same as that in (A.4) and (A.7); the coefficient of Δ9ij(πij) in (B.1) has four terms corresponding to those in (B.7), the first two terms are zero by the computation in Appendix A, by its symmetry in (k, l, m, n), the last two terms are

since

the first term above is zero. By expanding and check each term using (A.1) and (A.2), the second term above, and hence the coefficient of Δ9ij(πij) is

If π′ij=2, by (B.8), the coefficient of Δ7ij(πij) is the same as before. Now we compute the coefficient of Δ8ij(πij). We expand it in five terms as in (B.9). The first term is (1+f)2(1−ζ)σa2/2 by the computation in Appendix A. Expanding the same way as in Appendix A, the second term is

the last two terms above are zero by (A.1). Since

substitute this into the second and the third term in (B.12), it becomes −(1+f)2∑kαk2pk(qk−ζpk). By expanding the same way, the third term is

The last two terms in (B.14) are zero. Substitute (B.13) into the second and third term in (B.14), it becomes

now combine the second and the third terms gives

the fourth term is

the fifth term is

For the coefficient of Δ9ij(πij), we expand it in four terms according to (B.10), the first two terms are zero by the computation in Appendix A, so it reduces to

the first term above is zero since  , the second term, and hence the coefficient of Δ9ij(πij) is
, the second term, and hence the coefficient of Δ9ij(πij) is  , which is
, which is

The first term in the bracket above is

the second term is

the third term is

the fourth term is

Now collect terms, the coefficient of Δ9ij(πij) is

Appendix C
Let ξ=(α, β), ξ0=(α0, β), and define ξ1 and the hat notations for the corresponding estimates. Let Î(ξ) be the empirical Fisher information matrix evaluated at (ξ), by Taylor expansion,

and its is well known that, under H1, as n → ∞,

Also, since the familial structures are homogeneous, so

Thus under H1,

Rights and permissions
About this article
Cite this article
Yuan, A., Chen, G., Yang, Q. et al. Variance components model with disequilibria. Eur J Hum Genet 14, 941–952 (2006). https://doi.org/10.1038/sj.ejhg.5201645
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.ejhg.5201645