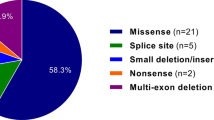
Abstract
Marfan syndrome (MFS; OMIM#154700) is a connective tissue disorder characterized by manifestations in the ocular, skeletal and cardiovascular systems. MFS is caused by mutation in the fibrillin-1 gene (FBN1; OMIM#134797) and more than 550 mutations have been identified so far. FBN1 is ∼230 kb in size and contains three evolutionarily conserved alternatively spliced exons B, A and C at the 5′end. In a first systematic attempt to associate sequence variations in the FBN1 5′ alternatively spliced exons with MFS, we investigated 41 individuals fulfilling the diagnostic criteria of Ghent nosology or with features of MFS including at least one major criterion or involvement of two organ systems but not fulfilling a strict interpretation of the Ghent nosology, and known to be negative for mutations in the FBN1 exons 1–65 as well as the TGFBR2 and TGFBR1 coding regions. We identified five novel and one previously reported variants in the six unrelated probands and provide preliminary evidence for their role in pathogenesis.
Similar content being viewed by others
Introduction
Fibrillin-1 is a 350-kDa extracellular matrix protein, a principal constituent of the 10-nm microfibrils, which are associated with maintenance of elastic fibers and anchoring epithelial cells to the interstitial matrix.1, 2, 3 This gene is ∼230 kb in size and is located on chromosome 15q21.4, 5 Mutations in the Fibrillin-1 (FBN1; OMIM#134797) coding regions cause Marfan syndrome (MFS; OMIM#154700), and more than 550 mutations have been identified so far.6 It is also established that FBN1 gene has alternatively spliced exons, B, A and C and a common exon M at the 5′ end (Figure 1),5 but little is known about the promoter region and any role of the 5′ alternatively spliced exons in transcriptional regulation and translation. It has been shown, however, that the FBN1 5′ alternatively spliced exons are evolutionarily conserved, suggesting that this region may have important regulatory functions and that mutations therein may play a role in MFS and/or other phenotypes.4 Recently, mutations in TGFBR1 and TGFBR2 have been reported in patients with MFS type 2 (MFS2; OMIM#154705)8 and a related disorder, Loeys–Dietz syndrome (LDS; OMIM#609192).
Schematic representation of human FBN1 upstream exons.7 Exon B, A and C are alternatively spliced exons. The presumptive initiation codon is indicated by ATG in exon M, which is being considered as the beginning of the exon 1 in the FBN1 mutations database (http://umd.be:2030/). The sequence homology between human, murine and porcine is given above each exon/intron.
In order to investigate the possible association between mutations in the 5′ alternatively spliced exons of FBN1 and MFS, we selected a cohort of 41 individuals fulfilling the diagnostic criteria of Ghent nosology or with features of MFS including at least one major criterion or involvement of two organ systems but not fulfilling a strict interpretation of the Ghent nosology,9 and negative for mutations in FBN1 exons 1–65 as well as the TGFBR2 and TGFBR1 coding regions, and 153 healthy controls.
We here report a total of five novel and one previously described sequence variants in the 5′ upstream region of the FBN1 gene in a total of six unrelated probands.
Materials and methods
In total, 41– unrelated individuals, including 35 German, four Swedish, one Swiss and one Turkish patients, fulfilling the diagnostic criteria of Ghent nosology or with features of MFS including at least one major criterion or involvement of two organ systems and not fulfilling a strict interpretation of the Ghent nosology, but without mutations in FBN1 exons 1–65 detectable by SSCP10 or in the TGFBR2 or TGFBR1 coding regions by sequencing (unpublished data), were subjected to mutation screening of 5′ alternatively spliced exons B, A and C of the FBN1 gene. A total of 153 randomly selected healthy blood donors were included as controls and subjected to the same mutation screening as the patients. Genomic DNA was extracted from blood using standard protocols. Primers were designed based on human sequence (accession number L19896) for amplification of ∼1.6 kb 5′ alternatively spliced exons B, A and C of the FBN1 gene (Table 1). Standard PCR conditions were initial denaturation at 95° for 10 min, followed by 33 cycles of 96°C for 1 min, 60–65°C for 1 min and 72°C for 1 min with final elongation for 10 min at 72°C in a 50-μl reaction mixture, containing 1 × buffer (Qiagen, Germany), 1 × Q solution (Qiagen, Germany), 20 pM each primer and 2.5 U Taq Polymerase (Qiagen, Germany). PCR products were purified with ExoSAP-IT (USB, USA) and both strands were sequenced with BigDye Terminator chemistry version 1.1 by standard protocol (ABI, USA). Sequencing reactions were carried out at 96°C for 10 s, 50°C for 5 s and 60°C for 4 min (25 cycles) (Biometra, Germany). The reaction mixtures were purified using DyeEx™ 2.0 Spin Kit (Qiagen, Germany) and analyzed on the ABI Genetic Analyser 3100 according to the supplier's instructions with the sequence analysis software (ABI, USA). Cloning was performed in TOPOTA cloning vector (Invitrogen, Germany) and the clones were sequenced. A search for transcription factor binding site was performed using http://www.cbrc.jp/research/db/TFSEARCH.html. Variants found upstream and downstream to exons are marked with ‘–’ and ‘+’ signs, respectively.
Results
A total of six, five novel and one previously reported, sequence variants were identified in six out of 41 individuals fulfilling the diagnostic criteria of Ghent nosology or with features of MFS including at least one major criterion or involvement of two organ systems but not fulfilling a strict interpretation of the Ghent nosology.
An A>G nucleotide replacement, 12 bp upstream to exon B (exon B, −12A>G) and an insertion of T at position 112 of exon B (exon B, 112insT) were found in a 17-year-old Swiss boy with dilatation of ascending aorta as one major criterion. The skeletal system was involved exhibiting arm span to height ratio of 1.06, positive wrist and thumb signs, pes planus, funnel chest and joint hypermobility. In the ocular system, divergent strabismus and a wide interpupillary distance were determined. As the family history is unknown and dural ectasia was not assessed, he did not fulfil the diagnostic criteria of Ghent nosology. Cloning of the PCR products showed that both variants occurred in cis. None of these variants was present in the 306 control chromosomes.
An exon A, +1G>A nucleotide replacement was found in a 19-year-old male proband of Swedish origin, who had enlargement of aortic root. The skeletal system was involved and displayed marfanoid body proportions, chest deformity, scoliosis, joint hypermobility and high arched palate with crowding of teeth. The ophthalmologic examination revealed no pathologic result except myopia. As a familial case, he fulfilled the diagnostic criteria of Ghent nosology. This variant occurred at the splice donor site of exon A, which has been found to be conserved in mouse (accession number L29454) and pig (accession number NM_001001771). This variant was not seen in the 306 control chromosomes.
Three further variants, intron C, +9C>T, intron C, +30C>T and intron C, +48 T>C were found in three unrelated probands. Intron C, +9 C>T was seen in one further proband, in whom the other two variants, intron C, +30 C>T and intron C, +48 T>C were absent. Cloning of the respective PCR products revealed that the three variants, if present, occurred on the same allele in these probands. These three variants occurred at a position, which is conserved between man (accession number L19896) and pig (accession number NM_001001771). Intron C, +9C>T and intron C, +30C>T were not seen in the 306 control chromosomes. The frequency of intron C, +48T>C did not differ between probands (allele frequency 0.25) and healthy controls (allele frequency 0.25).
The first proband identified with intron C, +9C>T, intron C, +30C>T and intron C, +48 T>C was a 28-year-old male of German origin with typical marfanoid skeletal features (reduced upper to lower segment ratio, positive wrist and thumb signs, scoliosis, highly arched palate) and facial appearance with downslanting palpebral fissures. If protrusio acetabuli had been ascertained on radiographs, the major criterion might be fulfilled in the skeletal system. The cardiovascular system was involved showing a mitral valve prolapse as a minor criterion. The ocular system was not involved. As a sporadic case, he did not fulfil the diagnostic criteria of Ghent nosology.
The second proband with intron C, +9C>T, intron C, +30C>T and intron C, +48T>C was a 60-year-old female of German origin with a negative family history, who underwent surgical replacement of ascending aorta and aortic valve because of dilatation of ascending aorta at the age of 48 years. The skeletal system was involved and showed scoliosis, pectus excavatum and highly arched palate. Her height was 181 cm with normal body proportions. The ocular system was not involved. This proband did not fulfil diagnostic criteria of Ghent nosology.
The third proband with intron C, +9C>T, intron C, +30C>T and intron C, +48T>C was a 46-year-old male proband of German origin, who underwent surgical replacement of ascending aorta because of dilatation. Furthermore, he had mitral valve prolapse, calcification of the mitral anulus and the mitral valve was replaced because of insufficiency. The skeletal system was involved, presenting pectus carinatum, highly arched palate and typical facial appearance. He had a height of 203 cm, but body proportions were not strikingly marfanoid. Striae atrophicae were present. The ocular system was not affected. As his mother presented with mitral valve prolapse with slight regurgitation and softening of the connective tissue as the only symptoms, she did not fulfil diagnostic criteria for MFS. As a sporadic case, he himself also did not fulfil the criteria.
The German proband with the variant intron C, +9C>T was a 47-year-old male with unknown family history. He had severe stenosis of the aortic valve, which was replaced at the age of 22 years. His mitral valve was replaced because of regurgitation at the age of 40 years. Dilatation of ascending aorta was diagnosed at the same time. The skeletal system fulfilled the major criterion exhibiting pectus carinatum, an arm span to height ratio of 1.06, scoliosis, reduced extension of the elbows, highly arched palate and typical facial appearance. Striae distensae were present. This proband fulfilled the diagnostic criteria of Ghent nosology.
Discussion
Direct sequencing revealed a total of six variants, either as a single alteration or in different combinations in the 5′ region of the FBN1 gene in six of 41 tested probands fulfilling the diagnostic criteria of Ghent nosology or with features of MFS including at least one major criterion or involvement of two organ systems but not fulfilling a strict interpretation of the Ghent nosology.
Variants exon B, −12A>G and exon B, 112insT occurred as a complex allele in one proband with strongly suspected MFS. Furthermore, he had hypertelorism and exotropia, symptoms also seen in Shprintzen–Goldberg (OMIM#182212) or LDS (OMIM#609192). In order to access possible functional consequences of exon B, −12A>G, a transcription factor binding site search was performed, but no known binding site was found to be associated with this variant in the respective region, whereas, variant exon B, 112insT leads to a frameshift in the conceptual amino-acid sequence and creates a stop codon (UGA) within the exon M (Figure 2). Occurrence of the variant exon B, −12A>G at an evolutionarily conserved position in the proband but not in controls suggests its functional relevance in MFS.
Exon A, +1G>A was found in a proband fulfilling the Ghent diagnostic criteria for MFS. This variant occurred at the donor splice site of exon A, leading to the disruption of the donor splice site (GT>AT). Splice site disruption is expected to alter the reading frame and inactivate the gene product. Conservation of this splice site in different species and the absence of this variant in controls suggests its pathogenic role in MFS.
A complex allele, intron C, +9C>T, intron C, +30C>T and intron C, +48T>C occurred in three unrelated probands with features suggestive of MFS. Intron C, +9C>T was found in one more proband with features of MFS. Intron C, +9C>T and intron C, +30C>T were not found in the controls. In order to access the functional relevance of these variants, a search for transcription factor binding sites was performed. Intron C, +9C>T and intron C, +30C>T were found to create a putative binding site for transcription factor cdxA (CTTTGTG; score 87.2) and Sp1 (CTGGCGGGGT; score 86.3), respectively. Presence of all three variants at the evolutionarily conserved position, their occurrence on the same allele and absence of intron C, +9C>T and intron C, +30 C>T in controls implicate them in the pathogenesis of MFS. However, the apparently unaffected father of the third proband, carrying intron C, +9C>T, intron C, +30C>T and intron C, +48T>C is a carrier of this complex variant. Given that intron C, +9C>T, intron C, +30C>T, have not been found in any of the 306 control chromosomes, and their expected function in transcription, it is possible that they act as predisposing or aggravating disease-related factors in conjunction with other unknown factors. Intron C, +48T>C, was the only variant found in 153 controls, which otherwise showed complete sequence homogeneity in the entire region, a finding in support of selective constraints acting on this gene region.
In summary, we have found five novel variants in the 5′ upstream region of the FBN1 gene in probands fulfilling the Ghent diagnostic criteria or with features of MFS including at least one major criterion or involvement of two organ systems but not fulfilling a strict interpretation of the Ghent nosology, and negative for mutations in the FBN1 exons 1–65, TGFBR2 and TGFBR1 coding regions. We have obtained preliminary evidence for a pathogenic role of some of these variants. Studies aiming to elucidate the role of FBN1 5′ upstream region variants at the transcript level are currently underway.
Accession codes
References
Sakai LY, Keene DR, Engvall E : Fibrillin, a new 350-kD glycoprotein, is a component of extracellular microfibrils. J Cell Biol 1986; 103: 2499–2509.
Pereira L, Andrikopoulos K, Tian J et al: Targetting of the gene encoding fibrillin-1 recapitulates the vascular aspect of Marfan syndrome. Nat Genet 1997; 17: 218–222.
Mecham RP, Heuser JE : The elastic fiber; in Hay ED (ed): Cell Biology of Extracellular Matrix. New York: Plenum, 1991, pp 79–109.
Biery NJ, Eldadah ZA, Moore CS, Stetten G, Spencer F, Dietz HC : Revised genomic organization of FBN1 and significance for regulated gene expression. Genomics 1999; 56: 70–77.
Corson GM, Chalberg SC, Dietz HC, Charbonneau NL, Sakai LY : Fibrillin binds calcium and is coded by cDNAs that reveal a multidomain structure and alternatively spliced exons at the 5′ end. Genomics 1993; 17: 476–484.
Boileau C, Jondeau G, Mizuguchi T, Matsumoto N : Molecular genetics of Marfan syndrome. Curr Opin Cardiol 2005; 20: 194–200.
Tan FK, Wang N, Kuwana M et al: Association of fibrillin 1 single-nucleotide polymorphism haplotypes with systemic sclerosis in Choctaw and Japanese populations. Arthritis Rheum 2001; 44: 893–901.
Mizuguchi T, Collod-Beroud G, Akiyama T et al: Heterozygous TGFBR2 mutations in Marfan syndrome. Nat Genet 2004; 36: 855–860.
De Paepe A, Devereux RB, Dietz HC, Hennekam RC, Pyeritz RE : Revised diagnostic criteria for the Marfan syndrome. Am J Med Genet 1996; 62: 417–426.
Rommel K, Karck M, Haverich A, Schmidtke J, Arslan-Kirchner M : Mutation screening of the fibrillin-1 (FBN1) gene in 76 unrelated patients with Marfan syndrome or Marfanoid features leads to the identification of 11 novel and three previously reported mutations. Hum Mutat 2002; 20: 406–407.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Singh, K., Shukla, P., Rommel, K. et al. Sequence variations in the 5′ upstream regions of the FBN1 gene associated with Marfan syndrome. Eur J Hum Genet 14, 876–879 (2006). https://doi.org/10.1038/sj.ejhg.5201620
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.ejhg.5201620
Keywords
This article is cited by
-
TGFBR3 variation is not a common cause of Marfan-like syndrome and Loeys-Dietz-like syndrome
Journal of Negative Results in BioMedicine (2012)
-
Clinical utility gene card for: Marfan syndrome type 1 and related phenotypes [FBN1]
European Journal of Human Genetics (2010)
-
Conservation of 5′-upstream region of the FBN1 gene in primates
European Journal of Human Genetics (2008)