Abstract
A total of 202 Sardinian male subjects were examined for 13 biallelic stable markers, the complex 49a,f/TaqI system and three microsatellites of the Y chromosome in order to investigate, through surname analysis, on a possible territorial heterogeneity inside the island. The study of geographical distribution and linguistic derivation of Sardinian surnames allow us to discover their ‘probable place of origin’ and reconstruct ancient genetic isolates which borders are, today, no more recognizable. The molecular analysis revealed that about 90% of the Sardinian Y chromosomes fell into haplogroups E-M35, G-M201, I-M26, J-12f2 and R-M269. In contrast with the territorial homogeneity of these haplogroups, when the individuals were distributed according to their birthplace, a significant difference between the three historically and culturally distinct geographical areas into which Sardinia can be subdivided was observed when the individuals were distributed according to the ancestral location of surnames. In particular, the major contribution to this heterogeneity is due to the ‘Sardinian-specific’ haplogroup I-M26 (almost completely associated with the 49a,f-Ht12/12f2-10Kb/YCAIIa-21/YCAIIb-11 compound haplotype), which shows both a significantly higher incidence in the central-eastern (archaic) area and a significantly lower frequency in the northern area. The results of this study agree with the hypothesis that the ancestral homeland of this specific subset of haplogroup I is the mountainous central-eastern area of Sardinia, where the population underwent a long history of isolation since ancient times, and highlight the informative power of the surname analysis.
Similar content being viewed by others
Introduction
In most societies surnames are transmitted from father to child, just like Y-chromosome genes. In this way, an analysis of the geographic diffusion of surnames can provide accurate estimates of migration rates1 and give information to reconstruct the paths followed by men (and their genes) from the beginning of surname history. In addition, if the ‘cultural’ nature of surnames is considered, it is possible to detect much more ancient migrations which, in some cases, point to a common genetic origin.2,3,4 In fact, surnames are linguistic attributions, reflecting local language, which indicated the identity of people in the area where they lived. For this reason, individuals bearing surnames that are derived from words of the same local dialect probably share the same place of origin and the same gene pool.
Sardinia, which shows a particular genetic and cultural pattern due to two contrasting factors: (i) the conquests it has undergone and (ii) its long history of isolation, represents a good test case to verify whether surname history can also provide clues in detecting gene history.
In this study, we examined a sample of 256 Sardinian male subjects for both surnames and Y-chromosome markers (13 biallelic polymorphisms, the complex 49a,f/TaqI system and three microsatellites), in order to evaluate the power of surname analysis in revealing the Y-chromosome ancestry.
Material and methods
The sample
The sample consists of 256 unrelated apparently healthy Sardinian male subjects who gave their informed consent. In total, 116 were conscripts from the Sassari province and were collected in Porto Torres (N=105) and Olbia (N=11); 140 were subjects sampled at the clinical analysis laboratories of the hospitals ‘Ospedale Regionale Microcitemie’ of Cagliari (N=123) and ‘Ospedale A. Segni’ of Ozieri (N=17).
Y-chromosome DNA analysis
The individuals were examined for 13 biallelic markers, namely: the 12f2 [DYS11],5 YAP [DYS287],6 and, in a hierarchical way, M9, M17, M26,7 RPS4Y,8 M35, M74,9 M89, M170, M173, M201,10 M269,11 for the 49a,f/TaqI [DYS1] RFLPs12 and for the variable microsatellites YCAlIa, YCAIIb13 and DYS19.14 The 12f2 and 49a,f/TaqI polymorphisms were analyzed according to Passarino et al15 and the Alu insertion (YAP) according to Hammer and Horai.16 The mutations M17, M26, M35, M170, M173, M201 and RPS4Y were detected by DHPLC as reported by Underhill et al.17 The M9, M74, M89 and M269 were typed through PCR/RFLP assay: M9 and M269 according to Cruciani et al,11 M89 according to Akey et al18 and M74 by using the primers 5′-ATG CTA TAA TAA CTA GGT GGT GAA G-3′ and 5′-AAT TCA GCT TTT ACC ACT TCT GAA-3′, followed by digestion with the restriction enzyme HpyCH4 V. Analyses of the YCAII and DYS19 microsatellites were performed as described by Mathias et al13 and Roewer et al14 who first reported the polymorphisms.
Surname analysis
A geo-linguistic research on the origin of Sardinian surnames was carried out, firstly, by analyzing the distributions of surname data derived from three different sources encompassing the whole territory of Sardinia at different times. The ancient data come from the collection of 48 470 consanguineous marriages celebrated between 1750 and 1950 in the 442 parishes of Sardinia. The more recent data consist of (1) surnames of electric power users in 1983 for all communes of Sardinia (N=484 484); (2) surnames of telephone users in 1993 (N=483 072). Only surnames present in each of the three data sets are considered and their distributions are analyzed in parallel allowing us to trace the dispersion of 4386 surnames throughout the 370 Sardinian communes and to estimate the parameters describing it: the place of maximum frequency and the center of dispersion area.
About 75% of the Sardinian surnames are dispersed on a very small area around the point of highest frequency corresponding to a specific commune. This type of surname is called ‘monophyletic’, assigning to this term the meaning of uniqueness of place of origin.
Moreover, data on surname linguistic origin are obtained searching either for identity (or similarity) between a surname and a toponym of ancient or recent origin in the same neighborhood,19,20,21 or for derivation of a surname from a lexical form,22,23 or for the presence of a surname in old Sardinian documents like the ‘Condaghe’ dating from XII to XIII centuries. For many surnames, these linguistic information may support or rectify the localization of the ‘probable territorial origin’ obtained from the study of their frequency distribution.
For each surname classified as monophyletic, a place of origin is attributed as a numerical code in order to warrant the individuals' privacy.
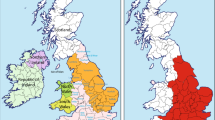
To emphasize the cultural and genetic heterogeneity of Sardinia, the three large areas that reflect its ancient history and geography were considered (Figure 1): the northern zone delimited by the mountain chain crossing Sardinia from the central-west to the north-east and linguistically different from the rest of the island; the south-western zone, delineated by the presence of many Phoenician and Carthaginian archeological sites24 and the central-eastern zone, asylum land of the ancient Sardinian population during invasions and domain of pastoral culture. This zone includes the more conservative, or ‘archaic’ area, defined by archaeological and linguistic25 studies and, more recently, also by studies in geo-linguistics and genetics26 (for a more detailed subdivision of Sardinia on the basis of genes, languages and surnames, see Cavalli-Sforza et al27).
Subdivisions of Sardinia based on geographic and historical criteria. N=northern zone; CE=central-eastern zone including the archaic region; SW=south-western zone. The black line indicates the mountain chain separating the northern zone from the rest of the island. The four centers from which samples were collected are also indicated.
Results
A total of 202 individuals of the initial sample corresponded to 175 monophyletic surnames of which 151 were associated to a single individual and 24 to more individuals. Subjects carrying the same surname, but not the same haplotype, were also included in the analysis, since it may originate from the same linguistic area from which their surnames derived. The remaining 54 individuals corresponded to 41 surnames (19%) for which it was not possible to assign a specific place of origin (polyphyletic surnames).
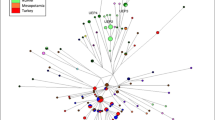
Figure 2 illustrates the world-wide Y-chromosome phylogeny (The Y-chromosome Consortium, 2002),28 where Sardinian Y chromosomes belonging to both monophyletic and polyphyletic surnames are introduced and compared with data from Italy and the Middle East. More than 95% of Sardinian samples fall into haplogroups E-M35, I-M170, J-12f2, G-201 and R-M173. Thus, the haplogroup composition of the Sardinian Y-chromosome pool is very similar to that of Italians and of other Europeans.9,17,29,30,31 The observed differences are therefore mainly quantitative, probably due to isolation and genetic drift. For example, haplogroup I-M170, which has its highest frequency in central-eastern Europe and lower frequencies in western Europe,9,17,32,33 is very frequent in Sardinia. Moreover, the majority of it harbor the additional mutation M26. As for the R-M173 chromosomes, the greater part belongs to the western European R-M269 subcluster, whereas the eastern European subcluster R-M17 is barely represented.
Phylogenetic tree of the Y-chromosome haplotypes and their percent frequencies in the Sardinian samples carrying ‘monophyletic’ and ‘polyphyletic’ (50 out of 54 individuals) surnames. Data on Italian and Middle Eastern samples30 are also given for comparison. Numbering of mutations is according to the YCC:28 those examined in the present study are shown in bold face type; those inferred are shown in italics. Capital letters indicate haplogroups according to the YCC.28 *Ten and ♯one of these chromosomes were not tested for M26 because DNA was finished. One major 49a,f-YCAIIa-YCACIIb-DYS19 compound haplotype characterizes each haplogroup: c-Ht 49a,f-Ht5/YCAIIa-22/YCAIIb-19/DYS19-13 for haplogroup E-M35; c-Ht 49a,f-Ht7/YCAIIa-22/YCAIIb-19/DYS19-14 for J-12f2, c-Ht 49a,f-Ht12/YCAIIa-21/YCAIIb-11/DYS19-17 for I-M26; c-Ht 49a,f-Ht15/YCAIIa-23/YCAIIb-19/DYS19-14 for R-M269 and c-Ht 49a,f-Ht8/YCAIIa-20/YCAIIb-20/DYS19-15 for G-M201.
The analyses of the 49a,f system and three microsatellites (data not shown) reveal the presence of one major compound haplotype per lineage (Figure 2 legend).
To search for an ancient territorial heterogeneity of these haplogroups, we distributed the 202 individuals carrying a monophyletic surname in the three zones described above, according to the surname place of origin. The resulting haplogroup frequencies are shown in Table 1. A general, significant heterogeneity in the distribution of all the haplogroups in the three areas is observed (χ2[10]=34.93, P<0.001). In particular, the cell χ2 analysis shows that the frequency of haplogroup G-M201 is significantly higher than expected in the northern zone (P<0.05), that of haplogroup I-M26 is significantly higher in the central-eastern zone (P<0.01) and lower in the northern zone (P<0.005), and that of haplogroup R-M269 is significantly lower in the central-eastern zone (P<0.05).
In contrast, when places of birth of sampled individuals were used to distribute the different Y-chromosome haplogroups, no heterogeneity among the three areas was detected: χ[10]2=13.36, P=0.204 underlining the effect of migration.
Discussion
Sardinia appears to be a particularly appropriate test case to evaluate the extent to which surnames are informative in identifying the history of Y-chromosome haplogroups.
As the other European populations, almost all the Sardinian Y chromosomes belong to haplogroups E-M35, I-M170, G-M201, J-12f2 and R-M269. Haplogroups E, G and J, which are believed to have an African (E) or Middle Eastern (G and J) origin and entered Europe through different migrations,30,34,35 show frequencies in the same range as other Mediterranean populations. By contrast, haplogroups I-M170 and R-M269 harbor unusual frequencies. Haplogroup R-M269 represents 20.8% of the Sardinian Y chromosomes, which is the lowest frequency in Western Europe (50–80%).30 On the contrary, haplogroup I-M170 shows the highest incidence (41.6%) among western European populations (3–22%),30 and most of it (91.9%) is represented by the subclade I-M26 which in addition is characterized by the compound haplotype 49a,f-Ht12, YCAIIa-21, YCAIIb-11 and DYS19-17 previously proposed as a ‘Sardinian’ marker.36,37 Outside Sardinia, this subclade was only observed at a very low frequency in the Basques,9 the Iberian Peninsula32 and, as inferred by the presence of the YCAIIb-11 (only observed in haplogroup I, and in particular in its subclade M26), in Béarnais, few Corsican and central-southern Italian subjects.38,39,40,41,42
In order to search for genetic heterogeneity inside the island, the effect of migration in the last centuries had to be considered. Indeed, demographic studies on the population evolution of Sardinian communes43 demonstrated that, from 1861 to 1991, the mountain area lost 20% of its population in favor of the plain. The distribution of individuals by birth place compared with that of their ancestors' place of origin seems to reflect this process of homogenization (Figure 3).
So, results shown in Table 1 may enlighten the genetic history of the different parts of Sardinia. The ‘Sardinian’ subhaplogroup I-M26, which is currently distributed almost uniformly in all parts of the island, shows a high heterogeneity between the areas when samples were redistributed according to the ancestral location of surnames. Interestingly, most surnames of individuals carrying this haplogroup seem to have originated in the central-eastern zone, which includes the archaic area. This supports the antiquity of this haplogroup. Indeed, history tells that indigenous populations retreated to the archaic area when Phoenicians and, later, Carthaginians colonized the southern part of the island, and this was followed by centuries of isolation which allowed genetic drift to increase the haplogroup frequency. Moreover, sub-haplogroup I-M26 shows a frequency significantly lower than that expected in the north and a nonsignificant increase in the southwest. Thus, ancient migrations could have brought this haplogroup from the central area towards the more open southern regions, separated only by a failing cultural barrier, more frequently than towards the northern regions, separated by the less accessible geographic barrier. Afterward, recent migrations have dispersed I-M26 all over the island.
The isolation of the central-eastern area could also explain the heterogeneous distribution of the R-M269 and G-M201 haplogroups. The low frequency of haplogroup R-M269 in the central-eastern area of Sardinia and its prevalence in the north suggest that R-M269 arrived to the Sardinian coasts from the continent, possibly after the occurrence and diffusion of the autochthonous I-M26 subhaplogroup, while the high frequency in the northern area of haplogroup G-M201, which is scarcely represented in Europe and in the Middle East,30 could be due to genetic drift.
In conclusion, new methods of sampling are called for and surnames, allowing the detection of the common genetic origin of the families, can bypass the effect of recent migrations and enlighten real genetic differences. Even if this analysis is obviously limited to the male component of the population, the obtained results on the genetic heterogeneity could be extended to the entire Sardinian population due to the smallness of the areas within which matrimonial exchanges occurred.44 Finally, this geo-linguistic approach, in a more general way, could be utilized to select samples of individuals as control for epidemiological studies.
References
Piazza A, Rendine S, Zei G, Moroni A, Cavalli-Sforza LL : Migration rates of human populations from surname distributions. Nature 1987; 329: 714–716.
Skorecki K, Selig S, Blazer S et al: Y chromosomes of Jewish priests. Nature 1997; 385: 32.
Thomas MG, Skorecki K, Ben-Ami H, Parfitt T, Bradman N, Goldstein DB : Origins of Old Testament priests. Nature 1998; 394: 138–140.
Sykes B, Irven C : Surnames and the Y chromosome. Am J Hum Genet 2000; 66: 1417–1419.
Casanova M, Leroy P, Boucekkine C et al: A human Y-linked DNA polymorphism and its potential for estimating genetic and evolutionary distance. Science 1985; 230: 1403–1406.
Hammer MF : A recent insertion of an Alu element on the Y chromosome is a useful marker for human population studies. Mol Biol Evol 1994; 11: 749–761.
Underhill PA, Jin L, Lin AA et al: Detection of numerous Y chromosome biallelic polymorphisms by denaturing high-performance liquid chromatography. Genome Res 1997; 7: 996–1005.
Bergen AW, Wang CY, Tsai J et al: An Asian-native American paternal lineage identified by RPS4Y resequencing and by microsatellite haplotyping. Ann Hum Genet 1999; 63: 63–80.
Underhill PA, Shen P, Lin AA et al: Y chromosome sequence variation and the history of human populations. Nat Genet 2000; 26: 358–361.
Shen P, Wang F, Underhill PA et al: Population genetic implications from sequence variation in four Y chromosome genes. Proc Natl Acad Sci USA 2000; 97: 7354–7359.
Cruciani F, Santolamazza P, Shen P et al: A back migration from Asia to sub-Saharan Africa is supported by high-resolution analysis of human Y-chromosome haplotypes. Am J Hum Genet 2002; 70: 1197–1214.
Ngo KY, Vergnaud G, Johnsson C, Lucotte G, Weissenbach J : A DNA probe detecting multiple haplotypes of the human Y chromosome. Am J Hum Genet 1986; 38: 407–418.
Mathias N, Bayes M, Tyler Smith C : Highly informative compound haplotypes for the human Y chromosome. Hum Mol Genet 1994; 3: 115–123.
Roewer L, Arnemann AJ, Spurr NK, Grzeschik KH, Epplen JT : Simple repeated sequences on the Y chromosome are equally polymorphic as their autosomal counterparts. Hum Genet 1992; 89: 389–394.
Passarino G, Semino O, Quintana-Murci L, Excoffier L, Hammer M, Santachiara-Benerecetti AS : Different genetic components in the Ethiopian population, identified by mtDNA and Y-chromosome polymorphisms. Am J Hum Genet 1998; 62: 420–434.
Hammer MF, Horai S : Y chromosomal DNA variation and the peopling of Japan. Am J Hum Genet 1995; 56: 951–962.
Underhill PA, Passarino G, Lin AA et al: The phylogeography of Y chromosome binary haplotypes and the origins of modern human populations. Ann Hum Genet 2001; 65: 43–62.
Akey JM, Sosnoski D, Parra E et al: Melting curve analysis of SNPs (McSNP): a gel-free and inexpensive approach for SNP genotyping. Biotechniques 2001; 30: 358–367.
Paulis G : I nomi di luogo della Sardegna. Sassari, Carlo Delfino Editore, 1987.
Wolf HJ : Toponomastica Barbaricina. Nuoro, Insula Editore, 1998.
Wolf HJ : Sardische Herkunftsnamen. in Schutzeichel R (ed) Beiträge zur Namenforschung. Heidelberg; Winter, 1988, pp 1–67.
Wagner ML : Dizionario etimologico sardo. Heidelberg, Winter, 1960.
Pittau M : I cognomi della Sardegna. Sassari, Carlo Delfino Editore, 199.
Barreca F : La Sardegna fenicia e punica. Sassari, Chiarella Editore, 1979.
Wagner ML : La lingua sarda. Storia spirito e forma. Berna, Francke, 1950.
Contini M, Cappello N, Griffo R, Rendine S, Piazza A : Géolinguistique et géogénétique, une démarche inter-disciplinaire. Géolinguistique 1989; 4: 129–197.
Cavalli-Sforza LL, Menozzi P, Piazza A : The history and geography of human genes. Princeton NJ, Princeton University Press, 1994.
The Y-chromosome consortium. A nomenclature system for the tree of human Y-chromosomal binary haplogroups. Genome Res 2002; 12: 339–348.
Brega A, Torroni A, Semino O et al: The p12f2/TaqI Y-specific polymorphism in three groups of Italians and in a sample of Senegalese. Gene Geography 1987; 1: 201–206.
Semino O, Passarino G, Oefner PJ et al: The genetic legacy of Paleolithic Homo sapiens sapiens in extant Europeans: A Y chromosome perspective. Science 2000a; 290: 1155–1159.
Passarino G, Semino O, Magri C et al.: The 49a,f haplotype 11 is a new marker of the EU19 lineage that traces migrations from northern regions of the Black Sea. Hum Immunol 2001; 62: 922–932.
Bosch E, Calafell F, Comas D, Oefner PJ, Underhill PA, Bertranpetit J : High-resolution analysis of human Y chromosome variation shows a sharp discontinuity and limited gene flow between Northwestern Africa and Iberian Peninsula. Am J Hum Genet 2001; 68: 1019–1029.
Wells RS, Yuldasheva N, Ruzibakiev R et al.: The Eurasian heartland: a continental perspective on Y-chromosome diversity. Proc Natl Acad Sci USA 2001; 98: 10244–10249.
Semino O, Passarino G, Brega A, Fellous M, Santachiara-Benerecetti AS : A view of the Neolithic demic diffusion in Europe through two Y chromosome-specific markers. Am J Hum Genet 1996; 59: 964–968.
Hammer MF, Karafet T, Rasanayagam A et al: Out of Africa and back again: nested cladistic analysis of human Y chromosome variation. Mol Biol Evol 1998; 15: 427–441.
Santachiara-Benerecetti AS, Semino O : Y-chromosome polymorphisms and history of populations. Cell Pharmacol 1996; 3: 199–204.
Caglià A, Novelletto A, Dobosz M, Malaspina P, Ciminelli BM Pascali VL : Y-chromosome STR loci in Sardinia and continental Italy reveal islander-specific haplotypes. Eur J Hum Genet 1997; 5: 288–292.
Ciminelli, BM, Pompei F, Malaspina P et al.: Recurrent simple tandem repeat mutations during human Y-chromosome radiation in Caucasian subpopulations. J Mol Evol 1995; 41: 966–973.
Malaspina P, Cruciani F, Ciminelli BM et al.: Network analyses of Y-chromosomal types in Europe, northern Africa, and western Asia reveal specific patterns of geographic distribution. Am J Hum Genet 1998; 63: 847–860.
Quintana-Murci L, Semino O, Poloni ES et al: Y-chromosome specific YCAII, DYS19 and YAP polymorphisms in human populations: a comparative study. Ann Hum Genet 1999; 63: 153–166.
Malaspina P, Cruciani F, Santolamazza P et al: Patterns of male-specific inter-population divergence in Europe, West Asia and North Africa. Ann Hum Genet 2000; 64: 395–412.
Scozzari R, Cruciani F, Pangrazio A et al.: Human Y-chromosome variation in the western Mediterranean area: implications for the peopling of the region. Hum Immunol 2001; 62: 871–884.
Angioni D, Loi S, Puggioni G, La popolazione dei comuni Sardi dal 1688 al 1991. Cagliari: C.U.E.C, 1997.
Gatti AM : L'area degli scambi matrimoniali in Sardegna tra XVII e XX secolo. in Oppo A (ed) Famiglia e matrimonio nella società sarda tradizionale. Nuoro, La Tarantola ed., 1990, pp 171–191.
Acknowledgements
We thank A Torroni for his comments on the manuscript, A Moroni for his invaluable work in collecting data of consanguineous marriages, ENEL for having supplied Sardinian surnames of electric power users, SEAT for Italian surnames listed in telephone directories, G Cossu and E Silini for providing us with blood samples. This research was supported by the Progetto Finalizzato C.N.R. ‘Beni Culturali’ the Italian Ministry of the University ‘Progetti Ricerca Interesse Nazionale’, and ‘Fondo d'Ateneo per la Ricerca’ dell'Università di Pavia (to AS S-B).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zei, G., Lisa, A., Fiorani, O. et al. From surnames to the history of Y chromosomes: the Sardinian population as a paradigm. Eur J Hum Genet 11, 802–807 (2003). https://doi.org/10.1038/sj.ejhg.5201040
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.ejhg.5201040
Keywords
This article is cited by
-
Complete mitochondrial sequences from Mesolithic Sardinia
Scientific Reports (2017)
-
Y-chromosomal evidence of the cultural diffusion of agriculture in southeast Europe
European Journal of Human Genetics (2009)
-
Y-STR genetic structure of the most common surnames in Korea
Genes & Genomics (2009)
-
Pathophysiology of ageing, longevity and age related diseases
Immunity & Ageing (2007)
-
Y-chromosomal STR haplotype analysis reveals surname-associated strata in the East-German population
European Journal of Human Genetics (2006)