Abstract
The pathophysiology of preeclampsia is incompletely understood, but the familial nature of the disease has long been recognized. Recent genome-scan studies have indicated linkage at the p23 region of chromosome 2. We have previously reported microsatellite marker association at chromosome region 2p13 in patients with obstetric cholestasis. We conducted population-based association screening with microsatellite markers to find potential preeclampsia-associated loci on chromosome region 2p13–p12 and to test whether preeclampsia and obstetric cholestasis share a single risk locus. The study was carried out among 115 unrelated control women, 133 preeclamptic women and 57 cholestatic women. Screening with microsatellite markers at the 2p13–p12 region revealed that the marker D2S286 was significantly associated with obstetric cholestasis in the overall association analysis (P=0.03), while it revealed only borderline association with preeclampsia (P=0.08). However, single allele association analysis indicated that both preeclampsia and obstetric cholestasis showed a statistically significant association with a common allele (P < 0.05), which was overrepresented in both the obstetric cholestasis (0.42) and preeclamptic (0.37) groups when compared with the control group (0.28). In conclusion, These findings suggest a possible genetic link between chromosome region 2p13–p12, preeclampsia and obstetric cholestasis. More specifically, these data suggest that there may be a common risk locus associated with both obstetric complications located in the vicinity of the 2p13–p12 association region.
Similar content being viewed by others
Introduction
Preeclampsia is a common multifactorial disorder, of unknown etiology, with an established genetic contribution.1,2 A variety of preeclampsia candidate genes such as HLA-DRbeta, HLA-G and the tumor necrosis factor alpha gene (chromosome 6), the angiotensin-converting enzyme gene (chromosome 17) and the CuZn superoxide dismutase gene (chromosome 21) have been examined on the basis of their pathophysiological effects, but so far evidence of association or linkage has been ambiguous, presumably because of differences in populations, small effects of disease alleles on risk, and confounding effects caused by gene–environment interactions.3,4,5,6,7 The genes for factor 5 and methylenetetrahydrofolate reductase lie on chromosome 1. Both have polymorphisms present at a significantly higher frequency in preeclamptic women, and it is likely that they are predisposing factors as regards the development of preeclampsia, without the necessity for it.8,9 The angiotensinogen gene also lies on chromosome 1 and it is in partial linkage disequilibrium with preeclampsia in some populations.6 Similarly, candidate gene studies have provided some controversial evidence of linkage to the eNOS gene on 7q36.10,11 Lachmeijer et al12 recently gave a systematic overview on the numerous candidate gene studies in this field and, overall, these observations appeared to be either population or experiment specific.
Recently, the results of genome-scan studies have indicated susceptibility loci on chromosome regions 4q and 2p in an Icelandic population, the locus at 2p being subsequently found also in a population from Australia and New Zealand.11,13,14 However, on a molecular genetic basis the loci in the Icelandic and Australian scan are very far apart and are likely to reflect different loci. Between the Dutch and Icelandic genomewide scan, two peaks overlapped in the same region at chromosome 3p and chromosome 15q but no overlap was seen on 2p.15 Consequently, replication studies are required across populations to investigate the involvement of the 2p region in the genetic contribution to preeclampsia. Finns are ideal for large-scale linkage disequilibrium mapping studies since they are derived from a small group of founders and are thus more genetically homogeneous than many other populations. Another interesting piece of information is that the 2p13 region, which has shown evidence of significance in preeclampsia, has previously been implicated in obstetric cholestasis in Finland.16 A familial tendency in obstetric cholestasis has been documented, suggesting that genetic factors contribute to its development.17 On the basis of the reported linkage between preeclampsia and this chromosomal region, we have carried out population-based association mapping to document that this linkage is present across populations. Another aim was to clarify whether or not preeclampsia and obstetric cholestasis share a common risk locus.
Materials and methods
Written approval for the study was obtained from the Ethics Committee of Kuopio University Hospital and the protocol was approved by the investigation review board. Informed consent was obtained from all subjects and documented.
Information was collected retrospectively in connection with 133 unrelated preeclamptic women and 57 cholestatic women who delivered at Kuopio University Hospital between January 1994 and December 1999. To ensure homogeneity of the genetic background, controls (N=115) originating from a regional population with no clinical signs of preeclampsia or obstetric cholestasis were enrolled by random selection in this study. The study and control groups were derived only from unrelated women with singleton deliveries at our hospital during the study period. Women with concomitant preeclampsia were excluded from the cholestatic data set; this was done in order to enhance confidence in the diagnosis of cholestasis. For each study and control subject, the outcome of pregnancy was reviewed.
Diagnosis of cholestasis was based on information obtained by clinical examination (generalized pruritus in the absence of any dermatologic condition), laboratory testing revealing a cholestatic pattern (serum aspartate and alanine transferase levels exceeding 40 U/l and serum total bile acid levels exceeding 8 μmol/l) that returned to normal following delivery, with no signs of viral hepatitis (negative results in assays for hepatitis B surface antigen and antihepatitis A and C antibodies), and normal ultrasonographic features of the liver and biliary tract. In this study, both clinical and biochemical disturbances (laboratory testing revealing elevated bile acid levels and transaminases) were required for enrollment. None of our patients underwent liver biopsy. The ancestors of each patient and their birthplaces were traced back three generations to make sure that they were unrelated.17
Hypertensive complications of pregnancy were classified as advocated by the US National High Blood Pressure Education Program Working Group on High Blood Pressure in Pregnancy.18 Preeclampsia was defined as the development of hypertension (>140/90 mmHg) and new-onset proteinuria (>300 mg of urinary protein in 24 h) in women with no proteinuria at baseline. Women with preexisting medical conditions, such as hypertension, diabetes or renal disease, or another complication of pregnancy associated with preeclampsia, such as gestational diabetes, were excluded from the study.
Genomic DNA was isolated from peripheral blood lymphocytes of patients and controls.19 A total of seven chromosome-2-specific fluorescently labeled microsatellite markers located at the 2p13–p12 region (D2S2109 (0 cM) D2S2112 (0.53 cM) D2S2110 (0 cM) D2S1394 (0 cM) D2S1374 (3.23 cM) D2S286 (5.36 cM) D2S1777; relative distances between loci are indicated in parentheses in Kosambi centiMorgans (cM)) were PCR-amplified using a multiplex approach with ATG-polymerase (Applied Bio-systems, Foster City, CA, USA). The sizes of the PCR products were determined using an ABI PRISM 310 genetic analyzer (Applied Biosystems, Foster City, CA, USA). The order of the markers and the estimated distances were obtained from http://marshmed.org/ and http://www.gdb.org/ sex-averaged maps.
Allelic association of the markers was calculated using (two-tailed) Fisher's exact test (Monte Carlo estimate with 99% confidence level), with SPSS software, version 9.0. Testing the significance of an individual marker allele was carried out by pooling the other alleles into one group and calculating the results using Fischer's exact test, and Bonferroni correction was applied by multiplying the P-values by the number of alleles examined. Odds ratios (ORs) as the estimates of relative risk of disease were calculated using 95% confidence intervals (CIs). The Hardy–Weinberg distribution of genotypes for marker D2S286 in the obstetric cholestasis, preeclampsia and control groups was found to be in equilibrium (Associate program, version 2.31). The level of statistical significance was defined as P<0.05.
Results
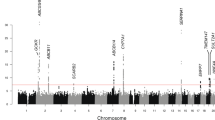
Genotyping of the seven microsatellite markers at the 2p13–p12 region showed that none of them was statistically significantly associated with preeclampsia, although there was a trend towards overall association with marker D2S286, with borderline significance (P=0.08) (Figure 1). When the alleles of the various loci were considered separately, only allele 6 of marker D2S286 was found to be statistically significantly associated with preeclampsia (P=0.04, Bonferroni-corrected P=0.39), giving an odds ratio of 1.48 (95% CI: 1.01–2.16). The frequency of allele 6 was 36.8% in the preeclampsia group and 28.3% in the control group (Table 1).
Association mapping results obtained with chromosome region 2p13–p12-specific microsatellite markers. Markers D2S2109, D2S2112, D2S2110, D2S1394 and D2S1374 have previously been genotyped in women affected by obstetric cholestasis.16
Markers D2S2109, D2S2112, D2S2110, D2S1394 and D2S1374 were genotyped in 47 women affected by obstetric cholestasis, in a previous study.16 In the present study, with an extended number of affected cholestatic women (N=57), marker D2S286 was also found to be associated with obstetric cholestasis (P=0.03). When the alleles of this marker were considered separately (Table 1), the frequency of allele 6 was higher in the cholestatic group than in the control group (42.1 vs 28.3%), indicating that allele 6 of marker D2S286 was significantly associated with obstetric cholestasis (P=0.01, Bonferroni-corrected P=0.09), at an odds ratio of 1.85 (95% CI: 1.15–2.95).
Discussion
In the present study, we tested the association between preeclampsia and obstetric cholestasis and microsatellite markers located at chromosome region 2p13–p12, the diseases being pregnancy-specific disorders with a complex genetic component. Two important aspects were taken into account when initially designing the study. First, the population of Finland is genetically isolated and we can hypothesize that only a small number of ancient changes in susceptibility genes are spread throughout the population.21 Second, we considered it appropriate to search for association between two pregnancy-specific disorders and the chromosomal region at 2p13. The rationale for this was based on the following studies: markers D2S2111 and D2S1394 at chromosome region 2p13 have been implicated in susceptibility to preeclampsia in a genome screen in Iceland.13 Independently of this, in a previous study we found that two microsatellite markers, D2S1394 and D2S1374, located at chromosome region 2p13, were associated with cholestatic phenotypes in the restricted population of Finland.16
The results of this study support the presence of a preeclampsia susceptibility locus at 2p13–p12. Plausible single-allele association was found for microsatellite marker D2S286, which was previously reported to have significant linkage with preeclampsia in families from Iceland.13 Furthermore, overall and single-allele association was found for D2S286 in the obstetric cholestasis group, which was an expected finding on the basis of our previous results,16 as this marker is located approximately 3.23 cM from the previously associated markers D2S1394 and D2S1374. Overall, D2S286 is the third associated marker at the 2p13–p12 region among our cholestatic data set, therefore strengthening the suggestion that this chromosomal region may encompass a novel susceptibility locus for obstetric cholestasis. Interestingly, more precise assessment at the allele distribution level indicates that both preeclampsia and obstetric cholestasis are associated with the same marker allele of D2S286, suggesting that there may be a common susceptibility locus for both preeclampsia and cholestasis.
The Finns represent a population that has developed from a few thousand ancestors and remained isolated for several centuries, and therefore differences in genetic material may explain a larger proportion of the phenotype than in a more heterogeneous population.22,23 Association studies in different ethnic groups may increase an understanding of the relationship between genetic susceptibility and preeclampsia. Regardless of variability of the genetic background in Iceland, Australia and Finland, we found suggestive association with chromosome region 2p13–p12 in preeclampsia. Thus, the association with preeclampsia proved to be replicative across populations, and therefore it is unlikely that these findings are false positives. Although current evidence points to a role for region 2p13–p12, the exact identity of the preeclampsia susceptibility genes has not been established. However, the detection of a locus with only a modest effect would most probably require a much larger sample size than was available in our study.
Although the precision of genome scans is somewhat poor, a single risk locus, occurring at the same position on the chromosome in groups affected by preeclampsia and obstetric cholestasis, suggests a genetic link or overlap between the two diseases. Accordingly, the present finding, with a single overrepresented allele, may represent evidence of the same locus segregating in women affected by preeclampsia and obstetric cholestasis. Although any theory remains speculative at this point, an unknown locus nearby may influence responsiveness to pregnancy-related metabolic or hormonal changes, resulting in the development of the complications. Thus, preeclampsia and obstetric cholestasis may share a common pathogenesis, which implies that a common group of genes may underlie these two diseases. Interestingly, Tyni et al24 have previously reported that asymptomatic carriers of long-chain 3-hydroxyacyl-coenzyme A dehydrogenase deficiency are at increased risk of developing both preeclampsia and obstetric cholestasis. The gene is located at the p23 region of chromosome 2. Although the manner in which a risk locus interacts to increase the risk of preeclampsia and cholestasis is unknown, this study may provide new insights into the genetic architecture of the two obstetric disorders. It is questionable whether the use of more markers will lead to the detection of candidate genes related to obstetric complications, and therefore a candidate gene approach or functional studies may be more appropriate in the future.
References
Arngrimsson R, Björnsson H, Geirsson R : Analysis of different inheritance patterns in preeclampsia/eclampsia syndrome. Hypertens Preg 1995; 14: 27–38.
Cincotta RB, Brennecke SP : Family history of pre-eclampsia as a predictor for pre-eclampsia in primigravidas. Int J Gynaecol Obstet 1998; 60: 23–27.
Wilton AN, Cooper DW, Brennecke SP, Bishop SM, Marshall P : Absence of close linkage between maternal genes for susceptibility to pre-eclampsia/eclampsia and HLA DR beta. Lancet 1990; 336: 653–657.
Humphrey KE, Harrison GA, Cooper DW, Wilton AN, Brennecke SP, Trudinger BJ : HLA-G deletion polymorphism and pre-eclampsia/eclampsia. Br J Obstet Gynaecol 1995; 102: 707–710.
Chen G, Wilson R, Wang SH, Zheng HZ, Walker JJ, McKillop JH : Tumour necrosis factor-alpha (TNF-alpha) gene polymorphism and expression in preeclampsia. Clin Exp Immunol 1996; 104: 154–159.
Ward K, Hata A, Jeunemaitre X et al: A molecular variant of angiotensinogen associated with preeclampsia. Nat Genet 1993; 4: 59–61.
Chen G, Wilson R, Boyd P et al: Normal superoxide dismutase (SOD) gene in pregnancy-induced hypertension: is the decreased SOD activity a secondary phenomenon? Free Radic Res 1994; 21: 59–66.
Dizon–Townson DS, Nelson LM, Easton K, Ward K : The factor V Leiden mutation may predispose women to severe preeclampsia. Am J Obstet Gynecol 1996; 175: 902–905.
Sohda S, Arinami T, Hamada H, Yamada N, Hamaguchi H, Kubo T : Methylenetetrahydrofolate reductase polymorphism and pre-eclampsia. J Med Genet 1997; 34: 525–526.
Lewis I, Lachmeijer G, Downing S et al: Failure to detect linkage of preeclampsia to the region of the NOS3 locus on chromosome 7q. Am J Hum Genet 1999; 64: 310–313.
Arngrimsson R, Hayward C, Nadaud S et al: Evidence for a familial pregnancy-induced hypertension locus in the eNOS-gene region. Am J Hum Genet 1997; 61: 354–362.
Lachmeijer AM, Dekker GA, Pals G, Aarnoudse JG, ten Kate LP, Arngrimsson R : Searching for preeclampsia genes: the current position. Eur J Obstet Gynecol Reprod Biol 2002; 105: 94–113.
Arngrimsson R, Sigurardottir S, Frigge ML et al: A genome-wide scan reveals a maternal susceptibility locus for pre-eclampsia on chromosome 2p13. Hum Mol Genet 1999; 8: 1799–1805.
Moses EK, Lade JA, Guo G et al: A genome scan in families from Australia and New Zealand confirms the presence of a maternal susceptibility locus for pre-eclampsia, on chromosome 2. Am J Hum Genet 2000; 67: 1581–1585.
Lachmeijer AM, Arngrimsson R, Bastiaans EJ et al: A genome-wide scan for preeclampsia in the Netherlands. Eur J Hum Genet 2001; 9: 758–764.
Heinonen S, Eloranta M-L, Heiskanen J et al: Maternal susceptibility locus for obstetric cholestasis maps to chromosome region 2p13. Scand J Gastroenterol 2001; 36: 766–770.
Eloranta M-L, Heinonen S, Mononen T, Saarikoski S : Risk of obstetric cholestasis in sisters of index patients. Clin Gen 2001; 60: 42–45.
Report of the National High Blood Pressure Education Program Working Group on High Blood Pressure in Pregnancy. Am J Obstet Gynecol 2000; 183: S1–S22.
Vandenplas S, Wiid I, Grobler-Rabie A et al: Blot hybridization of genomic DNA. J Med Genet 1984; 21: 164–172.
Sheffield VC, Weber JL, Buetow KH et al: A collection of tri- and tetranucleotide repeat markers used to generate high quality high, resolution human genome-wide linkage maps. Hum Mol Genet 1995; 4: 1837–1844.
Peltonen L, Jalanko A, Varilo T : Molecular genetics of the Finnish disease heritage. Hum Mol Genet 1999; 8: 1913–1923.
Norio R, Nevanlinna HR, Perheentupa J : Hereditary diseases in Finland; rare flora in rare soil. Ann Clin Res 1973; 5: 109–141.
Nevanlinna HR : The Finnish population structure. A genetic and genealogical study. Hereditas 1972; 71: 195–236.
Tyni T, Ekholm E, Pihko H : Pregnancy complications are frequent in long-chain 3-hydroxyacyl-coenzyme A dehydrogenase deficiency. Am J Obstet Gynecol 1998; 178: 603–608.
Author information
Authors and Affiliations
Corresponding author
Additional information
The study was supported by an EVO grant from Kuopio University Hospital.
Rights and permissions
About this article
Cite this article
Laasanen, J., Hiltunen, M., Romppanen, EL. et al. Microsatellite marker association at chromosome region 2p13 in Finnish patients with preeclampsia and obstetric cholestasis suggests a common risk locus. Eur J Hum Genet 11, 232–236 (2003). https://doi.org/10.1038/sj.ejhg.5200951
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.ejhg.5200951
Keywords
This article is cited by
-
Intrahepatic cholestasis of pregnancy as a risk factor for preeclampsia
Archives of Gynecology and Obstetrics (2020)
-
The Genetic Aspects of Pre-eclampsia: Achievements and Limitations
Biochemical Genetics (2008)
-
Genetics of preeclampsia: paradigm shifts
Human Genetics (2007)
-
Sequence variation in the ATP8B1 gene and intrahepatic cholestasis of pregnancy
European Journal of Human Genetics (2005)