Abstract
The ubiquitous vacuolar H+-ATPase, a multisubunit proton pump, is essential for intraorganellar acidification. Disruption of its function leads to disturbances of organelle function and cell death. Here, we report that overexpression of the B2 subunit of the H+-ATPase inhibits apoptosis. This antiapoptotic effect is not mediated by an increase in H+-ATPase activity but through activation of the Ras-mitogen-activated protein kinase (MAPK)-signaling pathway that results in the serine phosphorylation of Bad at residues 112 and 155. Increased Bad phosphorylation reduces its translocation to mitochondria, limits the release of mitochondrial cytochrome c and apoptosis-inducing factor and increases the resistance of the B2 overexpressing cells to apoptosis. Screening experiments of kinase inhibitors, including inhibitors of cAMP-activated protein kinase, protein kinase C, protein kinase B, (MAPK/extracellular signal-regulated (ERK) kinase) MEK and Ste-MEK113, a cell permeable ERK activation inhibitor peptide, revealed that the B2 subunit of H+-ATPase acts upstream of MEK activation in the MEK/ERK pathway to ameliorate apoptosis.
Similar content being viewed by others
Introduction
Vacuolar H+-ATPases are ubiquitous multisubunit complexes that mediate the acidification of many intracellular compartments including the lysosomes, Golgi apparatus, secretory vesicles and endosomes in virtually all of eukaryotic cells. Disruption of the acidic intraorganellar pH leads to disturbance of organelle function and often promotes cell death.1 Prior studies have demonstrated that specific inhibition of H+-ATPase (e.g. by bafilomycin or concanamycin) induces apoptosis.2, 3, 4, 5, 6 In contrast, increased expression of H+-ATPase was correlated with delayed apoptosis and cell differentiation.7, 8, 9 These observations suggest additional functions for H+-ATPase.
In this study, a stable human embryonic kidney (HEK) 293 cell line that overexpresses the B2 subunit of H+-ATPase (ATP6V1B2) as a green fluorescent protein (GFP) fusion protein was established and used to evaluate the potential role of this subunit in antagonizing apoptosis. Without any influences on expression or enzyme activity of the endogenous H+-ATPase, selective overexpression of GFP-B2 increased resistance of the cells to apoptosis triggered by serum starvation or staurosporine. This effect was mediated by blocking mitochondrial release of sequestered apoptogenic proteins including cytochrome c and apoptosis-inducing factor (AIF). The underlying mechanism by which B2 overexpression blocks cytochrome c and AIF translocation from the mitochondrial intermembranespace to the cytosol involves phosphorylation of Bad, a proapoptotic member of the Bcl-2 family, at position Ser-112 and -155 by ribosomal S6 kinase (RSK), the RSK, that was activated in cells that overexpress the B2 subunit. Inhibition of the MEK1-ERK-activating pathway reduced the protective effects of B2 both on mitochondrial membrane injury and apoptosis. Thus, the MEK/ERK/RSK/Bad cascade is one key pathway affected by B2 overexpression that modifies the cellular response to proapoptotic stimuli.
Results
Generation of stably transfected cell line that expresses GFP-B2 protein
To investigate the role of the B2 subunit of the H+-ATPase in apoptosis, a HEK 293 stably transfected cell line was established that expresses a fusion protein, GFP-B2, in which the carboxyl terminus of GFP protein is fused with the amino-terminus of H+-ATPase B2 subunit and is expressed under the control of human cytomegalovirus immediate–early promoter/enhancer. As a control, a GFP overexpressing HEK 293 cell line (293GFP) was also established. The GFP-B2 fusion protein in the clone (293B2) used in this study is an 81 kDa protein as documented by Western blot analysis (Figure 1a). This protein has a perinuclear distribution as shown by confocal microscopy (Supplementary Figure 2A B2).
Expression of GFP-B2 fusion protein in stably transfected HEK 293 (293B2) cells and its effect on endogenous H+-ATPase expression and activity. (a) Immunoblot analysis of the expression of GFP-B2 in cell homogenates from nontransfected (293) and GFP-B2 stably transfected (293B2) HEK 293 cells. The membrane was probed with an anti-GFP antibody. The expected 81 kDa GFP-B2 fusion protein is only detected in 293B2 cells. (b) Immunoblot analysis of H+-ATPase expression of subunit E (31 kDa) and B (57 kDa) in cell homogenates from GFP (293GFP) and GFP-B2 (293B2) stably transfected HEK 293 cells. Immunoblotting was performed as described utilizing anti-H+-ATPase E and B subunit and anti-actin antibodies. A representative experiment from four independent experiments is shown. (c) Quantification of band density of immunoblots (n=4) of the 31 kDa E and 57 kDa B subunits of H+-ATPase and actin in 293 GFP and 293B2 cells. Overexpression of B2 does not change the expression of the endogenous H+-ATPase since the 31 kDa E and the 57 kDa B subunits are equally expressed in the two cells types. (d) Effect of GFP-B2 overexpression on endogenous H+-ATPase activity. Cell homogenates from GFP (293GFP) and GFP-B2 (293B2) stable cell lines were analyzed for ATPase activity in the presence (−Ba) or absence of bafilomycin (1 μM) as described. Bafilomycin-sensitive H+-ATPase activities, as determined by subtraction of the ATPase activity in the presence of 1 μM bafilomycin from that in the absence of bafilomycin, were not different from each other in 293GFP and 293B2 cells. Inner medullary collecting duct cells (IMCD) were used in the assay as a positive control. These results are representative of six independent experiments
Overexpression of GFP-B2 does not effect intrinsic H+-ATPase expression
In order to determine whether transfection and expression of GFP-B2 interferes with endogenous H+-ATPase expression and ATPase activity, the cellular content of the 31 kDa E and 57 kDa B subunits of endogenous H+-ATPase as well as ATPase activity were determined in both 293GFP and 293B2 cells. Both cell lines had similar amounts of E and B subunits as determined by immunoblot (Figure 1b and c) and exhibited the same levels of bafilomycin-sensitive ATPase activity (Figure 1d). These results indicate that overexpression of a GFP-B2 fusion protein does not alter the pattern of the endogenous H+-ATPase gene expression or effect H+-ATPase activity.
Overexpression of GFP-B2 protects cells from apoptosis
To study the potential role of the H+-ATPase B2 subunit in apoptosis, 293B2 and 293GFP cells were subjected to one of two distinct, proapoptotic stimuli; serum starvation or staurosporine. In contrast to control (293GFP cells), apoptosis was markedly inhibited in 293B2 overexpressing cells after serum withdrawal or exposure to staurosporine (Figure 2a). Quantification of apoptosis by flow cytometry analysis of DNA degradation showed that serum starvation increase apoptosis from 0.6 to 14.3 and 69.8% after 0, 48 and 72 h of serum withdrawal in control 293GFP cells, whereas, in 293B2, apoptosis was detected in 0.7, 0.7 and 4.6% of cells at the same time points (Figure 2b). The rate of apoptosis was approximately 21 and 15 times lower in 293B2 cells than in the control 293GFP cells after serum deprivation for 48 and 72 h. Transient overexpression of GFP-B2 fusion protein in HEK 293 cells also increased the rate of cell survival during serum starvation (data not shown). Also, preliminary data showed that overexpression of the B1 subunit of H+-ATPase did not lower the rate of apoptosis in the transfected HEK293 cells after serum deprivation (Supplementary Figure 1). These results demonstrate that B2 subunit overexpression improves survival in cells exposed to proapoptotic stimuli.
Effect of B2 overexpression on serum withdrawal and staurosporine-induced apoptosis. (a) GFP- (293GFP, upper panel) or GFP-B2 (293B2, lower panel)-transfected cells either cultured in normal condition (10% FCS, left) or deprived of serum for 72 h (middle) or treated with 2 μM staurosporin for 4 h (right) were fixed and stained with Hochest dye #33342. Apoptotic cells are marked with white arrows. (b) Bar graph depicting the percentage of apoptosis determined by analyzing the DNA content in 293GFP and 293B2 cells before (0 h) and after serum starvation (48 and 72 h) (***P<0.01, compared to 0 h; ΔΔΔP<0.01, 293GFP cells compared to 293B2 cells, n=5). DNA content was determined by propidium iodide staining of permeabilized cells and analyzed by flow cytometry. Cells with sub-G1 DNA content (sub-G1 cells) representing fragmented DNA were considered to be apoptotic
Overexpression of B2 reduces mitochondrial release of cytochrome c and AIF during apoptosis
Apoptosis may be initiated through either a receptor-mediated or endogenous pathway that involves mitochondrial membrane injury.10 When triggered by the mitochondrial pathway, apoptogenic factors including cytochrome c and AIF are translocated from mitochondria into the cytosol. Cytochrome c triggers caspase-3 activation through formation of the cytochrome c/Apaf-1/caspase-9-containing apoptosome complex,11 whereas AIF translocates into nuclei, resulting in large-scale DNA degradation.12 These events produce many of the morphologic and biochemical features of apoptosis. In order to elucidate the mechanism of B2-mediated mitochondrial membrane protection, the release of cytochrome c and AIF was assessed by measuring the accumulation of these proteins in the cytosol during serum deprivation. Control 293GFP cells displayed a time-dependent increases in both cytochrome c and AIF release, whereas these same proapoptotic proteins did not translocate from the mitochondria of 293B2 cells despite serum deprivation (Figure 3). These results suggest that B2 subunit overexpression prevents apoptosis by limiting mitochondrial membrane injury.
Effect of B2 overexpression on AIF and cytochrome c release during serum deprivation. GFP- (control) or GFP-B2 (B2)-transfected HEK 293 cells were subjected to serum withdrawal (ss) for 0–72 h. Cytosolic fractions were analyzed by immunoblot analysis for AIF and cytochrome c as described in Materials and Methods. Results are representative of four independent experiments
B2 overexpression regulates the activity of Bad
B2 overexpression could prevent the mitochondrial release of apoptogenic proteins by one or more of several mechanisms including (1) direct interaction with and protection of the mitochondrial membrane; (2) modifying the activity of Bcl-2 family members by forming complexes with these family members or (3) indirectly modifying the activity of Bcl-2 family members by altering signal events that result in their activation.
First, we examined whether or not GFP-B2 fusion protein interacts directly with the mitochondrial membrane by determining the distribution of B2 by morphological (immunochemical) methods. GFP-B2 is localized to a compartment distinct from the mitochondrial compartment (Supplementary Figure 2A). The lack of localization of B2 to the mitochondria indicates that B2 does not protect mitochondria by direct mitochondrial interaction.
To determine whether the B2 subunit forms a complex with Bcl-2 family members including bcl-2, bax and Bad, co-immunoprecipitation studies were performed. GFP-B2 was immunoprecipitated and the precipitates were reprobed for the above Bcl-2 family members. Although bcl-2 family members could be detected in whole cell lysates from 293B2 and 293GFP, they could not be identified in GFP-B2 immunoprecipitates (Supplementary Figure 2B).
A third mechanism by which B2 overexpression may prevent apoptosis is by modifying the activation of bcl-2 family members. In order to alter membrane permeability, Bcl2 members must integrate with the mitochondrial membrane itself.10 To evaluate this mechanism, the effect of B2 overexpression on the fraction of membrane-bound pro- and antiapoptotic proteins was examined. For these studies, the digitonin-insoluble cell fraction was considered to be membrane bound. Insoluble bcl-2, bax and Bad fractions were assayed by immunoblot analysis before and after serum deprivation in both 293GFP and 293B2 cells (Figure 4a and c). Although membrane-bound bcl-2 and bax were similar in both cell lines and did not change during serum deprivation, the fraction of membrane-bound Bad was much lower in 293B2 cells than in 293GFP cells prior to serum starvation. Time-dependent increases in membrane-bound Bad were observed in 293GFP but not in 293B2 cells during serum starvation. These results indicate that B2 inhibits Bad activation and translocation from a soluble to membrane-associated fraction, a process that triggers apoptosis. To further characterize these observations, immunoblot analysis of purified mitochondria was performed before and after serum deprivation in 293B2 and 293GFP cells (Figure 4b and c). The Bad content in the mitochondrial fraction from 293GFP cells was greater than that of 293B2 cells prior to serum deprivation. Furthermore, an increase in mitochondrial Bad content occurred only in serum-deprived 293GFP cells. Taken together, these results indicate that B2 overexpression interferes with activation of Bad and its translocation to mitochondria membrane during a proapoptotic insult.
Effect of B2 overexpression on Bad inactivation. (a) Membrane-associated Bad content: equal amounts of digitonin insoluble protein obtained from 293GFP- (lanes 1, 3, 5 and 7) and 293B2- (lanes 2, 4, 6 and 8) transfected cells after serum withdrawal (0, 24, 48 and 72 h) were analyzed for bcl 2, bax and Bad content by immunoblot. Results are representatives of three separate experiments. (b) Bad content in cytosol and mitochondrial preparations before and after serum withdrawal in 293GFP- (lanes 1–4) and 293B2 (lanes 5–8)-transfected cells. Cytosolic (upper panel) and mitochondrial preparations (middle panel) were obtained from cells cultured in medium with (0 h, lane 1 and 5) or without serum for 24 h (lanes 2 and 6); 48 h (lanes 3 and 7); and 72 h (lanes 4 and 8). The lower panel is the same blot as the middle panel but was reprobed for cytochrome c oxidase (Cox, complex IV), a mitochondrial marker protein after it was stripped. Results are representative of three independent experiments. (c) Densitometric analysis of Bad distribution in the membranous and mitochondrial fraction in 293GFP and 293B2 cells before (0 h) and after (24, 48 and 72 h) serum deprivation. ***P<0.01 when compared to 293B2 cells; ΔP<0.05 when compared to the 293GFP cells without serum deprivation (0 h) (n=3). (d) Bad phosphorylation: equal amounts of cytosolic proteins harvested from 293GFP (lanes 1, 3, 5 and 7) and 293B2 (lanes 2, 4, 6 and 8) cells during serum withdrawal (0, 24, 48 and 72 h) were analyzed by immunoblot for Bad using phosphorylation-specific antibodies directed against serine residues ser-112, -155 and -136 as well as against total Bad. Results are representative of four independent experiments. (e) Densitometric analysis of Bad phosphorylation Ser-112 during serum withdrawal. ***P<0.01 when compared to 293GFP cells; Δ, ΔΔΔP<0.05 and 0.01 when compared to the 293B2 cells without serum withdrawal (0 h) (n=4)
B2 overexpression increases phosphorylation of Bad at Ser-112 and -155
Bad activity, as well as its relative distribution between cytosolic and membrane compartments, is regulated by serine phosphorylation.13, 14 Therefore, the phosphorylation levels of Bad at three functional sites, Ser-112, -136 and -155 was investigated. The amount of total Bad expressed by both cell lines was similar. However, the degree of phosphorylation of Ser-112 and -155 in 293B2 cells was greater than in 293GFP cells under serum-deplete conditions. In contrast, both cell lines showed similar levels of phosphorylation at Ser-136 (Figure 4d). Furthermore, a time-dependent increase in ser-112 phosphorylated Bad was detected during serum starvation in 293B2 cells (Figure 4e). These results indicate that B2 overexpression selectively increases the phosphorylation of Bad at Ser-112 and -155, and blocks its translocation from cytosol to mitochondria.
p90RSK pathway is activated by B2 overexpression
To identify potential kinases that might be involved in the Bad phosphorylation, active (phosphorylated), as well as total p42/44 MAPK, the downstream kinase p90RSK and AKT were measured and quantified in 293GFP and 293B2 cell lysates before and during serum starvation (Figure 5a and b). After 72 h of serum deprivation, the activity of AKT (phospho-AKT) decreased in both 293GFP and 293B2 cells, whereas p42/44 MAPK and p90RSK activity (phospho-p42/44 MAPK and p90RSK) increased in 293B2 but not in 293GFP cells. This suggests that MAPK and its downstream RSK activation mediates the effects of B2 overexpression on Bad phosphorylation and inhibits apoptosis in 293B2 cells. Unexpectedly, a significantly lower amount of phospho-AKT was detected in 293B2 cells compared to 293GFP cells before and during serum starvation. This decreased activity of AKT in B2 overexpressed cells may also elicit signals that result in the activation of p90RSK because of the negative influence of AKT on Raf1 and therefore subsequently on MEK/ERK/RSK activation.15
Activation of MEK/ERK signal pathway by B2 overexpression during serum starvation-induced apoptosis. Changes in active, phosphorylated forms of p42/44 MAPK (phospho/P-p42/44 MAPK), p90RSK (phospho/P-p90RSK) and AKT (p-AKT) as well as their total forms (p42/44 MAPK, p90RSK and AKT) were evaluated by Western blot (a) and the phosphokinases were quantified (b) in GFP and GFP-B2-transfected cells before (GFP and B2) and after 72 h of serum starvation (GFPss and B2ss). *, ***P<0.05 and 0.01 when compared to GFP cells; Δ, ΔΔΔP<0.05 and 0.01 when compared to nonstarved corresponding cells (n=4)
Involvement of ERK activation through MEK1 in B2 overexpressing cells
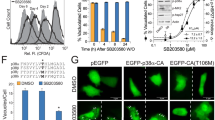
Bad is phosphorylated by AKT at ser-13616, 17 and by RSK at both ser-112 and -155.18, 19 RSK can be activated by ERK1/2 (MAPK), protein kinase C (PKC) or protein kinase A (PKA).20 To further identify the prosurvival pathway activated by B2, the effect of protein kinase inhibitors on apoptosis was examined. Of the inhibitors tested (bisindolylmaleimide I – PKC; U0126 – MEK1/MEK2; AKT inhibitor; H8 – PKA; and Ste-MEK1–13) at a two-fold higher concentration than the established IC50, only Ste-MEK1–13 antagonized the antiapoptotic effect of B2 overexpression. Ste-MEK1–13 is a cell-permeable, stearated 13-amino-acid peptide, corresponding to the N-terminus of MEK1 (MAPKK) that acts as a specific inhibitor of ERK activation, selectively binds to ERK2 and prevents its interaction with MEK.21 After serum deprivation for 24 h, control 293GFP cells displayed significant morphologic evidence of apoptosis as shown by annexin V staining (Figure 6a). In contrast, serum-deprived 293B2 cells only became apoptotic after treatment with Ste-MEK113 in a range of concentration from 7 to 56 μM. Ste-MEK1-13 was reported to have an IC50 of 13±3–30±5 μM for inhibiting ERK both in vitro and in vivo.21 This specific ERK inhibitor also increased both cytochrome c and AIF release (Figure 6b) and reduced the degree of Bad phosphorylation (Figure 6c) during serum starvation despite B2 overexpression. As shown, Ste-MEK1-13-induced apoptotic morphological changes in 293B2 cells after serum deprivation for 24 h at a concentration as low as 7 μM and showed a dose-dependent response in the range of 7–56 μM. In the presence of serum (10% fetal calf serum (FCS)) 56 μM Ste-MEK1–13 had no proapoptotic effects on either 293GFP or 293B2 cell lines. Neither of them displayed any apoptotic morphologic changes (Supplementary Figure 3A), indicating Ste-MEK1-13 induces apoptosis in serum-deprived 293B2 cells by antagonizing the antiapoptotic effect of B2 thereof. In this series of experiments, 200 nM U0126, another MEK1/MEK2 inhibitor induced only minor apoptotic changes in 293B2 cells as shown by annexin V staining and no significant increase in cytochrome c or AIF release (Figure 6b). In subsequent experiments, we increased the concentration of U0126 from 200 nM to 10 μM and also evaluated the effect a third MEK inhibitor, PD98059, at concentration of 25 μM.22 Both MEK inhibitors induced significant apoptosis in serum-deprived 293B2 cells, whereas an increased concentration of the PKC inhibitor, bisindolylmaleimide I at 1 μM did not (Supplementary Figure 3B). These results are consistent with our proposal that B2 overexpression activates the MEK1/ERK/RSK pathway at a point upstream MEK. The activation of this pathway results in the phosphorylation of Bad and therefore the antiapoptosis effect of B2.
Role of ERK activation in B2-mediated cytoprotection. 293GFP and 293B2 stably transfected cells were cultured in standard DMEM with 10%FCS (1 and 3) or without serum for 24 h (2 and 4). The remaining lanes show 293-B2 cells cultured in medium without serum that were treated with one of several protein kinase inhibitors for 24 h as follows: (5) Bisindolylmaleimide I, 20 nM; (6) U0126, 200 nM; (7) AKT inhibitor, 10 mM; (8) H8, 2.4 mM; (9) Ste-MEK1-13, 7 μM; (10) Ste-MEK 1–13, 14 μM; (11) Ste-MEK1–13, 28 μM; (12) Ste-MEK1-13, 56 μM. (a) Cells were stained with cy5-conjugated Annexin V (upper panels) before fixation and then stained with Hochest #33342 (lower panels) to assess the extent of apoptosis. (b) Cytosolic fractions of cells treated as described above were analyzed for cytochrome c, AIF and actin by immunoblot analysis. (c) Degree of Bad phosphorylation was determined by immunoblot with specific antiphospho-serine antibodies for 112, 136 and 155 in whole cell lysates from 293B2 and 293GFP cells treated as described above. Results are representative of three independent experiments
Discussion
We reported here, for the first time, that overexpression of B2, a subunit of the ubiquitous vacuolar H+-ATPase, enhances the resistance of HEK 293 cells to apoptosis. Overexpression of B2 activates the MEK/ERK/RSK/Bad signal pathway, enhances phosphorylation of RSK and thereby increases the serine phosphorylation of Bad at residues 112 and 15523 (Figure 7). When Bad is serine phosphorylated on these functional residues, its proapoptotic function is suppressed by increasing its association with 14-3-3ζ and by decreasing its association with bcl-xL or bcl-2 at the mitochondrial membrane.14, 24, 25, 26 Thus, B2 overexpression increases the threshold for mitochondrial release of sequestered apoptogenic proteins (cytochrome c and AIF) and markedly reduces apoptosis during serum starvation or staurosporine treatment.
Effect of B2 overexpression on the apoptotic-signaling pathway. Activation of Bad by apoptotic signals participates in the production of mitochondrial injury and the release of proapoptotic molecules such as cytochrome c and AIF. Overexpression of the B2 subunit of the H+-ATPase B2 activates the MEK1/ERK/RSK pathway at MEK or a point upstream of MEK. The activation of this pathway results in inactivation of Bad by phosphorylation at Ser-112 and -155 and amelioration, reduction in mitochondrial injury and thereby a reduction in apoptosis
In our study, the overexpression of the B2 subunit, as a GFP-B2 fusion protein, in stably transfected HEK cells is antiapoptotic. The introduction of this subunit neither altered intrinsic H+-ATPase gene expression, as demonstrated by the similarity in the expression level of endogenous B and E subunits of H+-ATPase in control (293GFP) and transfected cells (293B2) (Figure 1b and c) nor increased formation of assembled H+-ATPase enzyme since the bafilomycin-sensitive ATPase activity was also similar (Figure 1d). Thus, the antiapoptotic effect of B2 can be attributed to B2 overexpression per se and not to an alteration in either the expression level of other intrinsic H+-ATPase subunits or an increase in overall H+-ATPase activity.
The antiapoptotic effect exerted by a single subunit of H+-ATPase is a novel observation for this ubiquitously expressed B2 subunit. Free unassembled subunits of the H+-ATPase exist in a dynamic balance between the assembled and unassembled forms in cells.1 If subunits of this enzyme, in addition to forming the enzyme complex for proton transport, play other functional roles, then variations in the ratio between assembled and unassembled subunits during the cell cycle may determine, in part, whether a cell undergoes differentiation, proliferation or cell death. During these processes, cells with an elevated level of free B2 may be more resistant to signals that activate cell death. The lower baseline of cytosolic AIF and cytochrome c (Figure 3) and mitochondrial distribution of Bad (Figure 4) in the B2-overexpressing 293B2 cells suggests that increased level of cellular B2 may not only antagonize maneuvers that triggered apoptosis, but also silence the intrinsic programmed cell death pathway and thereby upgrade the threshold for apoptosis. The ability to regulate programmed cell death is critical for normal histogenesis and organ development. For example, the expression of the B2 subunit, as well as other subunits of the ubiquitous H+-ATPase, shows very early induction (embryonic day 13.5[E13.5] during nephrogenesis in contrast to the later (>E15.5) expression of the intercalated cell-specific B1 subunit.27 The B2 subunit can be upregulated independent of the other subunits by increased transcription of a novel factor through an AP-2 site during monocytic differentiation.8 Also, granulocyte colony-stimulating factor, a growth factor, upregulated expression of the 57-kDa B2 subunit and delay spontaneous neutrophil apoptosis.7 Taken together, these findings and our results implicate B2 as a potential regulatory factor during organ development. The mitogen-activated protein kinase (MAPK) pathway exists in all eukaryotes, and controls such fundamental cellular processes including proliferation, differentiation, survival and apoptosis.15 Activation by B2 overexpression may not only affect apoptosis, but other cellular processes, such as cell cycle regulation and cell transformation. Other potential functions of B2 subunit in cell biology remain open to investigation.
Materials and Methods
Generation of stable cell lines expressing GFP-B2 fusion protein or GFP
The rat gene encoding the sequence of B2 subunit of H+-ATPase (gi:2058353; Y12635) was cloned into the expressing vector, pcDNA 3.1/NT-GFP. TOPO (Invitrogen, CA, USA). The right insertion and gene sequence was confirmed by DNA sequencing. Recombinant and control empty plasmids were transfected into HEK 293 cells using lipofectamine 2000 (Invitrogen). Cells were selected in DMEM medium containing G418 (800 μg/ml) and 10% FCS. Stable clones expressing the GFP-B2 fusion protein or GFP were evaluated by fluorescence microscopy and Western blot analysis for GFP expression. Clones with homogenous expression of GFP-B2 or GFP, termed 293B2 and 293GFP, respectively, were used for all experiments.
Induction of apoptosis by serum deprivation or staurosporine
293B2 and 293GFP cells were cultured in DMEM containing 10% FCS at 37°C and 5% CO2. When confluent, serum starvation was initiated by washing and replacing the medium with DMEM containing no FCS. Cells were harvested at the time points (0, 24, 48 and 72 h) after serum withdrawal for analysis. For experiments in which staurosporine was used as the agonist to induce apoptosis, cells were cultured in DMEM medium containing 10% FCS plus 2 μM staurosporine for 4 h, then cells were collected for apoptotic evaluation.
Hochest staining
Cultured cells were fixed with an ice-cold solution of methanol/acetic acid, (3 : 1 vol : vol) for 5 min at 4°C. After brief washing with distilled water, cells were stained with Hoechst dye #33342 (5 μg/ml in distilled water) for 10 min at 4°C. Cells displaying condensed fluorescent chromatin were identified as apoptotic.
Flow cytometry analysis
Cells (106/ml) fixed in 70% ethanol were washed with ice-cold PBS and then resuspended in 1 ml of 0.25% Triton X-100 in PBS, vortexing samples and working in ice for 5 min. After washing with PBS and treatment with RNase A (50 μg/ml in PBS) at room temperature for 5 min, cells were stained with propidium iodide (50 μg/ml) for 30 min at room temperature, then analyzed with a FACScan II® and Cell Quest® software. Cells with fragmented sub-G1 DNA content were categorized as apoptotic.
Staining of mitochondria and confocal microscope study
Cells were grown on coverslips inside Petri dishes filled with DMEM containing 10% FCS and 5% CO2 at 37°C. Medium was removed from the dish and replaced with the prewarmed fresh medium containing MitoTracker®Red CMXRos (100 nM). Cells were incubated for 30 min at 37°C. After washing with prewarmed DMEM, live cells were viewed with a confocal microscope with spectral characteristics appropriate for the MitoTracker® at Ex 568 nm/Em 600 nm and for GFP at Ex 395 nm/Em 478 nm.
Preparation of digitonin-soluble and -insoluble protein samples for detection of cytochrome c and AIF release from mitochondria
Cells were grown to confluence in 60 mm2 dishes with or without 10% FCS. At serial time points, the medium was removed and the cells were washed with PBS × 1, then 300 μl of a buffer containing in mM: 120 KCl; 5 KH2PO4; 10 HEPES; 2 EGTA at pH 7.4, to which 0.15 mg/ml digitonin and Protease inhibitors cocktail were added. The cells were constantly agitated for 15 min at 4°C. The liquid was aspirated and then centrifuged at 12 000 × g for 10 min at 4°C to remove whole cells or fragments. The supernatants were saved and designated as a digitonin-soluble (cytosolic) fraction. Adherent cells were mechanically removed with a rubber policeman in the presence of 0.5 ml 2% SDS in PBS and sonicated to solubilize proteins, then centrifuged at 12 000 × g for 15 min at 4°C. The pellet was discarded and the supernatant was designated as the digitonin-insoluble (membrane associated) fraction. Cytochrome c and AIF accumulation in cytosolic fraction is an index of mitochondrial release.
Isolation and purification of mitochondria
Isolations of mitochondria were performed using a mitochondria isolation kit (Pierce, Rockford, IL, USA). Cells were grown in 150 mm2 plate and were subjected to apoptosis stimuli as described above. Cells (2 × 107) were harvested and subjected to mitochondria isolation following the instructions provided. The 12 000 × g centrifuged supernatants were collected as to estimate ‘cytosolic fraction’ and the mitochondria pellets were dissolved in IP buffer containing 1% SDS and protease inhibitors as previously described by our laboratory.28
Immunoblotting and immunoprecipitation studies
Cells were lysed in ice-cold lysis buffer containing in mM (Tris, 20; NaCl, 150; EDTA, 1; EGTA, 1; sodium pyrophosphate, 1; β-glycerophosphate, 1; Na3VO4, 1; and PMSF, 1; with 1% Triton X-100 and a 1 × protease inhibitor tablet added just before use) at pH 7.5. Cell lysates were incubated for 30 min on ice, then centrifuged at 14 000 × g for 15 min at 4°C. The protein concentrations of all samples were determined by Bradford protein assay (Boston Bioproducts Inc., Worchester, MA, USA). For immunoprecipitation, 1000 μg protein of cell homogenate was incubated with 5 μl anti-GFP antibody (BD, San Jose, CA, USA) at 4°C overnight followed with protein A beads for 2 h at 4°C. The immunoprecipitates were washed and then resuspended in SDS-loading buffer for SDS-PAGE separation, transfer onto nitrocellulose membranes and immunoblot analysis. The membranes were blocked using 5% nonfat dry milk in TBST and incubated overnight at 4°C with primary antibody (detailed below). After incubation with appropriate secondary antibodies conjugated to horseradish peroxidase, the blots were developed using the enhanced chemiluminescence method. The film were scanned and then subjected to band densitometry and quantification (NIH Image Software). Antibodies used in this study are as follows: rabbit antibody to the 31-kDa subunit of the H+-ATPase (a gift of D Brown, Massachusetts General Hospital, Charlestown, MA, USA); rabbit polyclonal antibody to the 57-kDa B1/2 subunit of the H+-ATPase and mouse monoclonal antibody to GFP (Santa Cruz Biotechnology, Santa Cruz, CA, USA); mouse monoclonal antibodies to bcl-2 and bax (Oncogene, San Diego, CA, USA); mouse monoclonal antibody to total Bad (BD Transduction Laboratories); rabbit antibodies to phospho-Bad ser-112, ser 136 and ser-155, to phospho-p90RSK, total p90RSK, total AKT and phospho-AKT (Cell Signaling Technology, Danver, MA, USA). The secondary antibodies for the immunoblotting were peroxidase-conjugated goat anti-rabbit and -mouse IgG (Jackson ImmunoResearch, West Grove, PA, USA).
ATP/NADH-coupled assay for ATPase activity
This assay is based on reaction in which the regeneration of hydrolyzed ATP is coupled to the oxidation of NADH. Following each cycle of ATP hydrolysis, regeneration system consisting of phosphoenolpyruvate (PEP) and pyruvate kinase (PK) converts one molecule of PEP to pyruvate when the ADP is converted back to the ATP. The pyruvate is subsequently converted to lactate by L-lactate dehydrogenase (LDH) resulting in the oxidation of one NADH molecule. The ATPase hydrolysis rates were determined at 25°C using the change in NADPH absorbance at 340 nm within 30 min in a reaction mixture consisting of 200 μg sample proteins supplemented with 1 × ATPase reaction buffer: 25 mM TEA (triethanolamine hydrochloride) pH 7.5, 13 mM magnesium acetate, 1.8 mM DTT, 5 mM ATP, 200 μg/ml BSA, 3 mM PEP, 20 U/ml PK/LDH, NADH (10 μl 20 mg/ml stock solution added to 500 μl to initiate the assay).29, 30 When the effect of bafilomycin is tested, 1 μl of 0.5 M bafilomycin stock solution (dissolved in DMSO) were added into each reaction mixture right before the initiation of the reaction.
Data are expressed as means±S.E. Data were analyzed by analysis of variance and paired or unpaired t-test as appropriate. Statistical significance was assumed at P<0.05.
Abbreviations
- MAPK:
-
mitogen-activated protein kinase
- ERK:
-
extracellular-signal-regulated kinase
- MEK:
-
MAPK/ERK kinase
- RSK:
-
ribosomal S6 kinase
- PKA:
-
cAMP-activated protein kinase
- PKC:
-
protein kinase C
- AKT:
-
protein kinase B
- AIF:
-
apoptosis-inducing factor
- GFP:
-
green fluorescent protein
References
Wagner CA, Finberg KE, Breton S, Marshansky V, Brown D and Geibel JP (2004) Renal vacuolar H+-ATPase. Physiol. Rev. 84: 1263–1314.
Hishita T, Tada-Oikawa S, Tohyama K, Miura Y, Nishihara T, Tohyama Y, Yoshida Y, Uchiyama T and Kawanishi S (2001) Caspase-3 activation by lysosomal enzymes in cytochrome c-independent apoptosis in myelodysplastic syndrome-derived cell line P39. Cancer Res. 61: 2878–2884.
Torigoe T, Izumi H, Ishiguchi H, Uramoto H, Murakami T, Ise T, Yoshida Y, Tanabe M, Nomoto M, Itoh H and Kohno K (2002) Enhanced expression of the human vacuolar H+-ATPase c subunit gene (ATP6L) in response to anticancer agents. J. Biol. Chem. 277: 36534–36543.
Aiko K, Tsujisawa T, Koseki T, Hashimoto S, Morimoto Y, Amagasa T and Nishihara T (2002) Involvement of cytochrome c and caspases in apoptotic cell death of human submandibular gland ductal cells induced by concanamycin A. Cell Signal 14: 717–722.
Zhan H, Yokoyama K, Otani H, Tanigaki K, Shirota N, Takano S and Ohkuma S (2003) Different roles of proteolipids and 70-kDa subunits of V-ATPase in growth and death of cultured human cells. Genes Cells 8: 501–513.
Ishisaki A, Hashimoto S, Amagasa T and Nishihara T (1999) Caspase-3 activation during the process of apoptosis induced by a vacuolar type H(+)-ATPase inhibitor. Biol. Cell 91: 507–513.
Niessen H, Meisenholder GW, Li HL, Gluck SL, Lee BS, Bowman B, Engler RL, Babior BM and Gottlieb RA (1997) Granulocyte colony-stimulating factor upregulates the vacuolar proton ATPase in human neutrophils. Blood 90: 4598–4601.
Lee BS, Krits I, Crane-Zelkovic MK and Gluck SL (1997) A novel transcription factor regulates expression of the vacuolar H+-ATPase B2 subunit through AP-2 sites during monocytic differentiation. J. Biol. Chem. 272: 174–181.
Anderson CL and Williams GT (2003) Apoptosis gene hunting using retroviral expression cloning: identification of vacuolar ATPase subunit E. Sci World J 3: 51–58.
Hengartner MO (2000) The biochemistry of apoptosis. Nature 407: 770–776.
van Loo G, Saelens X, van Gurp M, MacFarlane M, Martin SJ and Vandenabeele P (2002) The role of mitochondrial factors in apoptosis: a Russian roulette with more than one bullet. Cell Death Differ. 9: 1031–1042.
Susin SA, Lorenzo HK, Zamzami N, Marzo I, Snow BE, Brothers GM, Mangion J, Jacotot E, Costantini P, Loeffler M, Larochette N, Goodlett DR, Aebersold R, Siderovski DP, Penninger JM and Kroemer G (1999) Molecular characterization of mitochondrial apoptosis-inducing factor. Nature 397: 441–446.
Datta SR, Ranger AM, Lin MZ, Sturgill JF, Ma YC, Cowan CW, Dikkes P, Korsmeyer SJ and Greenberg ME (2002) Survival factor-mediated Bad phosphorylation raises the mitochondrial threshold for apoptosis. Dev. Cell 3: 631–643.
Zhou XM, Liu Y, Payne G, Lutz RJ and Chittenden T (2000) Growth factors inactivate the cell death promoter Bad by phosphorylation of its BH3 domain on Ser155. J. Biol. Chem. 275: 25046–25051.
Kolch W (2000) Meaningful relationships: the regulation of the Ras/Raf/MEK/ERK pathway by protein interactions. Biochem. J. 2: 289–305.
Datta SR, Dudek H, Tao X, Masters S, Fu H, Gotoh Y and Greenberg ME (1997) Akt phosphorylation of Bad couples survival signals to the cell-intrinsic death machinery. Cell 91: 231–241.
Blume-Jensen P, Janknecht R and Hunter T (1998) The kit receptor promotes cell survival via activation of PI3-kinase and subsequent Akt-mediated phosphorylation of Bad on Ser136. Curr. Biol. 8: 779–782.
Bertolotto C, Maulon L, Filippa N, Baier G and Auberger P (2000) Protein kinase C theta and epsilon promote T-cell survival by a rsk-dependent phosphorylation and inactivation of BAD. J. Biol. Chem. 275: 37246–37250.
Tan Y, Ruan H, Demeter MR and Comb MJ (1999) p90(RSK) blocks bad-mediated cell death via a protein kinase C-dependent pathway. J. Biol. Chem. 274: 34859–34867.
Harada H, Becknell B, Wilm M, Mann M, Huang LJ, Taylor SS, Scott JD and Korsmeyer SJ (1999) Phosphorylation and inactivation of Bad by mitochondria anchored protein kinase A. Mol. Cell 3: 413–422.
Kelemen BR, Hsiao K and Goueli SA (2002) Selective in vivo inhibition of mitogen-activated protein kinase activation using cell-permeable peptides. J. Biol. Chem. 277: 8741–8748.
Brognard J and Dennis PA (2002) Variable apoptotic response of NSCLC cells to inhibition of the MEK/ERK pathway by small molecules or dominant negative mutants. Cell Death Differ. 9: 893–904.
Bonni A, Brunet A, West AE, Datta SR, Takasu MA and Greenberg ME (1999) Cell survival promoted by the Ras-MAPK signaling pathway by transcription-dependent and independent mechanism. Science 286: 1358–1362.
Chiang CW, Kanies C, Kim KW, Fang WB, Parkhurst C, Xie M, Henry T and Yang E (2003) Protein phosphatase 2A dephosphorylation of phosphoserine 112 plays the gatekeeper role for Bad-mediated apoptosis. Mol. Cell Biol. 23: 6350–6362.
Schurmann A, Mooney AF, Sanders LC, Sells MA, Wang HG, Reed JC and Bokoch GM (2000) p21-activated kinase 1 phosphorylates the death agonist Bad and protects cells from apoptosis. Mol. Cell Biol. 20: 453–461.
Tan Y, Demeter MR, Ruan H and Comb MJ (2000) Bad Ser-155 phosphorylation regulates Bad/Bcl-XL interaction and cell survival. J. Biol. Chem. 275: 25865–25869.
Jouret F, Auzanneau C, Debaix H, Wada GH, Pretto C, Marbaix E, Karet FE, Courtoy PJ and Devuyst O (2005) Ubiquitous and kidney-specific subunits of vacuolar H+-ATPase are differentially expressed during nephrogenesis. J. Am. Soc. Nephrol. 16: 3235–3246.
Ruchalski K, Mao H, Singh SK, Wang Y, Mosser DD, Li F, Schwartz JH and Borkan SC (2003) HSP72 inhibits apoptosis-inducing factor release in ATP-depleted renal epithelial cells. Am. J. Physiol. 285: C1483–C1493.
Huang TG and Hackney DD (1994) Drosophilia kinesin minimal motor domain expressed in Escherichia coli. J. Biol. Chem. 269: 16493–16501.
Kiianitsa K, Solinger JA and Heyer WD (2003) NADH-coupled microplate photometric assay for kinetic studies of ATP-hydrolyzed enzymes with low and high specific activities. Anal. Biochem. 321: 266–271.
Acknowledgements
This research is supported by grants obtained from National Institutes of Health (National Institute of Diabetes and Digestive and Kidney Diseases): JH Schwartz (DK-52898) and (DK 59529) and SC Borkan (DK-53387). We also appreciate the gift of rabbit antibody to the 31 kD subunit H+-ATPase from Dr Dennis Brown.
Author information
Authors and Affiliations
Corresponding author
Additional information
Edited by JA Cidlowski
Supplementary Information accompanies the paper on Cell Death and Differentiation website (http://www.nature.com/cdd)
Rights and permissions
About this article
Cite this article
Li, G., Yang, Q., Krishnan, S. et al. A novel cellular survival factor – the B2 subunit of vacuolar H+-ATPase inhibits apoptosis. Cell Death Differ 13, 2109–2117 (2006). https://doi.org/10.1038/sj.cdd.4401970
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.cdd.4401970
Keywords
This article is cited by
-
Interaction between V-ATPase B2 and (Pro) renin Receptors in Promoting the progression of Renal Tubulointerstitial Fibrosis
Scientific Reports (2016)
-
The medaka mutation tintachina sheds light on the evolution of V-ATPase B subunits in vertebrates
Scientific Reports (2013)
-
NMR solution structure of subunit E (fragment E1–69) of the Saccharomyces cerevisiae V1VO ATPase
Journal of Bioenergetics and Biomembranes (2011)
-
Cytosolic pH is a second messenger for glucose and regulates the PKA pathway through V-ATPase
The EMBO Journal (2010)