Abstract
Three-dimensional DNA nanoscaffolds such as supramolecular tetrahedra can self-assemble from tris-oligonucleotidyls — synthetic three-armed building blocks in which three identical or non-identical short DNA sequences are connected by a tris-linking backbone1,2. Here we show that the connectivity information contained in these building blocks can be copied by using template-directed tris-linking. This finding is a crucial step towards the replication of nanoarchitectures that are based on tris-oligonucleotidyls and to the realization of artificially self-replicating systems on a nanometre scale.
Similar content being viewed by others
Main
The procedure is outlined in Fig. 1a. On the basis of the three template sequences of an asymmetric tris-15-nucleotide oligomer (ref. 2), we generated the three corresponding complementary linear sequences with a reactive group at their 5′ ends, and used these in different ligation reactions3,4,5 with a suitable tris-linker molecule.
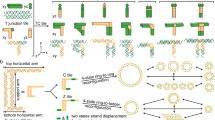
a, Left, chemical copying of connectivity involves a 3′-trisoligonucleotidyl template with three individually defined sequences, three linear complements with 5′ ends represented as blunt ends, and a tris-linker shown as a honeycomb cap. Centre, hybridization gives rise to a quartermolecular complex in which the blunt ends come into close spatial proximity. Right, the tris-linker has reacted to connect the 5′ ends in the copy. b, Chemical copying of connectivity on the basis of tris-hydrazone formation. Tris-linking of the 5′-hydrazide-modified oligonucleotides A, B and C with the tris-linker L in the presence of the 3′-tris-oligonucleotidyl template Y yields the tris-hydrazone H. c, Nucleic-acid staining (with SYBR gold), after denaturing polyacrylamide-gel electrophoresis, of an almost complete chemical copying of connectivity from a 10 µM solution of the components in 100 mM MOPS buffer (pH 5.6) after incubation for 2 h at room temperature in the presence or absence of template Y, an excess (2.5:1.0 equivalent) of tris-linker L, and sodium cyanoborohydride (NaCNBH3; 100 equivalents).
Template-directed tris-linking was rapid and robust — unhampered by hydrolysis — in the hydrazone-formation reaction shown in Fig. 1b. Three 5′-hydrazide-modified oligonucleotides, a 15-nucleotide molecule (15-mer; A), a 13-mer (B) and an 11-mer (C), the sequences of which are complementary to the respective arms of the tris-15-meric template Y, were synthesized by using standard solid-phase protocols. The 5′-hydrazide moieties were introduced by using a modifier amidite6.
For a statistical, non-template-directed synthesis, ten 5′-tris-linked species would be expected, ranging from a branched 33-mer to a branched 45-mer, as well as six bis-linked species containing between 22 and 30 nucleotides. A template-directed synthesis should yield a single branched 39-mer, as well as three of the six possible bis-linked species, if the reaction has not reached completion. Figure 1c shows that the predicted 39-mer is formed as the principal product in the presence of the template (confirmed by MALDI mass spectroscopy); in the absence of the template, there are no tris-linked products detectable by polyacrylamide-gel electrophoretic analysis. There is also evidence that the template directs the formation of the intermediate bis-linked species, the distribution of which differs from that in the statistical synthesis.
The copies are stable, even without reductive amination in the presence of sodium cyanoborohydride, which has been used successfully in other imine-based, template-directed ligation reactions7. We also found that short-armed tris-9-mers can be used as templates to generate asymmetrical tris-30-mers, which are otherwise not easily accessible by chemical synthesis. Tris-linking was also possible with 3 + 1-armed templates2, in which one arm is connected through its 5′ end. The 'Klenow' polymerase enzyme accepts the 3′ end of tris-15-mer copies hybridized to overhanging linear templates as extension sites.
A full replication cycle also requires reverse copying — that is, the tris-linking of 3′-modified oligonucleotides using 5′-tris-oligonucleotide templates. Once this step becomes feasible, surface-promoted replication and exponential amplification of DNA analogues (SPREAD)8 may be adapted to connectivity templating to achieve multiple rounds of replication. For self-assembling, non-covalent nanostructures, including tetrahedral nano-scaffolds of modular functions2, replication will depend only on the ability of the subunits to replicate. Non-covalent connectivity therefore constitutes transferable information that can instruct a subunit where to integrate during self-assembly. Covalent connectivity and/or connectivity by topological interweaving, as used in Seeman's DNA cube9, cannot replicate but ensures ultimate stability. Switchable duplex crosslinkers, such as those based on disulphide bonds10, may be suitable for stabilizing replicatable nano-objects.
References
Scheffler, M., Dorenbeck, A., Jordan, S., Wüstefeld, M. & von Kiedrowski, G. Angew. Chem. Int. Ed. 38, 3311–3315 (1999).
Dorenbeck, A., Scheffler, M., Wüstefeld, M. & von Kiedrowski, G. Angew. Chem. (in the press).
Dawson, P. E., Churchill, M. J., Ghadiri, M. R. & Kent, S. B. H. J. Am. Chem. Soc. 119, 4325–4329 (1997).
Stetsenko, D. A. & Gait, M. J. J. Org. Chem. 65, 4900–4908 (2000).
Cochran, J. R. & Stern, L. J. Chem. Biol. 7, 683–696 (2000).
Raddatz, S. et al. Nucleic Acids Res. 30, 4793–4802 (2002).
Li, X., Zhan, Z. Y. J., Knipe, R. & Lynn, D. G. J. Am. Chem. Soc. 124, 746–747 (2002).
Luther, A., Brandsch, R. & von Kiedrowski, G. Nature 396, 245–248 (1998).
Chen, J. & Seeman, N. C. Nature 350, 631–633 (1991).
van Aalten, D. M. F., Erlanson, D. A., Verdine, G. L. & Leemor, J. T. Proc. Natl Acad. Sci. USA 96, 11809–11814 (1999).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
brief communications is intended to provide a forum for both brief, topical reports of general scientific interest and technical discussion of recently published material of particular interest to non-specialist readers. Priority will be given to contributions that have fewer than 500 words, 10 references and only one figure. Detailed guidelines are available on Nature's website (http://www.nature.com/nature) or on request from nature@nature.com
Rights and permissions
About this article
Cite this article
Eckardt, L., Naumann, K., Matthias Pankau, W. et al. Chemical copying of connectivity. Nature 420, 286 (2002). https://doi.org/10.1038/420286a
Issue Date:
DOI: https://doi.org/10.1038/420286a
This article is cited by
-
Exponential growth and selection in self-replicating materials from DNA origami rafts
Nature Materials (2017)
-
Nanostructure evolution
Nature Materials (2017)
-
Controlled release of an anti-cancer drug from DNA structured nano-films
Scientific Reports (2014)
-
Fabrication of pRNA nanoparticles to deliver therapeutic RNAs and bioactive compounds into tumor cells
Nature Protocols (2013)
-
Systems chemistry: from chemical self-replication to trisoligo-based nanoconstruction
Journal of Cheminformatics (2010)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.