Abstract
Hepatitis C virus is a major risk factor for hepatocarcinogenesis in humans. In situ detection of the virus in early sequential lesions of hepatocarcinogenesis could provide information about the role of the virus in the transformation and promotion process. Parallel in situ detection of HCV proteins and RNA in human tissues were performed in 55 posthepatitis C cirrhosis, 17 dysplastic nodules (DN), and 25 hepatocellular carcinomas (HCC), using immunohistochemistry and tissue quantitative RT-PCR. A consistent cytoplasmic hepatocellular staining was obtained in 73% of cirrhosis cases (with or without HCC) and in 55% DN cases. A few tumoral hepatocytes were unambiguously stained in 28% HCC. The percentage of positive cells and the intensity of immunostaining significantly decreased from cirrhosis to HCC through DN, whereas there was no difference in the prevalence of positivity or the number of viral copies between cirrhosis and HCC using tissue-quantitative RT-PCR. Finally, RT-PCR levels were found parallel with the immunostaining in cirrhosis but not in HCC. These results suggest that HCV protein synthesis may persist but be down-regulated during sequential hepatocarcinogenesis. A putative role of HCV proteins on cell proliferation and differentiation during the early steps of carcinogenesis cannot therefore be excluded.
Similar content being viewed by others
INTRODUCTION
Studies of rodent models of hepatocarcinogenesis and, more recently, pathological examination of cirrhotic liver explants in humans have provided evidence that hepatocarcinogenesis is a sequential process (1). Indeed, in human cirrhotic livers, some of the hepatocellular carcinoma (HCC) may develop from early neoplastic lesions recently reported as dysplastic nodules (DN; 2). In Western countries and Japan, hepatitis C virus (HCV) is the main risk factor for cirrhosis and HCC, and DN are frequently diagnosed in HCV-induced cirrhosis (3, 4, 5). However, despite significant advances in the knowledge of HCV biology, little is known concerning the mechanisms involved in HCV-associated hepatocarcinogenesis. Clearly, chronic hepatitis, with its association of chronic inflammation, liver cell necrosis and regeneration, and extensive fibrosis, is a major step in this process (6). Another relevant hypothesis is that HCV acts more directly on hepatocytes via viral proteins. This is supported by a series of arguments: HCC can occur in HCV-positive livers without inflammation and cirrhosis (7) and some in vitro experiments have shown that HCV core proteins NS3 and NS5A may play a role in the modulation of cell phenotype, cell proliferation, viability, or apoptosis or may favor clonal cell expansion (8). However, most of these data come from cell culture models or molecular studies performed on blood or tissue extracts. Very few morphologic studies have tried to find relevant information in this field, and the results are still controversial (9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21).
In the present work, we have developed in situ detection of HCV proteins in human tissues from cirrhosis, DN, and HCC using an immunohistochemistry procedure (IHC), and we have quantitated HCV-RNA in the same tissue samples of cirrhosis and HCC, with a quantitative reverse transcription-polymerase chain reaction (qtRT-PCR) assay to find relevant information about HCV-associated sequential hepatocarcinogenesis in human tissues.
MATERIAL AND METHODS
Tissues
Fifty-five patients who underwent liver transplantation for cirrhosis secondary to HCV were the study population. After gross and microscopic examination of the 55 corresponding liver explants, they were divided into two groups: 31 with neoplasia, either DN or HCC (Group 1); and 24 without (Group 2). In both groups, tissue specimens from the different lesions (cirrhosis and, when present, DN and HCC) were taken, frozen in liquid nitrogen–cooled isopentane, and stored at −80°C before use. As a result, we could analyze 31 cirrhotic specimens, 17 DN specimens, and 25 HCC specimens from Group 1; and 24 cirrhotic specimens from Group 2, for a total of 97 specimens.
In Group 1, cirrhoses were mostly micronodular or mixed (n = 29/31), with moderate activity (n = 22/31) according to the METAVIR system (22) and frequent liver cell dysplasia (small- and/or large-cell dysplasia, n = 24/31). The mean diameter of the 25 HCC found in Group 1 was 17 mm (range, 2–60 mm). The tumors were almost always trabecular in architecture (n = 22/25) and moderately differentiated (n = 19/25); that is, they were Grade II or III according to Edmondson (23). The mean diameter of the 17 DN was 11 mm (range, 7–20 mm). Microscopically, 12 DN were classified as low grade, and 5 as high grade, as defined by the international consensus (2).
The cirrhotic livers in Group 2 (n = 24) had similar histological characteristics as in Group 1: the cirrhoses were mostly of micronodular or mixed type (n = 22/24), with moderate activity (n = 18/24), and areas of small- and/or large-cell dysplasia were found in half of the cases (n = 11/24).
Patients
All patients (n = 55) were HCV antibody (Ab) positive in serum (ELISA 2 and 3 and RIBA III tests). They were 38 males and 17 females, with a mean age of 54.5 years (range, 29 to 68). They underwent liver transplantation for either Child C–decompensated cirrhosis (n = 37) or known HCC in cirrhosis (n = 11); in seven cases, HCC was incidentally discovered on the explanted liver. In a small proportion of patients (9/31 in Group 1 and 6/24 in Group 2), there was a history of interferon therapy before transplantation. Results of qualitative HCV RNA detection in serum before transplantation were also available in 14 of the 25 patients with HCC and were positive in 10 cases (Table 1). The remaining patients were positive for HCV antibody in serum, had a medical history and histology in favor of HCV infection, and had no other factor of chronic liver disease.
Immunohistochemistry
IHC was performed in 97 samples (31 of cirrhosis with and 24 of cirrhosis without HCC; 17 DN; and 25 HCC), using a modification of a procedure described elsewhere (24). Briefly, 5-μm-thick frozen sections were incubated overnight at 4°C with a fluorescein-conjugated, purified IgG fraction from a pool of spontaneous human polyclonal antibody (Ab) isolated from a same HCV-human sera and directed against C100 (NS4), C33 (NS3), C22 (core), and NS5 viral proteins (same batch as described earlier, 24), which was diluted 1:20 in phosphate buffered saline (PBS) containing 50% normal human AB serum. After washing in PBS (twice for 10 min each), the sections were incubated for 30 minutes with a mouse monoclonal Ab to fluorescein (DAK-FITC4, DAKO A/S, Glostrup, Denmark) diluted 1:50 in PBS containing 5% normal human AB serum. After rinsing in PBS (twice for 10 min each), the revelation technique consisted in incubating the sections for 30 minutes with a drop of goat anti-mouse immunoglobulins conjugated to peroxidase-labeled polymer (EnVision TM+ anti-mouse HRP, DAKO), as stated by the manufacturer. After a rinse in PBS buffer, the colored reaction was developed for 3–4 minutes with diamino-benzidine (Liquid 3,3′-diaminobenzidine + large volume substrate-chromogen solution, DAKO). Sections were counterstained with hemalum, dehydrated in graded alcohol, cleared in xylol, and mounted with Eukitt. Negative controls consisted of 13 liver tissues from patients who were HCV-Ab negative in serum. Seven cases had cirrhosis due to hepatitis B virus (HBV; n = 3), alcohol (n = 3), or primary biliary cirrhosis (n = 1); another control showed lesions of HBV chronic hepatitis without cirrhosis, and the remaining five liver samples were histologically normal or subnormal (n = 5) and came from partial hepatectomies for colorectal metastasis. Other controls consisted of the use of a normal fluorescence isothiocyanate (FITC)-conjugated polyclonal Ab instead of the anti-HCV Ab and the omission of the primary Ab, both of which were applied to HCV-positive tissues.
For analysis of the results, the location of the staining was first evaluated within the tissues and at the cellular and subcellular levels. Thereafter, the percentage of positive cells was semi-quantitated as follows: 0, 0%; 1+, <10%; 2+, 10–50%; 3+, >50% positive cells. Furthermore, the intensity of staining, whether low, moderate, or strong, was appreciated. Because it was often heterogeneous in a single sample, the intensity observed in the largest number of cells was considered. This evaluation had been shown to be reproducible in a previous study with four observers (BLB, PBS, and two other members of our group; data not shown).
HCV RNA Quantitation in Frozen Liver Tissues
This technique was performed in the tissues from the 25 patients of Group 1 who presented with both cirrhosis and HCC. Tissue fragments were taken from the same frozen blocks used for IHC and digested for 1 hour at 55°C in 400 μl of proteinase K reagent (10 mM Tris, 1 mM ethylenediaminetetraacetic acid, 250 μg proteinase K, 1% SDS). The amount of HCV RNA in each biopsy was quantified by RT-PCR using the Amplicor HCV Monitor 1.0 assay (Roche Diagnostic Systems, Branchburg, NJ; 25). The technique was adapted from the Agence Nationale de Recherche sur le SIDA method used for HIV RNA quantitation in lymph nodes (26). Briefly, 100 μl of the proteinase K–digested specimens were mixed with 400 μl of Monitor lysis buffer containing guanidine thiocyanate with addition of the Roche-supplied internal quantitative standard (IQS) required for determining RNA levels. IQS is an in vitro–transcribed RNA molecule with HCV primer-binding sites and a unique probe-binding site. IQS is added at a known concentration to each specimen to provide a reference for quantitation of HCV-RNA.
After lysis, the RNA was precipitated by isopropyl alcohol and washed in 70% ethanol. At this stage, the pellet was digested with 4 U of an RNAse-free DNAse I (Promega, Madison, WI) for 60 minutes at 37°C to eliminate the residual DNA. The total RNA yield was determined by spectrophotometric readings at 260 nm, and the pellet was resuspended in 1 ml of specimen diluent. Fifty microliters of this suspension containing total RNA in known quantity was mixed with 50 μl of the PCR Master Mix. The combined RT-PCR was performed in a Perkin-Elmer GeneAmp PCR System 9600 thermal cycler (Perkin-Elmer, Norwalk, CT). When complete, 100 μl of denaturation solution was added to each PCR tube. After denaturation, a series of 5-fold dilutions of the biotinylated amplicons was performed in the appropriate wells coated with HCV-specific probe. HCV RNA was detected by addition of 100 μl of avidin horseradish peroxidase conjugate followed by 100 μl of substrate and stopping reagent with the appropriate washing solution. The concentration of native viral RNA was calculated using the manufacturer’s formula, which is based on the ratio of native-bound HCV RNA compared with the internal quantitation standard. For serum samples, the limit of detection (cutoff) of the assay quoted by the manufacturer is 5 × 102 HCV RNA copies per milliliter. The results, in HCV RNA copies per milliliter, which were obtained automatically using Roche’s computerized program, were converted into HCV RNA copies per microgram of total liver RNA. The results were calculated as follows: the value of HCV copies per milliliter was divided successively by 200, as optical densities were produced from 100 μl of serum and 50 μl of extracted RNA and converted automatically into copies per milliliter and by the quantity of total RNA present in the 50 μl of extracted RNA used for PCR reaction. The sensitivity of the assay was four copies per microgram of RNA, calculated as the minimum detectable quantity, which is five standard deviations above the mean copy number of five duplicate Amplicor HCV Monitor negative controls (27). The reproducibility of RT-PCR results was assessed by replicates performed in 10 cases. Negative- and positive-kit controls were included in each run. Two patients who received allogenic liver transplants for reasons other than HCV infection and who were proven to be negative by serum polymerase chain reaction were also included as negative controls.
Statistical Analysis
The results of IHC were compared using a χ2 test. The values of qtRT-PCR were expressed as mean ± SEM and as median and range. Comparison of quantitative data was performed using the non-parametric Mann-Whitney test and Kruskal-Wallis test as appropriate.
RESULTS
Immunohistochemistry
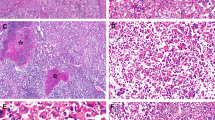
The specific staining was granular or powderlike, as described elsewhere (24). It was exclusively located in the cytoplasm of hepatocytes (Fig. 1A). On very rare occasions, a few hepatocytes also showed slight nuclear staining that was not found in negative controls. Lymphocytes, biliary, endothelial, and Kupffer cells were always negative (Fig. 1A). The positive hepatocytes were distributed at random in the tissue, either tumoral or nontumoral. The intensity of staining was generally heterogeneous (from low to strong) in a given case (Fig. 1B). Negative results were obtained with FITC-conjugated non-HCV normal polyclonal Ab and with the omission of the primary Ab (Fig. 1C) and in HCV-negative tissues (Fig. 1D).
Immunohistochemical detection of hepatitis C virus (HCV) in cirrhosis (A–D) and hepatocellular carcinoma (E–H). A, In most cirrhotic tissues, a staining was obtained in many hepatocytes distributed at random, without zonation, and other cell types (inflammatory, biliary, and Kupffer cells) were negative (80 ×). B, At higher magnification, the staining of hepatocytes was mostly cytoplasmic and granular, with variable intensity (low, moderate, or strong) from one cell to another (260 ×). Negative controls, consisting of the omission of the primary antibody on a positive tissue (C) or detection on a tissue from HVC-negative patients (D), were satisfactory (80 ×). Although most hepatocellular carcinomas (star) were negative for HCV immunohistochemistry, contrasting with the surrounding cirrhotic tissue (E, 130 ×), some tumors (star) displayed a variable number of scattered positive cells (F, 130 ×). The HCV proteins could be detected either in well-differentiated (G) or poorly differentiated tumors (H) and were located in the cytoplasm of tumor cells, sometimes around vacuoles of steatosis, and was usually low or moderate in intensity (G–H, 500 ×).
Seventy-three percent (40/55) of all cirrhosis cases were positive for HCV immunostaining. In these cases, the staining was rarely 1+ (7/40 = 17.5%) and was mostly 2+ (19/40 = 47.5%) or 3+ (14/40 = 35%). The intensity was mostly moderate (24/40 = 60%) and was strong on occasion (10/40 = 25%). There was no group difference in staining in the cirrhosis; that is, cirrhosis with DN and/or HCC (Group 1: 22/31), and cirrhosis without DN and/or HCC (Group 2: 18/24) were the same in terms of rate of positivity, percentage of positive cells, and intensity (Table 2).
Within Group 1, positive staining was observed in 70% (22/31), 59% (10/17), and 28% (7/25) of cirrhosis, DN, and HCC respectively (Table 3). In this group, the percentage of positive cells in cirrhosis was 2 to 3+ in the majority of cases (17/31, 55%), whereas in DN and HCC, it was ≦1+ in most cases (11/17, 65% and 22/25, 88%, respectively; χ2, P = .0039; degree of freedom = 2). The most frequent intensity found in cirrhosis was moderate to strong (19/22 = 86%), whereas in DN and HCC, it was low or undetectable (12/17 = 71% and 20/25 = 80%, respectively; χ2, P = .004; degrees of freedom = 2; Table 3; Fig. 1E–F). There was no difference in staining (in terms of rate of positivity, percentage of positive cells, and intensity of staining) in the group of DN according to their size and grade. There was also no difference in staining in the HCC group, whatever the size, grade, or architecture of the tumor (Fig. 1G–H), and the distribution of the positive cells was at random. In each individual case for which the three types of lesions were available (i.e., cirrhosis, DN, and HCC; n = 11), the percentage of positive cells and the intensity of staining were almost always lesser in the HCC specimen than in the cirrhosis specimen, with DN specimen(s) showing intermediate values (Table 4).
Quantitative RT-PCR
Using qtRT-PCR, the frequency of positivity was 84% (21/25) in cirrhosis and 72% (18/25) in HCC from the same livers previously tested by IHC. The mean values were 720 ± 211 (median = 154; range = 0 to 3720) HCV RNA copies per microgram of total extracted RNA in cirrhosis and 369 ± 119 (median = 54; range = 0 to 2151) per microgram of total extracted RNA in HCC, which was not significantly different (Mann-Whitney test = 257.50, P = .28, n.s.; Table 1). However, in 10 individual cases, qtRT-PCR value was lower in HCC than in the underlying cirrhosis. In eight cases, the values were identical in cirrhosis and HCC, and in the seven remaining cases, the number of HCV RNA copies was higher in HCC than in cirrhosis.
Comparison between IHC and qtRT-PCR
Results of qtRT-PCR were compared with those of IHC, defined either as a percentage of positive cells or as the intensity of staining, for cirrhotic (Fig. 2A–B) and carcinomatous tissues (Fig. 2C–D) from Group 1. In cirrhosis, when the percentage of positive cells by IHC was ≦1+, the mean value of qtRT-PCR was 9 ± 6 HCV RNA copies (median = 4, range = 0 to 55). When the percentage of positive cells was 2 or 3+, the mean value of qtRT-PCR was 800 ± 441 (median = 290.5, range = 4 to 3720) or 1440 ± 356 (median = 1274, range = 89 to 3118) HCV RNA copies, respectively. Comparison of the mean value of qtRT-PCR showed a statistically significant difference among the three groups (Kruskal-Wallis = 15.54, P = .0004; Fig. 2A). When the intensity of the staining was low or undetectable, the mean value of qtRT-PCR was 9 ± 7 copies (median = 2, range = 0 to 55), and when the intensity was moderate or strong, it reached a mean of 886 ± 361 (median = 282, range = 4 to 3720) or 1363 ± 432 (median = 1025, range = 299 to 3118) HCV RNA copies, respectively. Comparison of the mean value of qtRT-PCR showed a statistically significant difference among the three groups of intensity (Kruskal-Wallis test = 14.58, P = .0007; Fig. 2B).
Comparison of tissue-quantitative reverse-transcription polymerase chain reaction (RT-PCR) and immunohistochemistry in cirrhosis (A, B) and hepatocellular carcinoma (C, D) from Group 1. A, Comparison of RT-PCR and percentage of positive cells by IHC in cirrhosis indicates that the tissular viral load is significantly different in the three groups defined by the percentage of positive cells and that PCR and immunohistochemistry results grow parallel. B, RT-PCR and intensity of staining in cirrhosis: although viral load grows parallel in the three classes of intensity. C, RT-PCR and percentage of positive cells in HCC. D, RT-PCR and intensity of staining in HCC: no parallelism between RT-PCR and immunohistochemistry. Results are expressed as mean values ± SEM.
In HCC, the percentage of positive cells varied from 0 to 1+ (0 to 10%) to 2+ (10 to 50%), and corresponding qtRT-PCR levels were 324 ± 122 (median = 39, range = 0 to 2151) and 700 ± 447 (median = 553, range = 11 to 1537) copies per microgram of total extracted RNA, respectively. The results were not statistically different (Mann-Whitney = 19.000, P = .273, n.s.; Fig. 2C). The intensity of staining varied from low or undetectable to moderate and strong and corresponding qtRT-PCR, from 235 ± 95 (median = 21, range = 0 to 1446), 596 ± 333 (median = 418, range, 11 to 1537), and 2151 (one case) HCV RNA copies, respectively, without significant difference between the first two groups (Mann-Whitney = 19.000, P = .114, n.s.; Fig. 2D).
DISCUSSION
In this study, we have adapted two feasible and sensitive technics for the detection of HCV in tissues, one morphological and semiquantitative (IHC), and one quantitative (qtRT-PCR), which could be used in routine practice when frozen tissues are available. For IHC, we used a purified fraction of polyclonal IgG human isolated from a spontaneous human sera directed against structural (C22-3, corresponding to core protein) and nonstructural (C33 and C100-3, corresponding to NS3 and NS4, respectively, and NS5) viral proteins (24). Modification of the original protocol of IHC (24) consisted of changing the simple peroxidase-labeled anti-mouse immunoglobulin of the third step for a peroxidase-labeled dextran polymer to which many anti-mouse immunoglobulins are attached (DAKO, Envision system). This resulted in a superior sensitivity and specificity and allowed us to clearly reclassify some doubtful results into definitely weak positive staining or negative staining. Using this protocol, we obtained a high rate of positivity in the group of cirrhosis (73% of cases) with a high percentage of positive cells (>50% in more than one third of cases and >10% in 82% of cases). Taking into account the cirrhotic status of all our specimens, the probable long delay since viral contamination (28), and the history of antiviral therapy in some patients, these results suggest a high sensitivity. The pattern of staining was also very similar to that described by Ballardini and collaborators (24) and showed an excellent reproducibility. In the literature, papers about HCV in situ detection using IHC, in situ hybridization (ISH), or in situ RT-PCR are scarce, the protocols have not been standardized, and the results are controversial (9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21). Among the various techniques tested in our laboratory, such as ISH with radioactive or nonradioactive oligoprobes or riboprobes, in situ RT-PCR, or HIC with Tordji 22 antibody (data not shown), the IHC protocol described in this paper was the most feasible and specific in our hands. Although it is not suitable for formalin-fixed, paraffin-embedded tissues at this time, we hope that progress in antigen retrieval will allow such work very soon.
We also elaborated a method of detection of HCV RNA in tissues. Indeed, tissular RT-PCR is regarded as the most sensitive method for detecting HCV (9), but most of the studies have used qualitative RT-PCR (20, 29, 30, 31, 32). For our quantitative RT-PCR in tissue, we used a commercial kit previously elaborated for blood tests; the process was quite easy to perform, with good feasibility, in our own laboratory. In our hands, the sensitivity of qtRT-PCR in cirrhotic tissue was high, but not significantly superior to that of IHC (84 versus 72%). Recently, the first-generation Roche assay for HCV (Version 1.0) used in this study has been shown to underestimate viral loads in HCV infections with Genotypes 2 and 3 (25, 33, 34). The lack of correlation between IHC and qtRT-PCR for some patients (Patients 3, 10, and 12 for example) may be explained by an incomplete detection of Genotypes 2 and 3 with Version 1.0 assay. Unfortunately, in this retrospective study, serum samples were not available for all patients to confirm the genotypes. Finally, both methods, IHC and qtRT-PCR, were sensitive enough to detect HCV in tissues from patients who were HCV negative in serum (three cases in our study), a situation that has been observed elsewhere (35).
This study is also, to our knowledge, the first one to provide both in situ morphological and molecular data concerning the presence of HCV in the sequential lesions, in other words, cirrhosis, DN, and HCC, of virus-associated carcinogenesis in human cirrhosis (5). We have been able to unambiguously demonstrate expression of HCV proteins in cells clearly exhibiting a tumor phenotype. Thus, our report definitely establishes persistence and expression of the viral genome, at least in a subset of tumor cells. This confirms previous studies based on immunohistochemistry (20, 21, 36) or molecular biology (37).
Among the different viral proteins detected by our IHC technique, NS3, core, and NS5A proteins have been shown in the literature to be able to modulate liver cell proliferation and viability in vitro. For instance, in vitro, stable expression in NIH 3T3 cells of the N-terminal part of NS3 can induce a transformed phenotype (38). The nonstructural NS5A protein could also play a role in the control of cell growth and viability because it might bind to the double-stranded RNA–induced protein kinase R, a protein involved in the control of apoptosis, and could also contribute to cell transformation, as shown in some in vitro studies (39).
The HCV core, in addition to its role in the packaging of the viral RNA, modulates cellular transduction pathways such as tumor necrosis factor alpha (TNFα) signaling (40, 41) and transactivates NF-κB and AP-1 elements, interacting with the c-jun N-terminal kinase and MAPKK (42). This protein seemed also to be capable of inducing (43) or, in contrast, inhibiting apoptosis induced by various stimuli (44, 45). Some in vivo studies with transgenic mice have also raised the idea of the role of HCV core expression in the induction of HCC (46). Although HCV core is mostly a cytoplasmic protein located on endoplasmic reticulum membranes and around lipid vesicles, C-terminally truncated core translocates to the nucleus, where it might exert distinct biological effects (47, 48, 49). During natural HCV infection, some reports have suggested that such truncated core might be identified in the tumor tissues of patients with HCC (50, 51). The occasional faint nuclear staining obtained in our study might correspond to such a protein, but a specific immunostaining is needed for confirmation.
A further important issue in our study is the demonstration of a significantly decreased expression of the viral antigens in the sequential lesions of hepatocarcinogenesis, whereas HCV RNA was still detectable at similar levels in HCC and cirrhosis. These results suggest that the viral genome does persist within tumor cells but that the expression of viral proteins is down-regulated when clonal tumor cell expansion occurs as early as the dysplastic nodule stage. Thus, the expression of the viral proteins might be involved at early steps of the liver cell transformation. The final result on cell phenotype of the opposing effects of viral proteins, that is, cell proliferation and apoptosis, is depending on their intracellular concentration, and on various extracellular signals. It is therefore plausible that low levels, barely detectable, of HCV proteins would allow liver cell proliferation, whereas higher levels would sensitize infected cells to apoptosis.
In conclusion, the in situ demonstration of viral proteins within dysplastic and tumoral cells, even at a low level, makes possible the direct role of HCV proteins in the early stages of carcinogenesis, alone or in combination with other factors, such as the regenerative process. Further studies will be necessary to identify which specific viral proteins are expressed in these different cell types and which specific in vivo effects they may have on cell proliferation and apoptosis.
References
Bannasch P . Pathogenesis of hepatocellular carcinoma: sequential cellular, molecular and metabolic changes. Prog Liver Dis 1996; 14: 161–197.
International Working Party. Terminology of nodular hepatocellular lesions. Hepatology 1995; 22: 983–993.
Di Bisceglie AM . Hepatitis C and hepatocellular carcinoma. Hepatology 1997; 26 (1 Suppl):34S–38S.
Colombo M, Fasani P, Romeo R . Hepatitis C virus and hepatocellular carcinoma. Res Virol 1997; 148: 127–134.
Le Bail B, Belleannée G, Bernard PH, Saric J, Balabaud C, Bioulac-Sage P . Adenomatous hyperplasia in cirrhotic livers: histological evaluation, cellular density, and proliferative activity of 35 macronodular lesions in the cirrhotic explants of 10 adult French patients. Hum Pathol 1995; 26: 897–906.
Kew MC . Hepatocellular carcinoma with and without cirrhosis. A comparison in southern African blacks. Gastroenterology 1989; 97: 136–139.
De Mitri M, Poussin K, Baccarini P, Pontisso P, D’Errico A, Simon N, et al. HCV-associated liver cancer without cirrhosis. Lancet 1995; 345: 413–415.
Bréchot C . The direct interplay between HCV NS5A protein and interferon transduction signal: from clinical to basic science. J Hepatol 1999; 30: 1152–1154.
Komminoth P, Adams V, Long AA, Roth J, Saremaslani P, Flury R, et al. Evaluation of methods for hepatitis C virus detection in archival liver biopsies. Comparison of histology, immunohistochemistry, in situ hybridization, reverse transcriptase polymerase chain reaction (RT-PCR) and in situ RT-PCR. Pathol Res Pract 1994; 190: 1017–1025.
Negro F, Hadengue A . Détection de l’ARN du virus de l’hépatite C par hybridation in situ: une analyse critique. Gastroenterol Clin Biol 1997; 21: 93–97.
Blight K, Rowland R, Hall P, Lesniewski RR, Trowbridge R, LaBrooy JT, et al. Immunohistochemical detection of the NS4 antigen of hepatitis C virus and its relation to histopathology. Am J Pathol 1993; 143: 1568–1573.
Chamlian A, Benkoël L, Sahel J, Cherid A, Brisse J, Ikoli JF, et al. Immunohistochemical detection of hepatitis C related c100–3 and core antigens in formalin-fixed liver tissue. Cell Mol Biol 1996; 42: 557–566.
Gonzales Peralta RP, Fang JWS, Davis GL, Gish R, Tsukiyama-Kohara K, Kohara M, et al. Optimization for detection of hepatitis C virus antigen in the liver. J Hepatol 1994; 20: 143–147.
Hiramatsu N, Hayashi N, Haruna Y, Kasahara A, Fusamoto H, Mori C, et al. Immunohistochemical detection of hepatitis C infected hepatocytes in chronic liver disease with monoclonal antibody to core, envelope and NS3 regions of the hepatitis C virus genome. Hepatology 1994; 20: 143–147.
Krawczynski K, Beach MJ, Bradley D, Kuo G, Di Bisceglie AM, Houghton M, et al. Hepatitis C virus antigen in hepatocytes: immunomorphologic detection and identification. Gastroenterology 1992; 103: 622–629.
Nouri-Aria KT, Sallie R, Mizokami M, Portmann BC, Williams R . Intrahepatic expression of hepatitis C virus antigens in chronic liver disease. J Pathol 1995; 175: 77–83.
Nuovo GJ, Lidonici K, MacConnell P, Lane B . Intracellular localization of polymerase chain reaction (PCR)-amplified hepatitis C cDNA. Am J Surg Pathol 1995; 175: 77–83.
Tang L, Tanaka Y, Enomoto N, Marumo F, Sato C . Detection of hepatitis C virus RNA in hepatocellular carcinoma by in situ hybridization. Cancer 1995; 76: 2211–2216.
Uchida T, Shikata T, Tanaka E, Kiyosawa K . Immunoperoxidase staining of hepatitis C virus in formalin-fixed, paraffin-embedded needle liver biopsies. Virchows Arch 1994; 424: 465–469.
Sansonno D, Cornacchiulo V, Racanelli V, Damacco F . In situ simultaneous detection of hepatitis C virus RNA and hepatitis C virus-related antigens in hepatocellular carcinoma. Cancer 1997; 80: 22–33.
Uchida T, Shikata T . Hepatitis C virus appears to replicate not only in hepatocytes but also in hepatocellular carcinoma cells as demonstrated by immunostaining. Pathol Int 1994; 44: 832–836.
Bédossa P, Poynard T, the METAVIR Cooperative Study Group. An algorithm for the grading of activity in chronic hepatitis C. Hepatology 1996; 24: 289–293.
Edmondson HA, Steiner PE . Primary carcinoma of the liver: a study of 100 cases among 48900 necropsies. Cancer 1954; 7: 462–503.
Ballardini G, Groff P, Giostra F, Francesconi R, Miniero R, Ghetti S, et al. Hepatocellular codistribution of c100, c33, c22 and NS5 hepatitis C virus antigens detected by using immunopurified polyclonal spontaneous human antibodies. Hepatology 1995; 21: 730–734.
Erali M, Ashwood ER, Hillyard DR . Performance characteristics of the COBAS AMPLICOR hepatitis C virus MONITOR test, version 2.0. Am J Clin Pathol 2000; 114: 180–187.
Garrigue I, Pellegrin I, Hoen B, Dumon B, Harzic M, Schrive MH, et al. Cell-associated HIV-1-DNA quantitation after highly active antiretroviral therapy-treated primary infection in patients with persistently undetectable plasma HIV-1 RNA. AIDS 2000; 14: 2851–2855.
Martin J, Navas S, Quiroga JA, Colucci G, Pardo M, Carreno V . Quantitation of hepatitis C virus in liver and peripheral blood mononuclear cells from patients with chronic hepatitis C virus infection. J Med Virol 1998; 54: 265–270.
Duvoux C, Pawlowsky JM, Bastie A, Cherqui D, Soussy CJ, Dhumeaux D . Low replication levels in end-stage hepatitis C-virus-related liver disease. J Hepatol 1999; 31: 593–597.
Takeda S, Shibata M, Morishima T, Harada A, Nakao A, Takagi H, et al. Hepatitis C virus infection in hepatocellular carcinoma. Cancer 1992; 70: 2255–2259.
Gerber MA . Relation of hepatitis C virus and hepatocellular carcinoma. J Hepatol 1993; 17: S108–S111.
Saito K, Sullivan D, Haruna Y, Theise ND, Thung SN, Gerber MA . Detection of hepatitis C virus RNA sequences in hepatocellular carcinoma and its precursors by microdissection polymerase chain reaction. Arch Pathol Lab Med 1997; 121: 400–403
Kobayashi S, Hayashi H, Itoh Y, Asano T, Isono K . Detection of minus-strand hepatitis C virus RNA in tumor tissues of hepatocellular carcinoma. Cancer 1994; 73: 48–52.
Tong CY, Hollingsworth RC, Williams H, Irving WL, Gilmore IT . Effect of genotypes on the quantification of hepatitis C virus (HCV) RNA in clinical samples using the Amplicor HCV Monitor Test and the Quantiplex HCV RNA 2.0 Assay (bDNA). J Med Virol 1998; 55: 191–196.
Lunel F, Cresta P, Vitour D, Payan C, Dumont B, Frangeul L, et al. Comparative evaluation of hepatitis C virus RNA quantitation by branched DNA, NASBA, and Monitor assays. Hepatology 1999; 29: 528–535.
Dries V, von Both I, Müller M, Gerken G, Schirmacher P, Odenthal M, et al. Detection of hepatitis C virus in paraffin-embedded liver biopsies of patients negative for viral RNA serum. Hepatology 1999; 29: 223–229.
Haruna Y, Hayashi N, Kamada T, Hytiroglou P, Thung SN, Gerber MA . Expression of hepatitis C virus in hepatocellular carcinoma. Cancer 1994; 73: 2253–2258.
Paterlini P, Driss F, Nalpas B, Pisi E, Franco D, Berthelot P, et al. Persistence of hepatitis B and hepatitis C viral genomes in primary liver cancers from HBsAg-negative patients: a study of a low-endemic area. Hepatology 1993; 17: 20–29.
Sakamuro D, Furukawa T, Takegami T . Hepatitis C virus nonstructural protein NS3 transforms NIH3T3. J Virol 1995; 69: 3893–3896.
Gale M, Kwieciszewski B, Dossett M, Nakao H, Katze MG . Antiapoptotic and oncogenic potentials of hepatitis C virus are linked to interferon resistance by viral repression of the PKR protein kinase. J Virol 1999; 73: 6506–6516.
Zhu N, Khoshnan A, Schneider R, Matsumoto M, Dennert G, Ware C, et al. Hepatitis C virus core protein binds to the cytoplasmic domain of tumor necrosis factor (TNF) receptor 1 and enhances TNF-induced apoptosis. J Virol 1998; 72: 3691–3697.
You LR, Chen CM, Wu Lee YH . Hepatitis C virus core pro-tein enhances NF-KB signal pathway triggering by lymphotoxin-B receptor ligand and tumor necrosis factor alpha. J Virol 1999; 73: 1672–1681.
Shrivastava A, Manna SK, Ray R, Aggarwal BB . Ectopic expression of hepatitis C virus core protein differentially regulates nuclear transcription factors. J Virol 1998; 72: 9722–9728.
Ruggieri A, Harada T, Matsuura Y, Miyamura T . Sensitization to Fas-mediated apoptosis by hepatitis C virus core protein. Virology 1997; 229: 68–76.
Ray RB, Mayer K, Steele R, Shrivastava A, Aggarwal BB, Ray R . Inhibition of tumor necrosis factor (TNF-alpha)-mediated apoptosis by hepatittis C virus core protein. J Biol Chem 1998; 273: 2256–2259.
Marusawa H, Hijikata M, Chiba T, Shimotohono K . Hepatitis C virus core protein inhibits Fas- and tumor necrosis factor alpha-mediated apoptosis via NF-KB activation. J Virol 1999; 72: 4713–4720.
Moriya K, Fujie H, Shintani Y, Yotsuyanagi H, Tsutsumi T, Ishibashi K, et al. The core protein of hepatitis C virus induces hepatocellular carcinoma in transgenic mice. Nat Med 1998; 4: 1065–1067.
Matsuura Y, Harada T, Makimura M, Sato M, Aizaki H, Suzuki T, et al. Characterization of HCV structural proteins expressed in various animal cells. Intervirology 1994; 37: 114–118.
Suzuki R, Matsuura Y, Suzuki T, Ando A, Chiba J, Harada S, et al. Nuclear localization of the truncated core protein with its hydrophobic C terminus delated. J Gen Virol 1995; 76: 53–61.
Barba G, Harper F, Harada T, Kohara M, Goulinet S, Matsuura Y, et al. Hepatitis C virus core protein shows a cytoplasmic localization and associates to cellular lipid storage droplets. Proc Natl Acad Sci USA 1997; 94: 1200–1205.
Rüster B, Zuieem S, Roth WK . Hepatitis C virus sequences encoding truncated core proteins detected in hepatocellular carcinoma. Biochem Biophys Res 1996; 219: 911–915.
Horie C, Iwanata H, Horie T, Shimizu I, Yoshimoto K, Yogita S, et al. Detection of different quasispecies of hepatitis C virus core region in cancerous and noncancerous lesions. Biochem Biophys Res 1996; 218: 674–681.
Acknowledgements
This work was supported by the SNFGE (Société Nationale Française de Gastro-Entérologie), the FRM (Fondation pour la Recherche Médicale), and the Ligue Contre le Cancer-Comité Dordogne.
Among the members of the Liver Transplantation Unit of the University Hospital of Bordeaux, the authors especially thank Professors J. Saric and J. Carles (surgery), Dr. P-H Bernard, Dr. V. de Lédinghen, and Professor P. Couzigou (medical follow-up). We also wish to thank Professor Christian Bréchot for the review of this work and his considerable assistance with the discussion of the results.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Rullier, A., Trimoulet, P., Urbaniak, R. et al. Immunohistochemical Detection of HCV in Cirrhosis, Dysplastic Nodules, and Hepatocellular Carcinomas with Parallel-Tissue Quantitative RT-PCR. Mod Pathol 14, 496–505 (2001). https://doi.org/10.1038/modpathol.3880338
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.3880338