Abstract
Recent studies predict that tumor aneuploidy plays a direct role in tumor instability. The relationship between interphase cytogenetics, histology, grade, and tumor site was analyzed in 20 primary gastric carcinomas. Using fluorescence in-situ hybridization, the numerical changes of centromeric sequences of chromosomes 1, 3, 10, and 17 were directly analyzed in gastric biopsies. Polysomic copy numbers of chromosomes 1 and 17 were discovered in 63percnt; (10 of 16) and 59percnt; (10 of 17), respectively, of informative cancer cases. Chromosome 3 and 10 signal number changes were found in only 6percnt; (1 of 16) and 13percnt; (1 of 8), respectively, of informative cancer cases. There was a positive correlation between the appearance of polysomic nuclear target sites of chromosomes 1 and 17 (correlation coefficient r = 0.72; p < 0.005). Copy number changes were not significantly related to histologic subtypes of either the Laurén or WHO classifications. However, incidence of cancers having dual polysomic signal number abnormalities for both chromosomes 1 and 17 was significantly correlated to tumor location at the cardia. The data suggests that (i) human gastric cancer appears in two genomic groups that can be reliably diagnosed by fluorescence in-situ hybridization on routine biopsy sections, (ii) numerical aberrations of chromosomes 1, 3, 10, and 17 are largely independent of histologic subtypes, and (iii) polysomic copy number abnormalities of chromosomes 1 and 17 correlate to intragastric tumor site and are highest in cardia cancers, suggesting high tumor instability at this particular location.
Similar content being viewed by others
Introduction
Gastric carcinoma is of major importance worldwide. Despite a declining incidence in many countries, statistical data on mortality rates still identify stomach cancer among the 10 leading killer cancers (Peddanna et al, 1995). Gastric carcinoma is classified histologically according to the WHO and Laurén schemes. Cytogenetic studies on stomach cancer have been able to define numerical and structural changes involving a broad spectrum of chromosomes, some of which seem to be non-random (Peddanna et al, 1995; Wright et al, 1992). Mutations at defined proto-oncogene loci were described with varying frequency, with or without significant correlations to tumor histology (Nanus et al, 1990; Ogasawara et al, 1994; Ranzani et al, 1993; Tamura et al, 1996). The deletion of a number of specific loci on several chromosomes has been detected by studies using loss of heterozygosity (Cho et al, 1996; Ezaki et al, 1996; Sano et al, 1991). Recent hypotheses predict that genetic instability of neoplasms is proportional to their degree of chromosome aneuploidy (Duesberg et al, 1998; Lengauer et al, 1999).
Non-isotopic fluorescence in situ hybridization (FISH) with labeled DNA probes has become an important tool for the analysis of numerical and structural chromosome aberrations in interphase nuclei of neoplasms (Lichter et al, 1991). Probes specific to the repetitive centromeric sequences on individual chromosomes have proved particularly useful in assessing numerical chromosome abnormalities in interphase cells of solid tumors (Fringes et al, 1997; Lichter et al, 1991). Most interphase cytogenetic studies on gastric cancer have been performed on separated tumor cell nuclei derived from either disintegrated tissues or from cultured gastric tumor cell lines, all of which may be biased by the selection of particularly malignant cell clones and may not be representative of the original in-situ human neoplasm (Lichter et al, 1991). Gastric biopsies present the advantages of small tissue size and short time of overall processing (including fixation) that can improve hybridization efficiency (Fringes et al, 1997). The present study was performed to assess the numerical changes of four different chromosomes in primary human gastric carcinoma, and to test for a possible relationship between genotypic abnormalities, tumor phenotype, and tumor site. We used FISH and specific DNA probes for centromeric repeat regions on chromosomes 1, 3, 10, and 17 on stomach biopsy samples of 20 patients with primary gastric carcinomas categorized according to both the Laurén and WHO classifications.
Results
Clinical
Gastric carcinoma were histologically classified according to the criteria of the WHO and Laurén. Table 1 shows histologic type and grade, localization, age, sex, and interphase cytogenetics for the four chromosomes tested. Intestinal carcinomas were preferentially located in the antral region of the stomach; diffuse-type tumors predominated in the cardia and cardia/corpus area. The proportionate number of diffuse-type carcinoma in this study was higher than in studies published by other authors because we preferentially harvested diffuse-type carcinoma cases from our files to compare chromosome changes within the different cancer subtypes.
Interphase Cytogenetics
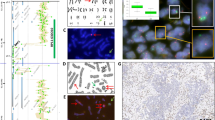
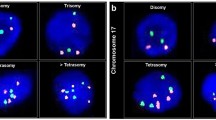
Digital photomicrographs generated in a laser scanning confocal microscope are shown in Figure 1. Control nuclei in normal gastric surface epithelia contained one to two signals per nuclear profile (Fig. 1a). Gastric neoplasms displayed a spectrum of signal distribution (Fig. 1b, c, e, and f). In some of the neoplastic cases, signal numbers counted on the tissue sections did not vary considerably from the control biopsies (Fig. 1d). The results of the quantitative microscopic analysis of fluorescent signal numbers are shown in Figure 2 and Table 2. The frequency distribution of the centromeric signals of each of the probes analyzed, estimated per nuclear profile and averaged from all cases that hybridized to the particular probe is depicted in Figure 2. Approximately 98percnt; of the nuclei in normal gastric surface epithelial cells contained one or two fluorescent signals per profile. Stomach cancer cells present in the biopsies of the same subject displayed a broad variety of signal numbers, ranging from one to six signals per nuclear profile, with a dominant nuclear fraction displaying three fluorescent signals. Figures 2a and 1a also show a fair amount of monosomic normal cells. This was expected because 5 to 7 μm-thick tissue sections were used throughout the study. Because the average diameter of a nucleus is 8 to 10 μm, some of the disomic nuclei would contain less than a complete nucleus and less than two signals. Additionally, the number of signals per trisomic neoplastic nuclei, likewise, might be higher than actually estimated on the nuclear profiles.
Photomicrographs of gastric neoplasms and control gastric mucosa after fluorescence in situ hybridization (FISH) with different biotinylated centromere repeat probes (green) and nuclear counterstaining with propidium iodide (red). Normal gastric surface epithelium (a) and diffuse-type gastric carcinoma rated as trisomic (b), both from Case 12, probed with pUC1.77 for chromosome 1 pericentromeric sequences. Diffuse-type gastric carcinoma (Case 13), rated as trisomic (c), and diffuse-type cancer (Case 11), scored as disomic (d), both probed for chromosome 17. Intestinal-type carcinoma (Case 8) (adenocarcinoma, poorly differentiated) hybridized to pUC 1.77 and scored as trisomic for chromosome 1 (e). Intestinal-type stomach carcinoma (Case 6) hybridized to the chromosome 17-specific probe p17H8, rated as trisomic (f). Scale bars: 100 μm in a, b, d, and f; 50 μm in c and e.
Frequency distribution of centromeric signals (mean ± sd) of all reactive gastric carcinoma cases analyzed on biopsy sections for chromosomes 1, 17, 3, and 10. a, Normal control epithelium, 98percnt; of the total nuclear fraction displays one to two signals/profile. b, Gastric cancers, rated as trisomic/tetrasomic with neoplastic nuclei, displaying a broad spectrum of signal distribution with a major nuclear fraction having three signals per nuclear profile. c, Gastric carcinoma cases, rated as disomic, displaying one to two signals/profile in their major nuclear fractions. Differences between trisomic fractions in gastric cancers from panels a, b, and c were significant (p < 0.005). The number over each bar is the chromosome number; n is the number of cases.
Two significantly different patterns of signal number distributions were registered, which was confirmed with all of the four DNA probes hybridized. Quantitative differences between the two patterns were most obvious when the fractions of neoplastic nuclei displaying three and four signals per nuclear profile were compared. In a number of tumors, the frequency distribution of nuclei consistently showed that 35percnt; to 40percnt; of cells contained three signals per nuclear profile, whereas, in many other cases, this nuclear fraction only reached an average of 6percnt; of cells (p < 0.0005). Differences were also significant when the percentages of tumor cells with four signals per nuclear profile were compared (p < 0.005). We defined the first group of tumors as polysomic, ie, trisomic and tetrasomic (Fig. 2b), and the second group as eusomic, ie, disomic (Fig. 2c), for the particular chromosomes investigated. If the actual fraction of nuclear profiles with three signals of each of the cases is displayed, the tumors that scored as trisomic cluster above the group of tumors that scored as disomic (Fig. 3). The threshold value was calculated at 21percnt; of trisomic nuclear profiles (using the formula given in “Materials and Methods”), and allows a clear separation between the two genomic gastric cancer groups. Signal number changes of chromosomes 1 and 17 were generally correlated with each other. There was a positive correlation between trisomic copy number changes of chromosomes 1 and 17 (r = 0.72; p < 0.005). See also Table 1.
Actual trisomic nuclear fraction (three signals/nuclear profile) for each of the gastric cancers and each of the chromosomes analyzed is shown. Neoplasms rated as trisomic cluster above the group of gastric cancers rated as disomic. The statistical threshold value calculated for trisomic tumors was 21percnt; of trisomic nuclei. N-trisomic, N-disomic, control epithelium of trisomic and disomic cases, respectively; CA-trisomic, CA-disomic, carcinoma rated as trisomic or disomic, respectively (p < 0.05).
Trisomic signal numbers for chromosomes 1 or 17 were detected in 63percnt; (10 of 16) and 59percnt; (10 of 17) of all neoplastic cases, respectively. Only 6percnt; (1 of 16) and 13percnt; (1 of 8) of the neoplasms contained trisomic signal numbers for chromosomes 3 and 10, respectively (Table 2). In the total incidence of gastric cancer cases that were trisomic for either or both chromosomes 1 and 17, two genotypic types of gastric carcinoma were registered: one group of 65percnt; (13 of 20) that contained trisomic copy numbers of chromosomes 1 and 17, and a second group of 35percnt; (7 of 20) that did not show polysomic copy number changes of chromosomes 1 and 17 (Table 1, Fig. 2).
Statistical analysis of the mean signal number per nuclear profile showed that, for both diploid and triploid gastric cancer cases, there were significant differences between controls and tumors but not between chromosomes (2-way ANOVA with chromosome type and group). With respect to the mean signal numbers per profile, variance analysis also showed that there were significant differences between the diploid and triploid cancer cases and that these differences resided principally in the tumor cells and not in the controls.
Tumor Histology, Grade, and Interphase Cytogenetics
When gastric carcinomas were subdivided according to Laurén, a slightly higher number of diffuse-type carcinoma cases (67percnt;, 6 of 9) than intestinal-type cancers (57percnt;, 4 of 7) were scored as trisomic/polysomic for chromosome 1 (Table 1). When the neoplasms were classified according to the criteria of the WHO, the highest fraction of tumors with trisomic signal numbers for chromosome 1 (2 of 7 reactive cases, 29percnt;; 2 of 3 of all poorly differentiated tumors, 67percnt;), and chromosome 17 (3 of 8 reactive cases, 38percnt;; 3 of 4 of all poorly differentiated cases, 75percnt;) were found to belong to the poorly differentiated type of gastric adenocarcinoma, and to the neoplastic cases diagnosed as signet ring-cell carcinoma (4 of 5 reactive cases, 80percnt;, for chromosome 1; and 3 of 5 reactive cases, 60percnt;, for chromosome 17) (Fig. 4). Using chi-square and Fisher’s exact probability tests on chromosomes 1 and 17 individually did not reveal significant differences in histologic subtypes of either classification.
Relationship between histologic type of gastric neoplasms, classified according to the WHO, and interphase cytogenetic results for chromosomes 1 and 17. Well (open bar), moderately (striped bar), and poorly differentiated (black bar) adenocarcinoma; signet ring cells carcinomas rich (shaded bar) and poor (horizontal bar) in the amount of signet ring cells.
Tumor-Site and Interphase Cytogenetics
Because the copy number changes of chromosome 1 and 17 were correlated, we assessed the number of reactive cases in which chromosomes 1 and 17 (i) were both scored as trisomic (C1/C17-dual trisomic), (ii) either one was scored as trisomic (C1/C17-single trisomic), (iii) were scored as disomic/trisomic (C1/C17-unbalanced trisomic), (iv) were scored as disomic for both (C1/C17-dual disomic), or (v) either one was scored as disomic (C1/C17-single disomic). The number of cases with dual, single, and unbalanced trisomic or disomic signal numbers for chromosomes 1 and 17 were summed to give a total C1/C17-signal number index, and these sums were expressed as a percentage of the total number of reactive cases. Table 3 shows that the highest number (85.7percnt;) of gastric cancers with total trisomic signal numbers for both chromosome 1 and 17 was found in tumors localized to the cardia. In contrast, in all other parts of the stomach, dual trisomic cases were much less frequent (53.9percnt;) than at the cardia. The difference was significant at the 0.05 level using Fisher’s exact probability test. Table 3 shows that cases with dual trisomic signal numbers for chromosomes 1 and 17 were also frequent at higher age ranges and in male subjects. Differences in C1/C17-dual trisomic signal numbers between intestinal and diffuse-type cancers were low. Dual trisomic copy numbers for chromosome 1 and 17 simultaneously were also more frequently observed in adenocarcinomas that were rated as poorly differentiated and in signet ring cell carcinomas. Applying the Fisher Yates test, the results showed a tendency toward the diffuse-type cancer located at the cardia being more frequently dual-trisomic for chromosomes 1 and 17 than diffuse-type cancers located in the antral region (p < 0.3).
Discussion
Numerical aberrations in 20 primary human gastric carcinomas have been characterized in routine stomach biopsies using FISH with DNA probes specific for the (peri)centromeric sequences on chromosomes 1, 3, 10, and 17. Application of this quantitative interphase cytogenetic approach to tissue sections identified two genotypic types of gastric carcinoma, a major group with polysomic signal numbers for chromosomes 1 and 17 and a smaller group of stomach cancer cases with disomic copy numbers for chromosomes 1 and 17. Chromosomes 3 and 10 remained numerically unchanged in nearly all of the neoplastic cases. There was a strong positive correlation between the polysomic signal number changes of chromosomes 1 and 17. Chromosome aberrations did not correlate with histologic subtypes, but significantly correlated with tumor location within the gastric wall. They also correlated to some extent with the WHO tumor grade.
The appearance of two genomic types of gastric cancers has also been reported by other investigators, applying FISH (Gomyo et al, 1995; Van Decken et al, 1990), banding methods (Peddanna et al, 1995; Wright et al, 1992), or RFLP analyses (Ezaki et al, 1996; Ranzani et al, 1993; Sano et al, 1991). The incidence of gastric cancers with aneusomic signal number changes for chromosomes 1 and 17 observed in our study equaled those in studies using RFLP analysis that showed a loss of heterozygosity on both the long arm of chromosome 1 and on 17p in 67percnt; of gastric adenocarcinomas (Sano et al, 1991). Chromosome 17 harbors the human proto-oncogene c-erbB-2 and the p53 locus, which have been shown to be deleted in 68percnt; and 40percnt; to 77percnt;, respectively, of advanced stomach carcinomas (Gomyo et al, 1996; Kobayashi et al, 1996; Sano et al, 1991; Shiao et al, 1994). Recently a 17q12–21 amplicon was discovered in the intestinal type of gastric carcinoma (Kokkola et al, 1997). Numerical changes of chromosome 3 and 10 were infrequent in this study. However, a tumor suppressor gene on chromosome arm 3p and an amplified genomic domain on chromosome 10 were observed in gastric adenocarcinomas of the poorly differentiated type (Mor et al, 1993; Schneider et al, 1995).
One of the goals of the present analysis was to test whether a correlation could be detected between tumor histologic phenotype and genotype, as represented by the two different classifications of gastric cancers and the numerical changes represented by copy number changes of four targeted chromosomes. There was no significant difference in the frequency of signal number distribution or in the incidence of cases with polysomic centromeric copy numbers for defined chromosomes between the two types of the Laurén classification or between the diverse subtypes of the WHO classification. Similarly, Sano et al (1991) did not find significant differences between histologic subtypes of stomach cancers when analyzing loss of heterozygosity in several chromosomes, including chromosomes 1 and 17. Studies on chromosome 17 allelic loss at the p53 locus and p53 expression also did not find significant associations with the histologic subtypes of the Laurén classification (Gabbert et al, 1995; Kim et al, 1995). Thus, we conclude that numerical abnormalities of chromosomes 1 and 17 in human gastric carcinoma are independent of histologic phenotype.
The incidence of gastric cancer cases with dual-polysomic copy number changes for chromosomes 1 and 17 correlated significantly with a cardial location of the neoplasms. Cancers located in the upper third of the stomach have a worse outcome compared with stomach neoplasms originally localized in the antral part of the stomach (Ikeguchi et al, 1995). This has been mainly attributed to the facts that cardia and fundus carcinoma are more often diffuse-type cancers and/or need a more aggressive surgical intervention of total gastrectomy (Ogasawara et al, 1994; Wright et al, 1992). Our study demonstrates that cardia cancers may represent a subgroup of malignant stomach neoplasms that contain abnormalities of copy numbers for particular chromosomes at a higher case incidence than cancers located in the lower part of the gastric interior. Additional implications that intragastric location is a major factor in biological behavior and outcome of stomach cancers comes from our observation that diffuse-type cancers located at the cardia were even more frequently polysomic than diffuse-type cancers located in the antrum. Recent studies shed a new light on chromosome aneuploidy, suggesting that genetic instability of cancer cells is proportional to their degree of aneuploidy, and that aneuploidy may be sufficient to explain genetic instability (Duesberg et al, 1998; Lengauer et al, 1999). We propose that the cardia may be susceptible to dietary carcinogens in a different manner from the antral part, and thus harbors cancers with higher frequencies of numerical chromosome abnormalities, higher genetic instability, and more aggressive behavior. Additionally, our study supports the hypothesis that genomic instability of gastric neoplasms is reflected by chromosome aneuploidy in the neoplastic cells and, within the gastric pouch, may also be strongly dependent on the intragastric location of the stomach cancers.
Materials and Methods
Clinical Material
Formalin-fixed, paraffin-embedded routine gastric biopsies from a total of 20 stomach cancer patients were examined. Histologic classification was established according to the criteria of Laurén and the WHO. The diagnoses made on the biopsy specimen were compared with those made on the respective surgically resectioned stomach neoplasms. Carcinomas that contained both intestinal and diffuse-type cancer components were classified according to the dominant tumor cell type seen within the biopsy specimen analyzed by the FISH method.
FISH Procedure
Multiple 6 μm-thick sections were cut, attached to slides precoated with aminopropyltriethoxysilane (Aldrich Chemical Company, Milwaukee, Wisconsin), and dewaxed in xylene. Pretreatment procedures according to Brigati et al (1983) were applied. Briefly, slides were immersed in 0.2N HCl and extracted in 0.5percnt; Triton X-100, then digested at room temperature with pronase solution (0.25 to 0.35 mg/ml) (Calbiochem, La Jolla, California) and treated with RNAase A (100 μg/ml) and RNase T1 (5 μg/ml) for 30 minutes at room temperature and for an additional 30 minutes at 37° C. Tissue sections were postfixed in buffered 4percnt; paraformaldehyde, dehydrated in graded ethanols and air dried. A set of four DNA plasmid probes that specifically decorate the centromeric/pericentromeric heterochromatin of their respective chromosomes was used. Those for chromosomes 1 (pUC1.77) and 17 (P17H8) were described by Cooke and Hindley (1979) and Waye and Willard (1986), respectively. A probe specific for chromosome 10 (pa10RP8) was described by Devilee et al (1988). The DNA probe for chromosome 3 (P3–5), has been mapped to the centromere region in the laboratory of Dr. D. C. Ward. All probes were labeled by nick translation with biotin-11-dUTP. Probe specificity was tested by hybridizing the plasmids to metaphase spreads of stimulated peripheral human lymphocytes. Labeled probe DNA (20 to 40 ng) was combined with salmon sperm DNA, ethanol precipitated, and redissolved in 10 μl of hybridization mixture. Probe and specimen DNA were denatured together at 86° C to 94° C for 6.5 minutes and hybridized overnight at 37° C. Biotinylated hybridization sites were detected by fluorescein isothiocyanate-avidin (Vector Laboratories, Burlingame, California), applying one cycle of signal amplification. Intermediate and final washing solutions contained 2x SSC and 0.1percnt; Tween-20 at 42° C. Nuclei were counterstained with propidium iodide and embedded in antifade mounting solution.
Microscopy and Signal Acquisition
Hybridization signals in interphase nuclei were visualized using a Nikon Optiphot microscope equipped with FITC epifluorescence optics, interfaced to a laser scanning confocal imaging system (Bio-Rad Lasersharp MCR-500 scanner, Bio-Rad, Hercules, California) using a 60× 1.4 Plan Apochromatic oil-immersion objective. Evaluation of signal numbers was performed by using the conventional epifluorescence mode. Digital images of specimens were generated in the scanner mode. For the scoring of signal numbers in interphase-nuclei of control and neoplastic gastric epithelial cells, these conditions were met: (i) a total of one to four biopsies containing tumor tissue was available per case, control and neoplastic biopsies of each subject were processed together; (ii) the number of signals was assessed in an average of 50 to 70 nuclear profiles taken at random from both the tumor containing areas and from the non-tumorous gastric surface epithelial cells of each case, most non-tumorous biopsies were from an additional biopsy specimen, distant from the neoplastic region; (iii) hybridization signals were estimated per individual nuclear profile.
Statistical Analysis
Student’s t test for unequal samples and the chi-squared test were used for statistical analysis. For significance of the differences regarding tumor site, histology, sex, and age-range, Fisher’s exact probability test was applied. Variance analysis was performed using the average signal numbers in control and neoplastic tissues of all cases and chromosomes. The threshold value for trisomy (three signals/nuclear profile) was calculated for each of the chromosome probes analyzed by using the formula in

where d3 are disomic cancer nuclei having three signals; d3c are control nuclei with three signals (false trisomy); t3 are trisomic nuclei of gastric cancers with 30percnt; or more trisomic nuclei; sd is the standard deviation. This value was used to test the reliability of the separation between trisomic and disomic gastric cancer cases.
References
Brigati DJ, Myerson D, Leary JJ, Spalholz B, Travis SZ, Fong CKY, Hsiung GD, and Ward DC (1983). Detection of viral genomes in cultured cells and paraffin-embedded tissue sections using biotin-labeled hybridization probes. Virology 126: 32–50.
Cho JH, Noguchi M, Ochiai A, and Hirohashi S (1996). Loss of heterozygosity of multiple tumor suppressor genes in human gastric cancers by polymerase chain reaction. Lab Invest 74: 835–841.
Cooke HJ and Hindley J (1979). Cloning of human satellite III DNA. Different components are on different chromosomes. Nucleic Acids Res 6: 3177–3197.
Devilee P, Kievits T, Waye JS, Pearson PL, and Willard HF (1988). Chromosome-specific alpha satellite DNA: Isolation and mapping of a polymorphic alphoid repeat from human chromosome 10. Genomics 3: 1–7.
Duesberg P, Rausch C, Rasnik D, and Hehlmann R (1998). Genetic instability of cancer cells is proportional to their degree of aneuploidy. Proc Natl Acad Sci U. S. A. 95: 13692–13697.
Ezaki T, Yanagisawa A, Ohta K, Aiso S, Watanabe M, Hibi T, Kato Y, Nakajima T, Ariyama T, Inazawa J, Nakamura Y, and Horii A (1996). Deletion mapping on chromosome 1p in well differentiated gastric cancer. Br J Cancer 73: 424–428.
Fringes B, Gates J, and Gomez F (1997). Technical aspects of non-radioactive in-situ hybridization (interphase-FISH) to normal and neoplastic human gastric biopsy specimen. Pathol Res Pract 137/2: 99 (Suppl).
Gabbert HE, Müller W, Schneiders A, Meier S, and Hommel G (1995). The relationship of p53 expression to the prognosis of 418 patients with gastric carcinoma. Cancer 76: 720–726.
Gomyo Y, Andachi H, Nagao K, Ikeguchi M, and Ito H (1995). Interphase cytogenetics of gastric carcinoma: Fluorescence in situ hybridization (FISH) applied to cells obtained from formalin-fixed, paraffin-embedded tissues. Pathol Int 45: 227–232.
Gomyo Y, Osaki M, Kaibara N, and Ito H (1996). Numerical aberration and point mutation of p53 gene in human gastric intestinal metaplasia and well-differentiated adenocarcinoma: Analysis by fluorescent in situ hybridization (FISH) and PCR-SSCP. Int J Cancer 29: 594–599.
Ikeguchi M, Ohfuji S, Oka A, Tsujitani S, Maeta M, and Kaibara N (1995). Aneuploidy of tumor cells in cases of gastric cancer with esophageal invasion: Another indicator of poor prognosis. J Surg Oncol 58: 83–90.
Kim CJ, Kim WH, Kim CW, Lee JB, Lee CK, and Kim YL (1995). Detection of 17p loss in gastric carcinoma using polymerase chain reaction. Lab Invest 72: 232–236.
Kobayashi M, Kawashima A, Mai M, and Ooi A (1996). Analysis of chromosome 17p13 (p53 locus) alterations in gastric carcinoma cells by dual-color fluorescence in situ hybridization. Am J Pathol 149: 1575–1584.
Kokkola A, Monni O, Puolakkainen P, Larramendy ML, Victorzon M, Nordling S, Haapiainen R, Kivilaakso E, and Knuutila S (1997). 17q12-21 amplicon, a novel recurrent genetic change in intestinal type of gastric carcinoma: A comparative genomic hybridization study. Genes Chromosomes Cancer 20: 38–43.
Lengauer C, Kinzler KW, and Vogelstein B (1999). Genetic instability in colorectal cancers. Nature 386: 623–627.
Lichter P, Boyle AL, Cremer T, and Ward DC (1991). Analysis of genes and chromosomes by nonisotopic in situ hybridization. GATA 8: 24–35.
Mor O, Ranzani GN, Rotman G, Gutman M, Manor A, Amadori D, Gouldsworth J, Hollstein M, and Schwab M (1993). DNA amplification in human gastric carcinomas. Cancer Genet Cytogenet 62: 111–114.
Nanus DM, Kelsen DP, Mentle IR, Altorki N, and Albino AP (1990). Infrequent point mutations of ras oncogenes in gastric cancers. Gastroenterology 98: 955–960.
Ogasawara S, Maesawa C, Tamura G, and Satodate R (1994). Lack of mutations of the adenomatosis polyposis coli gene in oesophageal and gastric carcinomas. Virchows Arch 424: 607–611.
Peddanna N, Holt S, and Verma RS (1995). Genetics of gastric cancer. Anticancer Res 15: 2055–2064.
Ranzani GN, Renault B, Pellegata NS, Fattorini P, Magni E, Bacci F, and Amadori D (1993). Loss of heterozygosity and K-ras gene mutations in gastric cancer. Human Genet 92: 244–249.
Sano T, Tsujino T, Yoshida K, Nakayama H, Haruma K, Ito H, Nakamura Y, Kajiyama G, and Tahara E (1991). Frequent loss of heterozygosity on chromosomes 1q, 5q, and 17p in human gastric carcinomas. Cancer Res 51: 2926–2931.
Schneider BG, Pulitzer DR, Brown RD, Prihoda TJ, Bostwick DG, Saldivar V, Rodriguez-Martinez HA, Gutierrez-Diaz ME, and O’Connell P (1995). Allelic imbalance in gastric cancer: An affected site on chromosome arm 3p. Genes Chromosomes Cancer 13: 263–271.
Shiao YP, Rugge M, Correa P, Lehmann HP, and Scheer WD (1994). p53 alteration in gastric precancerous lesions. Am J Pathol 144: 511–517.
Tamura G, Ogasawara S, Nishizuka S, Sakata K, Maesawa C, Suzuki Y, Terashim M, Saito K, and Satodate R (1996). Two distinct regions of deletion on the long arm of chromosome 5 in differentiated adenocarcinomas of the stomach. Cancer Res 56: 612–615.
Van Decken H, Pizzolo JG, Kelsen DP, and Melamed MR (1990). Targeted cytogenetic analysis of gastric tumors by in situ hybridization with a set of chromosome-specific DNA probes. Cancer 66: 491–497.
Waye JS and Willard HF (1986). Structure, organization, and sequence of alpha satellite DNA from human chromosome 17: Evidence for evolution by unequal crossing-over and an ancestral pentamer repeat shared with the human x chromosome. Molec Cell Biol 6: 3156–3165.
Wright PA, Quirke P, Attanoos R, and Williams GT (1992). Molecular pathology of gastric carcinoma. Hum Pathol 23: 848–859.
Acknowledgements
The expertise of F. Gomez, C. Simoes, and A. Cirillo in technical assistance and retrieval of the patient data is gratefully acknowledged by the authors.
Author information
Authors and Affiliations
Corresponding author
Additional information
This work was supported by grants from the Deutsche Forschungsgemeinschaft, Bonn (Fr 563/2-) and from the Boehringer-Ingelheim Fonds, Stuttgart, Germany to BF and by Grant GM-40115 to DCW.
Rights and permissions
About this article
Cite this article
Fringes, B., Mayhew, T., Reith, A. et al. Numerical Aberrations of Chromosomes 1 and 17 Correlate with Tumor Site in Human Gastric Carcinoma of the Diffuse and Intestinal Types. Fluorescence In Situ Hybridization Analysis on Gastric Biopsies. Lab Invest 80, 1501–1508 (2000). https://doi.org/10.1038/labinvest.3780159
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1038/labinvest.3780159