Abstract
Lung cancer has a considerable impact on morbidity and mortality throughout the world. Despite extensive effort, no lung cancer-specific cytogenetic changes, such as lineage-specific translocations or inversions, have been described to date. In this study we used multiplex fluorescence in situ hybridization (M-FISH), comparative genomic hybridization, and multicolor bar coding to analyze eight cell lines derived from non-small cell lung cancers. M-FISH did not identify any balanced translocations, which are the dominating feature in leukemias and lymphomas. Instead, M-FISH unraveled an enormous number of numerical and structural aberrations, with each tumor having its own “private” pattern of chromosomal changes. In contrast, comparative genomic hybridization demonstrated similarities between tumors, because each cell line shared some chromosomal segments that were commonly gained or lost. One of these involved chromosome 12. Chromosome 12 specific bar code probe sets were constructed and used to demonstrate that breaks on chromosome 12 occur preferentially within specific bands. With the progressive use of higher resolution approaches, more information can be gained about the chromosomal alterations in cancer.
Similar content being viewed by others
Introduction
In industrialized countries lung cancer is the most common cause of cancer death among both women and men, with an increasing incidence during the past few decades (Pisani et al, 1993). Lung cancer is subdivided into small cell lung cancers (SCLCs) and non-small cell lung cancers (NSCLCs). NSCLCs comprise three major groups depending on the cell type: squamous cell carcinoma (30% of all lung cancer), adenocarcinoma (40%), and large cell carcinoma (15%).
Because of the high impact of lung cancer on morbidity and mortality, considerable efforts have focused on understanding the underlying genetic change related to the pathophysiology of lung cancer. Karyotype analysis has provided a fruitful approach to characterizing tumor cells. The accumulating data from hematological malignant diseases and bone and soft tissue tumors support the idea that each cancer type may be characterized by specific genetic alterations (Barr, 1998). However, such tumor-specific alterations have generally not been found in the common epithelial malignancies. Despite technical advances in cell culture, the rate of successful karyotypic analysis of lung carcinomas has remained low because of karyotype complexity and chromosome morphology. In NSCLCs, one of the most common aberrations is the deletion of 3p, as first documented by Zech et al (1985). Overall, the emerging picture indicates that, in NSCLCs, deletions of chromosome arms 3p, 9p, and 17p, trisomy 7, and isochromosomes i(5)(p10) and i(8)(q10) are recurrent themes within the plethora of genetic alterations (Johansson et al, 1995; Testa et al, 1997). Comparative genomic hybridization (CGH) and allelotyping studies have confirmed many of these karyotypic imbalances and revealed, in addition, several other recurrent abnormalities, such as DNA copy number decreases on chromosome arms 1p, 3p, 4q, 5q, 6q, 8p, 9p, 10q, 13q, 14q, 18q, and 21q, and DNA gains on chromosome arms 1q, 5p, 7p, 7q, 8q, 11q13, 12p, 16p, 17q, 19q, and 20q (Petersen et al, 1997a, 1997b, 1998). However, no cytogenetic alterations specific for NSCLCs have yet been detected.
In this study, multiplex fluorescence in situ hybridization (M-FISH) (Speicher et al, 1996a) was used for color-karyotyping of metaphase spreads from four new NSCLC cell lines and four established NSCLC cell lines. In M-FISH, at least five fluorophores are needed to distinguish the 24 human chromosomes, and each of the five fluorophores is imaged separately by using a series of optical filters (Lichter, 1997; Speicher et al, 1996a, 1996b). The five images are merged into a composite in which each chromosome is assigned a distinct color based on its unique spectral signature (Bolzer et al, 1999; Eils et al, 1998; Speicher et al, 1996a, 1996b; Uhrig et al, 1999). The M-FISH results were combined with CGH data. Because chromosome 12 was often involved in structural aberrations, a chromosome 12-specific bar code was used to determine whether breaks on this chromosome occurred in a completely random fashion. The molecular cytogenetic approaches presented here use progressively higher resolutions and, therefore, permit a more precise description of the spectrum of chromosome alterations in NSCLCs.
Results
CGH Analysis
A summary of genetic imbalances observed in the cell lines is shown in Figure 1. Vertical lines to the left of each chromosome indicate loss and lines to the right indicate gain of chromosomal material. Gains and losses were observed on all chromosomes. Frequent losses were observed on chromosome arms 4q, 8p, 9p, 9q, 13q, and 15q, whereas frequent gains were observed on chromosome arms 8q, 12p, 12q, 17q, and 20q. The CGH results from our newly established cell lines were compared with the primary tumors from which they were derived, and each showed an almost identical CGH profile (data not shown).
Summary of gains and losses detected by comparative genomic hybridization (CGH) in primary cell cultures (lines 1–4: D51, D54, D97, D117) and cell lines from the American Type Culture Collection (ATCC) and German Collection of Microorganisms and Cell Cultures (DSMZ), respectively (lines 5–12: A427, BEN, Colo-699, DV-90). Bars on the right side of a chromosome indicate gains, whereas bars on the left side indicate losses. Frequent losses were observed at chromosome regions 4q, 8p, 9p, 9q, 13q, and 15q, and frequent gains on chromosome arms 8q, 12p, 12q, 17q, and 20q.
Multiplex FISH
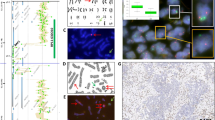
Karyotypes were generated based on the spectral signature of chromosomes. With the exception of DV-90, all cell lines revealed a large number of numerical and structural abnormalities. A detailed description of the composite karyotypes, in the long nomenclature of the International Standing Committee on Human Cytogenetic Nomenclature (ISCN) (Mitelman, 1995), is shown in Table 1. Only abnormalities observed in at least two metaphase spreads were included in the karyotype designation. Figure 2 shows a representative karyotype for cell lines Colo-699 (a), A427 (b), D54 (c), and D117 (d).
M-FISH did not reveal a significant difference in chromosomal complexity between cell lines with a low passage number and the cell lines with a high passage number. For example, cell lines D54 and D117 (Fig. 2, c and d) were analyzed at passages 3 and 19, respectively. In both cases the changes were already as complex as in the commercial cell lines (Fig. 2, a and b). To further unravel the underlying organization of cytogenetic abnormalities in NSCLCs, several questions were addressed:
How Often Are Individual Chromosomes Involved in Structural Abnormalities?
The frequency of interchromosomal abnormalities was evaluated by counting chromosomes participating in a translocation or insertion. For example, if a translocation t(2;7) was consistently observed in a particular tumor, participation in a structural abnormality was counted for both chromosome 2 and chromosome 7. Using M-FISH, only large intrachromosomal changes are detectable if chromosomes are homogeneously stained with the color combination of only one chromosome, and yet the size and/or 4′, 6-diamidino-2-phenylindole (DAPI) banding indicates structural abnormalities, such as deletions and duplications. These intrachromosomal aberrations were included in our summary shown in Figure 3. Chromosomes could be subdivided into three groups with chromosomes 1, 2, 3, 11, 12, and 22 as the group of chromosomes with the most often observed intra- or interchromosomal changes (>14 times each). Chromosomes 1, 5, 6, 7, 8, 9, 16, 17, 18, 19, and 20 participated less often (8–14 times), and chromosomes 4, 10, 13, 14, 15, 21, X, and Y, the least often (7 times or fewer each).
Plot of number of intra- (light bars) and interchromosomal (dark bars) changes per chromosome. Chromosomes most frequently involved in structural rearrangements were chromosomes 1, 2, 3, 11, 12, and 22 (>14 times each). Chromosomes 5, 6, 7, 8, 9, 16, 17, 18, 19, and 20 participated less often (8–14 times), whereas chromosomes 4, 10, 13, 14, 15, 21, X, and Y were rearranged at an even lower frequency (7 times or fewer).
Do Certain Chromosomes or Parts of Chromosomes Tend to Fuse Particularly Frequently Either as Translocations or Insertions?
The occurrence of fusions of different chromosomes or parts of chromosomes, either in the form of translocations or insertions, was analyzed in Figure 4. A fusion among two chromosomes is indicated in the respective field. For example, a translocation t(3;17) appears in column 17, row 3. The number in each field indicates how often such an interchromosomal exchange was observed. Only fusion chromosomes observed in at least two metaphase spreads were included. Fusions among parts of chromosomes 11 and 12 were observed most frequently (n = 7). Other frequently occurring fusions were, in descending order, between chromosomes 19 and 22 (n = 5); 1 and 15, 3 and 11, 3 and 12, 3 and 17, and 12 and 22 (n = 4 each). Note that this analysis does not indicate the breakpoints. Thus, the derivative chromosomes with chromosome 11 and 12 material may not have the same breakpoints.
Interchromosomal rearrangement scoring: summary of all fusions between different chromosomes. A fusion among two chromosomes is indicated in the respective field. For example, a translocation t(3;7) appears in column 7, row 3. The number in each field shows how often such an interchromosomal exchange occurred.
Can NSCLCs Be Classified According to the Percentage of Inter- and Intrachromosomal Changes?
The percentage of chromosomes participating in structural abnormalities for each tumor was analyzed (Fig. 5). In the majority of tumors, about 20% to 35% of chromosomes showed structural abnormalities (A427, Colo-699, D51, D54, and D117). However, in two tumors (BEN and D97), more than half of all chromosomes were involved. The only exception, with one unbalanced translocation, was DV-90. Most tumors (A427, BEN, Colo-699, D51, D97, and D117) were dominated by interchromosomal changes; however, in D54, intrachromosomal changes represented the majority.
Graph showing the percentage of chromosomes participating in structural abnormalities (interchromosomal, light; intrachromosomal, dark) for each tumor. About half of all chromosomes were involved in these abnormalities in tumors BEN and D97. For the other tumors, changes are in the range of 20% to 35%, with the exception of DV-90, which showed an unbalanced translocation as the only detectable structural alteration. In most tumors, interchromosomal changes represent the majority of observed changes; only D54 exceeded 50% intrachromosomal changes.
Chromosome 12-Specific Bar Code
To assess whether breakpoints along a chromosome occur at preferential locations, two chromosome 12-specific bar codes (Table 2a) were applied to cell lines A427, Colo-699, D51, D54, D97, and D117. The M-FISH results were used as a guide to locate specific marker-chromosomes in the metaphase spreads. Figure 6 shows examples of applications of the chromosome-12 specific bar code to Colo-699 (a) and A427 (b). The chromosome in the first column in Figure 6a is a normal chromosome 12 displaying the expected number of seven yeast artificial chromosome (YAC) signals. This figure shows that YAC clones are already so close to each other that the simultaneous hybridization of a larger number of YACs would result in overlapping signals. The der(12)t(12;22) (Fig. 2a) resulted in a missing signal of the p-terminal YAC 924, (Fig. 6a, column 2). Surprisingly the bar code revealed that the large chromosome 12 inserts in the der(17) (Fig. 2a) were caused by a duplication of chromosome region 12p11.2-p12 (Fig. 6a, columns 3 and 4).
Examples demonstrating the potential of the chromosome 12-specific bar code (Table 2a) in identifying chromosomal subregions involved in derivative chromosomes. The figure shows the 4′, 6-diamidino-2-phenylindole (DAPI)-stained chromosomes with chromosome 12 material to the left and the respective bar code fluorescence signals with the probe names to the right. (a) Set 2 of the chromosome 12 bar code hybridized to cell line Colo-699. The first column shows a normal chromosome 12 with all seven expected signals visible. On the chromosome in the second column, the signal for yeast artificial chromosome (YAC) clone 924 hours12 (12p12) is missing. This chromosome corresponds to the der(12)t(12;22) shown in Figure 2a. An amplification of chromosome region 12p11.2-p12 as indicated by the duplication of the signal for YAC 952a6 is shown in columns three and four. These chromosomes correspond to the two der(17) chromosomes in Figure 2a. (b) Set 2 of the chromosome 12 bar code hybridized to cell line A427. The chromosome in the first column corresponds to the der(3;13) in Figure 2b. The small chromosome 12 material at the distal tip was derived from chromosome band 12q24.3. The chromosome in the second column corresponds to the der(11) (Figure 2b) and contains chromosome 12 region 12q13-q15. The third column shows a der(3)(12?::3p11∼3q28::12?) with 12p12-q15 material at the proximal and 12p12 material at the distal end. The chromosome in the fourth column corresponds to the der(12)(12pter∼q12::11?::12?), with 12p12 in the proximal and 12q24.3 in the distal region.
Table 2b summarizes the frequency of observed breaks between adjacent YAC clones, and the results are also illustrated in Figure 7. Using the combined information of the two alternating bar code sets, two breakpoint cluster regions were identified in regions 12p11.2-p12.3 and 12q15-q21.3 (Fig. 7).
Summary of results obtained with the two sets of chromosome 12-specific YAC clones (Table 2). YACs and their map position are indicated on the left side. Bars on the right side indicate the space between adjacent YAC clones for each bar code set (black, Set 1; gray, Set 2). The thickness of the bars corresponds to the frequency of breaks between the two YACs. The combined information of the two bar code sets allowed us to narrow down the breakpoints. For example, with Set 2, breaks between YACs 952a6 and 754a1 were observed nine times, but no breaks were seen with Set 1 between YACs 933e11 and 928a12. This suggests that breaks preferentially occurred distal to YAC 933e11. Altogether, two major breakpoint cluster regions were identified: regions 12p11.2-p12.3 and 12q15–12q21.3.
Discussion
The large number of gains and losses detected by CGH (Fig. 1) reflects the enormous genomic instability encountered in this tumor entity. Similar CGH patterns for NSCLCs were previously obtained from primary NSCLCs (Petersen et al, 1997a, 1997b). The CGH results, showing specific gains and losses for some chromosomal regions, suggest that within the plethora of numerical changes some underlying specific patterns exist.
The genomic instability in NSCLCs was also reflected in the M-FISH results. With the exception of cell line DV-90, the structural and numerical changes observed were very complex (Fig. 2 and Table 1), and no common pattern was evident from the M-FISH data per se. Cell lines with short passage numbers (Fig. 2, c and d) demonstrated already a complexity similar to that of the commercially available cell lines with much higher passage numbers (Fig. 2, a and b).
Balanced translocations were not observed in any cell line. Instead, the data suggest that the development of NSCLCs is more dependent on an intricate combination of deletions and amplifications of multiple chromosomal segments. As a result, each cell line had its own, “private” pattern of numerous structural abnormalities. The only evidence for some nonrandomness was suggested by the preferential involvement of some chromosomes in structural inter- and intrachromosomal rearrangements (Fig. 3) and the clustering of breakpoints as shown for chromosome 12 (Figs. 6 and 7). Based on the facts that some chromosomes have a special preference to break, that the breakpoints occur preferentially in certain regions, and that there is some evidence that particular fusions of double-strand breaks occur more frequently, one can speculate that this could result in the formation of fusion proteins in epithelial cancers. However, the significance of such putative fusion proteins for tumor initiation and progression and their real frequency is currently completely elusive. The data suggest that tumors with preferential interchromosomal rearrangements (A427, BEN, Colo-699, D51, D97, and D117) occur with a higher frequency as tumors with a rather intrachromosomal instability (D54, as shown in Figure 5).
The application of well-defined chromosomal bar codes is an important addition to M-FISH. The delineation of a chromosomal subregion participating in a breakage is often not possible based on the M-FISH results and a comparison of the banding pattern alone. For example, in Figure 6a, the wide DAPI-light band was caused by a duplication of chromosome region 12p11.2-p12. This could not have been concluded by any other method. In the past the extent of complex chromosomal rearrangements was assessed by banding techniques alone. While M-FISH results indicate that the true extent was in fact underestimated, the additional application of a bar code improves resolution and increases the degree of discernible rearrangements even further. This is documented in the fact that the chromosome 12 breakpoint cluster regions were not identified before in NSCLC (Mitelman et al, 1997) (see also database about breakpoints in epithelial cancers: http://www.ncbi.nlm.nih.gov/CCAP/NG/mitelmanSum.html). We anticipate that, for a wide range of epithelial cancers, the precise location of many breakpoints is not known yet. Thus, assuming that the high resolution mapping of breakpoints may be the key in the identification of lineage specificity in epithelial tumors, a cascade of experiments, as described here with the progressive use of higher resolution approaches, should gain new information about the chromosomal alterations in cancer.
Materials and Methods
Cell Lines
Eight cell lines were analyzed. Four (D51, D54, D97, and D117) were primary cultures established at the Institute of Pathology at the Charité (Berlin, Germany), one (A427) was purchased from the American Type Culture Collection (ATCC, Rockville, Maryland), and three (BEN, Colo-699, and DV-90) were purchased from the German Collection of Microorganisms and Cell Cultures (DSMZ, Braunschweig, Germany). Five tumors were adenocarcinomas (Colo-699, D51, D54, D117, and DV-90), and one was a large cell carcinoma (D97). The others showed phenotypical features of NSCLC in cell culture without distinct histotypic differentiation.
CGH
CGH was done as described previously (Petersen et al, 1997a, 1997b). Details are available at the Web site http://amba.charite.de/cgh/.
Metaphase Preparation
Metaphase spreads were prepared from the primary cell cultures at passages 56 (D51), 3 (D54), 39 (D97), and 19 (D117). The remaining cell lines have been in culture for an extended period and passage number data are unavailable. Preparation of metaphase spreads was according to standard protocols with the exception of the hypotonic treatment, which was done using 0.8% sodium citrate for approximately 20 minutes.
M-FISH
M-FISH was done as described previously (Eils et al, 1998). In brief, the complex M-FISH probe mixture was generated by pooling the painting probes according to our combinatorial labeling scheme. For example, all painting probes with fluorescein isothiocyanate (FITC) as part of the identifier tag were combined in a “FITC-DNA-pool,” and the same was done for Cy3, Cy5, biotin, and digoxigenin. Exact probe concentrations were described by Eils et al (1998). The five fluorochrome DNA pools were labeled by degenerate oligonucleotide primed PCR (Telenius et al, 1992), mixed, and ethanol precipitated overnight. The precipitate was resuspended in a hybridization mix, consisting of 50% formamide, 15% dextran sulfate, and 2 × SSC. The probe mixture and slides were denatured and hybridized for one or two nights at 37° C. After post-hybridization washes and blocking with 3% BSA, biotin and digoxigenin pools were detected using avidin-Cy3.5 (1:300; Amersham Pharmacia Biotech, Uppsala, Sweden) and anti-digoxigenin-Cy7 (1:200; Amersham Pharmacia Biotech, Freiburg, Germany), respectively. FITC-, Cy3-, and Cy5-labeled probes were imaged directly with appropriate filter sets. The slides were counterstained with DAPI and mounted in p-phenylenediamine dihydrochloride antifade solution.
Chromosome 12-Specific Bar Code
YAC clones for a chromosome 12-specific bar code were selected from the Centre d'Etude du Polymorphisme Humain library (CEPH, Paris, France) based on previously published data (Bray-Ward et al, 1996). The relative distance of different YACs to each other was tested by hybridizing two or three YACs in different colors simultaneously. The YACs and their band assignment are summarized in Table 2. To avoid overlapping signals that may hamper unequivocal interpretation in tumor metaphase spreads with condensed chromosomes, two different chromosome 12 bar codes were generated. Each bar code set consisted of seven YACs (Table 2a), with complementary map positions to cover most regions of chromosome 12 and to achieve a maximum resolution.
Epifluorescence Microscopy
Images were captured using the motorized epifluorescence microscope Leica DMRXA-RF8 (Leica Microsystems, Wetzlar, Germany) equipped with an eight-position filter wheel and a new generation of filter sets (Chroma Technology Corporation, Brattleboro, Vermont). Specifications for these filters were published previously (Eils et al, 1998). For image capturing, a Sensys CCD camera (Photometrics, Tucson, Arizona) with Kodak KAF 1400 chip (Kodak, Rochester, New York) was used. Both the camera and microscope were controlled by the Leica QFISH software (Leica Microsystems Imaging Solutions Ltd., Cambridge, United Kingdom).
Image Analysis
M-FISH image analysis was done applying an adaptive region-oriented approach for spectral classification of fluorescence signals (Eils et al, 1998). The classification process is integrated into the Leica MCK image analysis package (Leica Microsystems Imaging Solutions Ltd.). Chromosome 12-specific bar codes were generated by pseudocoloring and overlaying gray images using the Leica QFISH software (Leica Microsystems Imaging Solutions Ltd.).
Change history
01 August 2000
In the July issue the legend for the cover figure was incorrect. The correct legend for the July cover figure appears below. We regret the error. Cover: Representative karyotypes of NSCLC cell lines (a) Colo-699 and (b) A427. Numbers adjacent to a derivative chromosome indicate the origin of the translocated or inserted material.
References
Barr FG (1998). Translocations, cancer and the puzzle of specificity. Nature Genet 19:121–124.
Bolzer A, Craig JM, Cremer T, and Speicher MR (1999). A complete set of repeat-depleted, PCR-amplifiable, human chromosome-specific painting probes. Cytogenet Cell Genet 84:233–240.
Bray-Ward P, Menninger J, Lieman J, Desai T, Mokady N, Banks A, and Ward DC (1996). Integration of the cytogenetic, genetic, and physical maps of the human genome by FISH mapping of CEPH YAC clones. Genomics 32:1–14.
Eils R, Uhrig S, Saracoglu K, Sätzler K, Bolzer A, Chassery JM, Ganser M, and Speicher MR (1998). An optimized, fully automated system for fast and accurate identification of chromosomal rearrangements by multiplex-FISH (M-FISH). Cytogenet Cell Genet 82:160–171.
Johansson M, Jin Y, Mandahl N, Hambraeus G, Johansson L, Mitelman F, and Heim S (1995). Cytogenetic analysis of short-term cultured squamous cell carcinomas of the lung. Cancer Genet Cytogenet 81:46–55.
Lichter P (1997). Multicolor FISHing: What's the catch. Trends Genet 13:475–479.
Mitelman F (ed) (1995). ISCN 1995: An international system for human cytogenetic nomenclature. Basel, Switzerland: S. Karger.
Mitelman F, Mertens F, and Johansson B (1997). A breakpoint map of recurrent chromosomal rearrangements in human neoplasia. Nat Genet 15:417–474.
Petersen I, Langreck H, Wolf G, Schwendel A, Psille R, Vogt P, Reichel MB, Ried T, and Dietel M (1997a). Small cell lung cancer is characterized by a high incidence of deletions on chromosomes 3p, 4q, 5q, 10q, 13q and 17p. Br J Cancer 75:79–86.
Petersen I, Bujard M, Petersen S, Wolf G, Goeze A, Schwendel A, Langreck H, Gellert K, Reichel M, Just K, du Manoir S, Cremer T, Dietel M, and Ried T (1997b). Patterns of chromosomal imbalances in adenocarcinoma and squamous cell carcinoma of the lung. Cancer Res 57:2331–2335.
Petersen S, Rudolf J, Bockmühl U, Gellert K, Wolf G, Dietel M, and Petersen I (1998). Distinct regions of allelic imbalances on chromosome 10q22–q26 in squamous cell carcinoma of the lung. Oncogene 17:449–454.
Pisani P, Parkin DM, and Ferlay J (1993). Esimates of the worldwide mortality from eighteen major cancers in 1985. Implications for prevention and projections of future burden. Int J Cancer 55:891–903.
Speicher MR, Ballard SG, and Ward DC (1996a). Karyotyping human chromosomes by combinatorial multi-fluor FISH. Nat Genet 12:368–375.
Speicher MR, Ballard SG, and Ward DC (1996b). Computer image analysis of combinatorial multi-fluor FISH. Bioimaging 4:52–64.
Telenius H, Pelmear AH, Tunnacliffe A, Carter NP, Behmel A, Ferguson-Smith MA, Nordenskjöld M, Pfragner R, and Ponder BAJ (1992). Cytogenetic analysis by chromosome painting using DOP-PCR amplified flow-sorted chromosomes. Genes Chromosomes Cancer 4:257–263.
Testa JR, Liu Z, Feder M, Bell DW, Balsara B, Cheng JQ, Taguchi T (1997). Advances in the analysis of chromosome alterations in human lung carcinomas. Cancer Genet Cytogenet 95:20–32.
Uhrig S, Schuffenhauer S, Fauth C, Wirtz A, Daumer-Haas C, Apacik C, Cohen M, Müller-Navia J, Cremer T, Murken J, and Speicher MR (1999). Multiplex-FISH (M-FISH) for pre- and postnatal diagnostic applications. Am J Hum Genet 65:448–462.
Zech L, Bergh J, and Nilsson K (1985). Karyotypic characterization of established cell lines and short-term cultures of human lung cancers. Cancer Genet Cytogenet 15:335–347.
Acknowledgements
We are grateful to Prof. M. Ferguson-Smith, Cambridge, United Kingdom, for the generous gift of chromosome-specific painting probes. We are indebted to Drs. Tricia Bray-Ward and David Ward for propagating the YAC clones to us.
Author information
Authors and Affiliations
Corresponding author
Additional information
This work was supported by the Alfried Krupp von Bohlen und Halbach Stiftung, the Deutsche Krebshilfe (10–1392-Pa1) and the Deutsche Forschungsgemeinschaft (Sp 460/3–1 and Pe 602/1–2).
Rights and permissions
About this article
Cite this article
Speicher, M., Petersen, S., Uhrig, S. et al. Analysis of Chromosomal Alterations in Non-Small Cell Lung Cancer by Multiplex-FISH, Comparative Genomic Hybridization, and Multicolor Bar Coding. Lab Invest 80, 1031–1041 (2000). https://doi.org/10.1038/labinvest.3780108
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1038/labinvest.3780108
This article is cited by
-
Hierarchical discovery of large-scale and focal copy number alterations in low-coverage cancer genomes
BMC Bioinformatics (2020)
-
Morphologische und molekulare Pathologie des Lungenkarzinoms
Der Pathologe (2010)
-
Chromosomal analysis of non-small-cell lung cancer by multicolour fluorescent in situ hybridisation
British Journal of Cancer (2004)