Abstract
The use of five histochemical stains (cresyl violet, thionin, hematoxylin & eosin, silver stain, and acridine orange) was evaluated in combination with an expression profiling paradigm that included regional and single cell analyses within the hippocampus of post-mortem human brains and adult mice. Adjacent serial sections of human and mouse hippocampus were labeled by histochemistry or neurofilament immunocytochemistry. These tissue sections were used as starting material for regional and single cell microdissection followed by a newly developed RNA amplification procedure (terminal continuation (TC) RNA amplification) and subsequent hybridization to custom-designed cDNA arrays. Results indicated equivalent levels of global hybridization signal intensity and relative expression levels for individual genes for hippocampi stained by cresyl violet, thionin, and hematoxylin & eosin, and neurofilament immunocytochemistry. Moreover, no significant differences existed between the Nissl stains and neurofilament immunocytochemistry for individual CA1 neurons obtained via laser capture microdissection. In contrast, a marked decrement was observed in adjacent hippocampal sections stained for silver stain and acridine orange, both at the level of the regional dissection and at the CA1 neuron population level. Observations made on the cDNA array platform were validated by real-time qPCR using primers directed against β-actin and glyceraldehyde-3 phosphate dehydrogenase. Thus, this report demonstrated the utility of using specific Nissl stains, but not stains that bind RNA species directly, in both human and mouse brain tissues at the regional and cellular level for state-of-the-art molecular fingerprinting studies.
Similar content being viewed by others
Main
The study of optimally prepared (eg, fixed, sectioned, and stained) tissues is a principal method for histological and histopathological investigation. Several histochemical staining techniques have been developed empirically on their ability to increase contrast of specific regional, cellular, or subcellular tissue constituents to enable the visualization of distinct cellular structures and cell types, as most populations of cells are relatively translucent and difficult to discern in unstained preparations. Although histological stains are typically not specific to an individual cell type or protein, much information can be gleaned by utilizing classical histological preparations in conjunction with contemporary protein (eg, immunocytochemistry) and molecular biological methodologies. However, the information gathered through morphological observation (in particular, clinicopathological diagnosis within the central nervous system) and molecular biological methods have proven difficult to compare and correlate in the same brain tissue section. The problem arises mainly from the perception that methods requisite for morphological investigation and molecular- and/or cellular-based studies are mutually exclusive. Thus, morphological observation via histochemical staining and molecular procedures, such as RNA amplification followed by cDNA array analysis or quantitative real-time polymerase chain reaction (qPCR), could not be performed on the same tissue section or identified cells simultaneously. Therefore, neuroscientists relied on in situ hybridization (ISH) histochemistry procedures to examine gene expression in tissue sections. ISH is an extremely useful technique, especially when applied to well-processed brain sections, allowing for the localization and estimation of abundance of specific mRNA transcripts within single cells.1 Similar to protein assessments using immunocytochemistry, quantitation of reaction product density in the case of nonradioactive probes such as digoxigenin, or densitometry of silver grains from radiolabeled probes, can be applied. Owing to several technical considerations including limited sensitivity and relative inability to assess multiple mRNAs simultaneously, ISH is best suited for regional assessments of gene expression patterns, or as a companion technique to PCR- or linear amplification-based analyses.2, 3
Problems with the combination of cellular visualization procedures and molecular-based techniques has been partially obviated by combining immunocytochemical procedures for labeling specific proteins in tissue sections with microdissection and subsequent microarray analysis.4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14 A question remains as to whether histochemical stains can be used in conjunction with RNA amplification and cDNA array analysis. Several groups have used hematoxylin & eosin (H&E) staining on peripheral tissues in cancer-related paradigms in combination with laser capture microdissection (LCM) and PCR-based detection as well as cDNA microarray platforms using RNA amplification methods.15, 16, 17, 18 The efficacy of several nuclear dyes in PCR-based detection of genomic DNA and transcripts on LCM-acquired cells has also been performed in peripheral tissues,19, 20, 21 with varying degrees of success depending upon methods of tissue accession and preparation. In brain, few reports using histological stains exist, although successful cDNA microarray analysis has been performed on dorsal root ganglia neurons stained with cresyl violet (CV).22 Moreover, qPCR has been performed on hippocampal neurons stained with hematoxylin, nuclear fast red, or immunostained with an antibody directed against NeuN.23 Notably, RNA yields did not differ significantly in samples stained with either of the two dyes or NeuN.23 Despite these encouraging observations, an important question looms as to the utility of specific histochemical stains in brain when applied in conjunction with expression profiling technologies including RNA amplification and cDNA microarray analysis. The use of histochemical stains is particularly indicated when employing single cell and/or population cell analyses whereby neighboring cell types can be discriminated morphologically and microaspirated unequivocally in a histochemical-based preparation, but not readily by chemical phenotype analysis.
In the present report, five histochemical procedures have been employed along with immunocytochemistry using an antibody directed against neurofilaments (NF) in adjacent serial sections of human and mouse hippocampus to evaluate the effects of histochemical stains (Figure 1). Specifically, histochemistry was employed in conjunction with RNA amplification and gene expression profiling, as assessed on custom-designed cDNA array platforms containing approximately 220 cDNAs relevant to neuroscience.4, 5, 6, 24 Three stains that visualize Nissl substance (basophilia), CV, thionin (Th), and H&E were used. Although these dyes stain Nissl substance by binding to stacks of rough endoplasmic reticulum in the cytoplasm, we hypothesized that accession to mRNAs (at least the polyadenylated tail and 5′ regions) would not be impeded significantly by dye binding. A modified Gallyas silver stain (SiS) was selected because of its historical perspective in neuroscience and its usefulness in identifying neuronal processes as well as neuropathological hallmarks of neurodegenerative disorders. In addition, SiS has been used to detect minute quantities of RNA25, 26 as well as proteins subjected to gel electrophoresis. A histofluorescent dye, acridine orange (AO), was also employed. AO is a fluorescent dye that intercalates selectively into nucleic acids27, 28 and has been used to detect RNA and DNA in brain tissues.29 AO has been used by this group as a diagnostic measure in tissue sections to visualize intact RNA species prior to engaging in laborious RNA amplification and cDNA array procedures in individual human autopsy cases.30, 31 We hypothesize that AO will affect negatively gene expression levels as compared to Nissl stains because of the intercalation of the dye directly into RNA species. Similarly, the employment of SiS is predicted to cause a reduction in signal due to the ability of silver conjugates to bind nucleic acids directly. Importantly, all five histochemical stains are evaluated in relation to an immunocytochemical preparation on fixed hippocampal tissue sections that has demonstrated previously excellent hybridization signal intensity.4, 5, 12, 24
Materials and methods
Subjects
Tissue accession and accrual procedures were in accordance with IRB and IACUC standards at the Nathan Kline Institute and NYU School of Medicine. Hippocampi were accessed from human autopsy cases (Table 1; n=6 males; mean age 72.3±10.7 (s.d.) years; mean post-mortem interval (PMI) 13.0±5.2 h) and from adult male C57BL/6 mice (n=8; 20–30 g). The human case materials were obtained from neuropathologically confirmed normal controls without history of neurological or major psychiatric illness as described previously.4, 5, 6, 11, 12 The brains used in this study were accrued from normal control subjects accessed through the established collection at the Center for Neurodegenerative Disease Research, University of Pennsylvania School of Medicine.
Histological Processing
For the human studies, regional hippocampal dissections and individual CA1 hippocampal neurons were isolated from 6 μm-thick fixed (fixed in 70% ethanol plus 150 mM sodium chloride overnight at 22°C) paraffin-embedded sections of hippocampus. Mice were given an overdose of ketamine and xylazine and perfused transcardially with ice-cold 70% ethanol plus 150 mM sodium chloride. Brains were removed from the skull and post fixed in the same solution for 2 h. Tissue blocks containing the dorsal hippocampus were paraffin embedded. Tissue sections (6 μm-thick) were cut in the coronal plane on a rotary microtome for histochemistry and immunocytochemistry. All the tissue samples were harvested using the same methods and procedures. Each human and mouse brain was confirmed to have abundant cytoplasmic RNAs by AO histofluorescence as described previously.7, 30, 31
Histochemical and Immunocytochemical Preparation
The procedures for the histochemical stains employed in this study were described briefly, as they are fairly conventional and well-established methods. The pertinent dye substrates and reagents were purchased from Sigma (St Louis, MO, USA) unless noted otherwise. Briefly, all human and mouse tissue sections were deparaffinized in xylene, and rehydrated through a descending ethanol series followed by immersion in 18.2 mega Ohm RNase-free water and stored in an RNase-free 0.1 M Tris (pH 7.6) solution prior to initiating staining protocols. CV was performed at a final concentration of 0.02% in acetate buffer as described previously.32, 33 Th was used at a final concentration of 0.1% in acetate buffer (pH 3.0) for 20 min followed by differentiation in 70% ethanol containing glacial acetic acid.34, 35 H&E was performed using undiluted hematoxylin (Gill's #2; Electron Microscopy Sciences, Fort Washington, PA, USA) followed by immersion in a working solution of 0.3% eosin.5, 36 For AO staining, tissue sections were placed in a 0.2 M dibasic sodium phosphate/0.1 M citric acid (SC; pH 4.0) solution for 5 min prior to staining with AO (10 μg/ml) in SC buffer for 15 min. The sections were rinsed three times in SC buffer and immersed briefly in 50% ethanol in 0.1 M Tris.28, 30, 31 A Gallyas SiS modified for tissue sections adhered to glass slides was employed as described previously to detect fibers.34, 37, 38 Finally, the immunocytochemical protocol consisted of blocking tissue sections in a 0.1 M Tris (pH 7.6) solution containing 2% donor goat serum (DGS) and 0.01% Triton X-100 for 1 h and then incubating with a monoclonal antibody directed against nonphosphorylated NF proteins (RMdO2039) to delineate CA1 pyramidal neurons and hippocampal lamination patterns in a 0.1 M Tris/2% DGS solution overnight at 4°C in a humidified chamber. Sections were processed with the ABC kit (Vector Labs, Burlingame, CA, USA) and developed with 0.05% diaminobenzidine, 0.03% hydrogen peroxide, and 0.01 M imidazole in Tris buffer for 10 min as described previously.4, 5, 6, 12 Stained tissue sections were not coverslipped and were immersed in RNase-free 0.1 M Tris until regional dissection and/or microaspiration and subsequent TC RNA amplification was performed. Histochemical control studies in conjunction with TC RNA amplification have been reported previously.4, 40 In addition, adjacent unstained (U) tissue sections were deparaffinized and used to evaluate the effects of histochemical and immunocytochemical labeling procedures on cDNA array hybridization signal intensity. Reaction products were assessed microscopically, and images were archived digitally using a photomicroscope (E800, Nikon, Japan) interfaced with a CCD camera (CoolSnap, Roper, Trenton, NJ, USA) and imaging software (MetaMorph, Universal Imaging Corporation, Downingtown, PA, USA).
Regional hippocampal dissections were performed on stained tissue sections using a scalpel blade under microscopic visualization as described previously.24, 41 Three to five dissections were performed per human (n=6) and mouse (n=8) brain. Hippocampal dissections included the Ammon's horn subfields and the dentate gyrus. Entorhinal cortex and the subicular complex were excluded from the dissection. Individual CA1 pyramidal neurons were microaspirated using LCM instrumentation (PixCell IIe, Arcturus, Mountain View, CA, USA) (Figure 2). A total of 23–25 cells were captured per reaction.4, 24 A total of 3–5 reactions (containing 23–25 LCM-captured CA1 neurons each) were performed per human (n=6) and mouse (n=8) brain.
Microaspiration by LCM of a representative CA1 pyramidal neuron from a fixed tissue section of the mouse hippocampus stained with CV. The tissue section was dehydrated in xylenes and not coverslipped (note the grainy appearance of the tissue). (a) CA1 pyramidal neuron to be microaspirated is identified (arrow). Following one laser pulse of the near-infrared laser (b), the thermoplastic transfer film on the cap was adhered to the CA1 neuron (arrow). The cap was removed from the LCM, leaving behind a space (arrow) where the CA1 neuron was microaspirated (c). The captured cell can be visualized (d) by placing the cap on a clean slide for contrast. Scale bar a–d: 25 μm.
TC RNA Amplification
The amplification of RNA from regional hippocampal dissections and individual CA1 pyramidal neurons was performed using a new terminal continuation (TC) RNA amplification methodology developed in this laboratory.4, 14, 40 TC RNA amplification provides high fidelity, linear amplification with significant representation of long transcripts under a variety of staining conditions using a myriad of cell types as evidenced by bioanalysis and cDNA array analysis.4, 40 TC RNA amplification entailed synthesizing first strand cDNA complementary to the RNA template, generating second strand cDNA complementary to the first strand cDNA, and finally in vitro transcription using the double-stranded cDNA as template.14, 40 Dissected hippocampal regions and individual hippocampal neurons were incubated in 250 μl of proteinase K solution (Ambion, Austin, TX, USA, 50 μg/ml) for 12 h at 37°C prior to extraction in Trizol reagent (Invitrogen, Carlsbad, CA, USA). RNAs were reverse transcribed in the presence of the poly d(T) primer (10 ng/μl) and TC primer (10 ng/μl) in 1X first strand buffer (Invitrogen), 1 mM dNTPs, 5 mM DTT, 20 U of RNase inhibitor, and 5 U reverse transcriptase (Superscript III; Invitrogen). The synthesized single-stranded cDNAs were converted into double-stranded cDNAs by adding into the reverse transcription reaction the following: 10 mM Tris (pH 8.3), 50 mM KCl, 1.5 mM MgCl2, and 0.5 U Rnase H (Invitrogen) in a total volume of 99 μl. Samples were placed in a thermal cycler and second strand synthesis proceeded as follows: Rnase H digestion step 37°C, 10 min; denaturation step 95°C, 3 min, annealing step 50°C, 3 min; elongation step 75°C, 30 min Taq polymerase 5 U (1 μl) (PE Biosystems, Foster City, CA, USA) was added to the reaction at the initiation of the denaturation step (ie, hot start).40 The reaction was terminated with 5 M ammonium acetate. The samples were extracted in phenol:chloroform:isoamyl alcohol (25:24:1) and ethanol precipitated with 5 μg of linear acrylamide (Ambion) as a carrier. The solution was centrifuged at 14 000 rpm and the pellet washed once with 95% ethanol and air-dried. The cDNAs were resuspended in 20 μl of RNase-free H2O and drop dialyzed on 0.025 μm filter membranes (Millipore, Billerica, MA, USA) against 50 ml of 18.2 mega Ohm RNase-free H2O for 2 h. The sample was collected off the dialysis membrane and hybridization probes were synthesized by in vitro transcription using 33P incorporation in 40 mM Tris (pH 7.5), 7 mM MgCl2, 10 mM NaCl, 2 mM spermidine, 5 mM of DTT, 0.5 mM of ATP, GTP, and CTP, 10 μM of cold UTP, 20 U of RNase inhibitor, T7 RNA polymerase (1000 U, Epicentre, Madison, WI, USA), and 40 μCi of 33P-UTP (Amersham Biosciences, Piscataway, NJ, USA). The reaction was performed at 37°C for 4 h. Radiolabeled TC RNA probes were hybridized to custom-designed cDNA arrays without further purification.
Custom-designed cDNA Array Platforms and Data Analysis
Array platforms consisted of 1 μg of linearized cDNA purified from plasmid preparations adhered to high-density nitrocellulose (Hybond XL, Amersham Biosciences). Each cDNA and/or expressed sequence-tagged cDNA (EST) was verified by sequence analysis and restriction digestion. cDNA clones and ESTs from mouse, rat, and human were employed. Approximately 220 cDNAs/ESTs were utilized on the current array platform. Classes of transcripts included: cytoskeletal elements (n=12); glutamate receptors, transporters, and interacting proteins (n=35); neurotrophins and neurotrophin receptors (n=13); Alzheimer's disease-related genes (n=36); synaptic-related markers (n=18); immediate-early genes (n=9); cell death markers (n=28); glial-associated markers (n=9); GABA synthesis, receptors, and transporters (n=20); potassium channels (n=8); protein phosphatases/kinases (n=21); and others (n=3). The majority of genes are represented by one transcript on the array platform. However, several genes have representation at 3′ and 5′ regions, including the high-affinity neurotrophin receptors (trkA, trkB, and trkC) and the neurofilament subunits (NF-L, MF-M, and NF-H) to assess relative expression levels from separate regions of the gene and evaluate potential RNA degradation.5 For example, ESTs that encode the tyrosine kinase domain (TK) and extracellular domain (ECD) were employed for trkA, trkB, and trkC.5 Arrays were prehybridized (2 h) and hybridized (12 h) in a solution consisting of 6X SSPE, 5X Denhardt's solution, 50% formamide, 0.1% sodium dodecyl sulfate (SDS), and denatured salmon sperm DNA (200 μg/ml) at 42°C in a rotisserie oven.4, 5, 6 Following hybridization, arrays were washed sequentially in 2X SSC/0.1% SDS, 1X SSC/0.1% SDS and 0.5X SSC/0.1% SDS for 20 min each at 42°C. Arrays were placed in a phosphor screen for 24 h and developed on a phosphor imager (Amersham Biosciences). Hybridization signal intensity was quantified by subtracting background using empty vector (pBs). Expression of TC amplified RNA bound to each linearized cDNA (approximately 220 cDNAs/ESTs) was expressed as a ratio of the total hybridization signal intensity of the array (a global normalization approach). Global normalization effectively minimizes variation due to differences in the specific activity of the synthesized TC probe as well as the absolute quantity of probe present.4, 5, 6 Data analyzed in this manner does not allow the absolute quantitation of mRNA levels. However, an expression profile of relative changes in mRNA levels was generated. Relative changes in total hybridization signal intensity and in individual mRNAs were analyzed by one-way analysis of variance (ANOVA) with post hoc analysis (Newmann–Keuls test; P<0.01 is the determinant for statistical significance) for individual comparisons.4, 5, 7, 11, 12, 14, 41 Expression levels were analyzed and clustered using a bioinformatics and graphics software package (GeneLinker Gold, Predictive Patterns Inc., Kingston, ON, USA).
Real-time qPCR
qPCR was performed using mouse hippocampal dissections stained with CV, Th, H&E, NF, SiS, and AO as starting material. PCR primers were designed for two genes, the cytoskeletal element β-actin (ACTB) and the housekeeping gene glyceraldehyde-3 phosphate dehydrogenase (GAPDH). The primer sequences are as follows: β-actin, amplicon length 101 bp, forward primer: ctt cct tct tgg gta tgg aat cc; reverse primer: cgt aaa gac ctc tat gcc aac aca. GAPDH, amplicon length 95 bp, forward primer: tga gca aga gag gcc cta tcc; reverse primer: cca taa taa cag gag ggg cct a. The PCR primers were designed to span intron–exon junctions to avoid potential genomic DNA contamination. Samples were run on a real-time PCR cycler (7900HT, ABI, Foster City, CA, USA) using SYBR green dye chemistry as a reporter (SYBR Green PCR Master Mix, ABI) as per the manufacturer's instructions. The qPCR assay was performed in triplicate on hippocampal regional dissections obtained from 3–4 mouse brains per staining condition on a 96-well optical plate (ABI). Standard curves and cycle threshold (Ct) were measured using standards obtained from total mouse brain RNA. Similar to the cDNA array analysis, relative changes in PCR product synthesis were analyzed by one-way ANOVA with post hoc analysis (Newmann–Keuls test) for individual comparisons. Amplicon specificity was evaluated by subcloning the β-actin and GAPDH amplicon products (Zero Blunt, Invitrogen) and performing sequence analysis.
Results
A description of the experimental design is illustrated in Figure 1. Adjacent serial tissue sections from human post-mortem hippocampus and mouse hippocampus were stained with CV, Th, H&E, SiS, AO, and immunocytochemically labeled with anti-NF for microscopic evaluation and subsequent downstream genetic analyses using TC RNA amplification coupled with custom-designed cDNA microarrays. The hippocampal formation was readily discernible from temporal neocortex and subcortical structures in all five of the histochemical preparations. Examination of NF-immunoreactive tissue sections throughout the human and mouse temporal lobe revealed a distinct laminar pattern of immunoreactivity confined to the somatodendritic region of neurons throughout Ammon's horn subfields of the hippocampal formation. No apparent differences in the staining intensity or reaction product distribution were observed between the normal human cases used in this study. AO staining did not reveal any significant differences in the presence of nucleic acid content between the six human cases, similar to previous observations by our group using some of these same case materials.5, 12 In addition, robust AO staining of nucleic acids was observed in the mouse brains, consistent with previous observations in fixed tissue sections.4
Regional analysis of the post-mortem human hippocampus revealed robust hybridization signal intensity for hippocampi stained with Nissl stains (ie, CV, Th, and H&E), and NF immunocytochemistry, whereas very low, but consistent, levels of hybridization signal intensity were detected for adjacent serial sections stained with SiS and AO (Figure 3a). Quantitative analysis revealed no significant differences in relative hybridization signal intensity between the individual Nissl stains and NF immunocytochemistry. All four displayed highly significant hybridization signal intensity levels as compared to SiS and AO (P<0.0001). The staining procedures did not impair signal recovery, as no differences were observed between the Nissl stains, NF immunocytochemistry, and adjacent unstained sections (Figure 4). However, there was much greater variability in the unstained preparations, likely due to the variations in the tissue microdissections as landmarks were relatively difficult to discriminate in unstained preparations relative to stained tissue sections. No significant differences were detected between SiS and AO staining conditions. Similar results for overall expression patterns were obtained from mouse hippocampus (Figure 4). Specifically, no significant differences in relative hybridization signal intensity were observed for CV, Th, H&E, and NF immunocytochemistry in mouse tissue sections, whereas a marked decrease was found in the SiS and AO conditions. Little or no RNA degradation was apparent, as expression levels for the TK and ECD domains of the trkA, trkB, and trkC ESTs did not display significant differences in relative expression levels in adjacent tissue sections stained for CV, Th, H&E, and NF immunocytochemistry. Expression levels of the trk receptors (TK and ECD) were below the limit of detection for tissue sections stained with SiS and AO.
Histograms illustrating global hybridization signal intensity from adjacent human post-mortem brain tissue sections prepared for histochemical stains and NF immunocytochemistry. (a) Regional hippocampal dissections and (b) individual populations of CA1 pyramidal neurons are depicted. Note the marked decrement in signal obtained from SiS- and AO-stained preparations (single asterisk denotes significance at P<0.0001; double asterisk denotes significance at P<0.0005). In contrast, no significant differences were detected for global hybridization signal intensity between CV, Th, H&E, and NF immunocytochemistry, indicating the utility of these stains in this paradigm.
Global hybridization signal intensity levels obtained from regional dissections of mouse hippocampus expressed as a percentage of hybridization signal intensity garnered from NF immunostained sections (set at 100%±standard deviation). Similar to human post-mortem material, no significant differences were detected between NF immunocytochemistry and CV-, Th-, and H&E- stained sections. Unstained (U) sections also displayed similar hybridization signal intensity levels, albeit a much larger variability, likely due to less precise microdissections. In contrast, highly significant decreases in SiS and AO were observed (asterisk denotes P<0.0001). No significant differences between SiS and AO were detected.
Expression profiling of microaspirated human and mouse CA1 pyramidal neurons revealed relatively equivalent hybridization signal intensity for the Nissl stains and NF immunocytochemistry for global gene expression (Figure 3b) as well as individual genes. In contrast, the SiS and AO stains displayed significantly lower hybridization signal intensity (P<0.0005). The relative hybridization levels for 50 representative genes (including cyto-skeletal elements, glutamate receptors, glutamate transporters, and glutamate receptor-interacting proteins) are depicted from regional hippocampal dissections (Figure 5a) and from 24 representative genes (including Alzheimer's disease-related genes) obtained from microaspirated CA1 neurons (Figure 5b). Note the wide dynamic range of relative signal intensities from low, moderate to high in the Nissl stained and NF immunocytochemistry preparations.
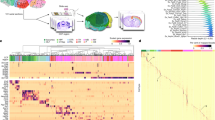
Color coded matrix plots illustrating relative expression levels for individual genes in different histochemical and immunocytochemical staining conditions in human regional hippocampal dissections and CA1 pyramidal neurons. (a) Depiction of expression levels of 50 genes, illustrating the relatively high expression in the Nissl stains and NF immunocytochemistry in contrast to SiS and AO from regional dissections. Note the low to moderate levels of variability across the Nissl stains and NF immunocytochemistry conditions. Abbreviations: ACTB, β-actin; ACTG, γ-actin; TUBA, α-tubulin; TUBB, β-tubulin; MAP1b, microtubule-associated protein 1b; MAP2, microtubule-associated protein 2; NF-L, neurofilament light subunit; NF-M, neurofilament medium subunit; NF-H, neurofilament heavy subunit; AKAP1, A kinase anchor protein 1; NES, nestin; UTRN, utrophin; GRIA1, α-amino-3-hydroxy-5-methyl-4-isoxazole propionate (AMPA) receptor GluR1; GRIA2, GluR2; GRIA3, GluR3; GRIA4, GluR4; GRIA5, kainate (KA) receptor GluR5; GRIA6, GluR6; GRIA7, GluR7; GRIK1, KA1; GRIK2, KA2; EAAT1, excitatory amino-acid transporter 1 (GLT-1) EAAT2, excitatory amino-acid transporter 2 (GLAST); EAAT3, excitatory amino-acid transporter 3 (EAAC1); GRIN1, N-methyl D-aspartate (NMDA) receptor NR1; GRIN2A, NMDAR2A; GRIN2B, NMDAR2B; GRIN2C, NMDAR2C; GRIN2D, NMDAR2D; GRM1a, glutamate receptor, metabotropic (GluR1α; GRM2, mGluR2, GRM3, mGluR3, GRM4, mGluR4; GRM5a, mGluR5α, GRM6, mGluR6; GRM7, mGluR7; GRM8, mGluR8; GRIP1, glutamate receptor ineracting protein 1; GRIP2, glutamate receptor interacting protein 2; PSD-95, postsynaptic density protein 95; SAP-102, synapse-associated protein 102; SAP-90; synapse-associated protein 90; HMR2A, homer2A; den-180; densin-180 CIT; citron; RBL2, p130. (b) Illustration of 24 genes from CA1 pyramidal neurons depicting the discrepant hybridization signal intensity levels observed in the Nissl stains and NF immunocytochemistry vs SiS and AO staining conditions. Similar to (a), note the relatively moderate levels of variability across the staining paradigm. Abbreviations: APLP1, amyloid precursor-like protein 1; APLP2, amyloid precursor-like protein 2; APP695, amyloid precursor protein; PRNP, prion protein; BACE, beta-site APP-cleaving enzyme; X11, APP-binding, family A, member 1; Fe65, APP-binding, family B, member 1; amylin, islet amyloid polypeptide; SAP, serum amyloid P-component; SAA4, serum amyloid A 4; CSEN, calsenilin; PSEN1, presenilin1; PSEN2, presenilin2; ALEX2, armadillo repeat protein 2; LRP1, low-density lipoprotein-related protein 1; HDLBP, high-density lipoprotein binding protein; A2 M, α2-macroglobulin; B2 M, β2-microglobulin; CSPG, chondroitin sulfate proteoglycan; DSPG, dextran sulfate proteoglycan; HSPG, heparan sulfate proteoglycan; HSP-40, heat shock protein 40; HSP-60, heat shock protein 60; HSP-70, heat shock protein 70.
Validation of the gene expression results was obtained by qPCR. Specifically, assessments of the cytoskeletal gene ACTB and housekeeping gene GAPDH were performed using hippocampal dissection samples acquired from adjacent stained tissue sections of mouse brain. The primers utilized identified only one PCR product that was sequence verified to be 100% identical to the corresponding wild-type sequence of ACTB and GAPDH, respectively. Results indicated similar levels of ACTB PCR product obtained from unstained mouse sections, Nissl-stained mouse sections, and NF immuno-labeled mouse sections. Statistically significant lower levels for SiS- and AO stained hippocampi (P<0.0005) were found (Figure 6a), consistent with the aforementioned array results. Moreover, comparable levels of GAPDH PCR product was observed between Nissl stains and NF immuno-cytochemistry, whereas significantly lower levels of GAPDH PCR product was detected using AO histochemistry (P<0.0001) (Figure 6b).
Histograms illustrating β-actin (a) and GAPDH (b) PCR products following real-time qPCR using mouse regional hippocampal dissections as input RNA sources. Similar to observations gleaned through cDNA array analysis, no differences were observed between U-, CV-, Th-, H&E-, and NF- stained hippocampal sections, whereas statistically significant lower levels of β-actin PCR products (asterisk denotes P<0.0005) were found for SiS and AO. In addition, no differences in GAPDH PCR products were found for Nissl stains and NF immunocytochemistry. However, a significant reduction was observed (double asterisk denotes P<0.0001) for AO-stained hippocampi.
Discussion
The present study was designed to test the hypothesis that distinct histochemical stains can be used in combination with state-of-the art molecular and cellular-based approaches to provide hybridization signal intensity detection that is similar to the standard set by immunocytochemical preparations. CV and hematoxylin have been employed successfully in concert with LCM studies using rodent brains,22, 23 and several histochemical and nuclear dyes have been used in studies with peripheral tissues as sources of input RNA.15, 16, 19 However, a systematic evaluation of the efficacy of histochemical stains in combination with RNA amplification and cDNA microarray analysis was lacking. Importantly, little or no information was available to evaluate histochemical stains in relation to immunocytochemical preparations, especially within the context of human brain tissue sections. The present results indicated that several Nissl stains work well along with NF immunolabeling to identify regions and neurons for subsequent RNA amplification and downstream genetic analyses. Although SiS and AO are useful for detecting nucleic acids (notably RNA species) in gels and tissue sections, these stains are not feasible for combination with tissue microdissection and cDNA array platforms.
The methodology employed in the present study comprised staining of the accrued fixed tissues and cells with a variety of histological stains followed by extracting total RNA through regional dissections and microaspiration via LCM. mRNA was then selectively converted to double-stranded cDNA and the starting mRNA was amplified linearly through in vitro transcription driven by a promoter of prokaryotic origin annealed to the converted cDNA.4, 40 Amplified RNA can be utilized for various downstream genetic manipulations and investigations, such as, but not limited to, gene profiling, qPCR, expression library construction, protein expression, and probe generation.4, 14, 40 In the present study, TC RNA amplification was followed by hybridization to custom-designed cDNA arrays for expression profiling analysis and qPCR. The CA1 molecular fingerprints were consistent in terms of signal intensity and expression level variability with published results using individual CA1 pyramidal neurons from post-mortem human brains.4, 5, 12
In terms of technology advancement, data sets derived from the present report indicate that Nissl stains and immunocytochemical procedures can be used successfully for tissue microdissection, RNA amplification, and downstream genetic analyses. These findings are particularly relevant for studying neuronal subtypes that have been difficult to identify based solely upon immunocytochemical procedures and phenotype analysis. Although many neuronal populations can be identified morphologically, for example, on the basis of cytoskeletal labeling (eg, neurofilaments) or by classes of calcium-binding proteins (eg, GABAergic interneurons), the ability to employ histochemical staining for gene expression profiling studies imparts a great deal of flexibility to the investigator. The process of selecting an appropriate dye or antibody will rely squarely on the cell type(s) to be identified in tissue sections. Specifically, populations of neurons that express proteins selectively such as cholinergic basal forebrain neurons7 or midbrain dopaminergic nigral and ventral tegmental neurons42, 43 are identified best by immunocytochemistry. However, many neuronal populations do not have a prominent and/or selective signature phenotype that differentiates them from other neuronal subtypes, and may be discriminated best by Nissl-stains. For example, subnuclei within the amygdala, hypothalamus, and thalamus fall into this latter category. Moreover, lamination patterns of the hippocampus, cerebral cortex, and cerebellar cortex are readily appreciable in Nissl stained preparations as well, thus are well suited for microdissection procedures. In addition to genomic-based studies, the proteomic potential of combining LCM with two-dimensional gel electrophoresis has been evaluated in peripheral tissues.44 In this paradigm, the use of histochemical stains will be important to identify regions, laminae, subnuclei, and cell types for subsequent proteomic analyses. The present results also indicated that the SiS used to detect proteins in gels for proteomic-based analyses will not be suitable to stain cells in tissue sections for subsequent downstream genetic analyses due to a relative paucity of hybridization signal intensity as compared to Nissl stains or immunocytochemical preparations.
The results of the present report indicate a vast discrepancy between the utility of CV, Th, and H&E vs SiS and AO on adjacent 6 μm-thick serial sections prepared to assess recovery of global hybridization signal intensity (on approximately 220 transcripts) as well as to assess individual genes. The experimental design was employed to minimize differences in mRNA concentration across the staining conditions (eg, due to trauma, fixation, or other variables), as it was relatively unlikely that the loss of hybridization signal intensity observed in the SiS and AO conditions arose from intrinsic differences in total RNA content from adjacent serial sections. A plausible mechanism for the observed effects is that the dyes themselves determined the availability and accessibility of mRNA species in situ. CV, Th, and H&E all label basophilia by binding to stacks of rough endoplasmic reticulum, or ‘Nissl’ substance.45 H&E stains nuclei as well, and this facet of the dye chemistry also does not appear deleterious to RNA accession.19, 21 In contrast, SiS appears to bind directly to RNA species, likely making accession to the polyadenylated or 5′ region of mRNAs more difficult for the TC primers and subsequent RNA amplification that is requisite when using minute amounts of input RNA. Specifically, Fourier transform infrared spectroscopy has been employed to demonstrate that silver binds to guanine, and to a lesser degree, cytosine, within RNA molecules with high affinity, approximately twice that of DNA.25 Further, SiS has been used for years to identify RNA bands on agarose gels subjected to electrophoresis.26, 46, 47 The observed decrement in hybridization signal intensity using AO-stained tissues may also be explained by the binding of the dye molecules to RNAs that are to be amplified from tissue sections. AO is a fluorescent dye that intercalates selectively into nucleic acids. Upon excitation with ultraviolet spectra, AO that intercalates into RNA emits an orange-red fluorescence (excitation at 440-470 nm, emission at 510 nm), whereas AO that intercalates into DNA (excitation at 480 nm, emission at 510 nm) emits a yellowish-green fluorescence.27, 28, 48 In brain tissue sections, labeled processes are contrasted by the pale green background of surrounding neuropil and white matter tracts that lack abundant nucleic acids.29, 30, 31 AO has the property of intercalating directly into nucleic acids (particularly double stranded) as well as binding electrostatically to the phosphate backbone,49, 50 presumably making primer accession difficult.
In conclusion, the combination of regional and single cell microdissection, TC RNA amplification, and cDNA microarrays enables high resolution and high throughput expression profiling of hundreds to thousands of genes simultaneously from a single neuron or a neuronal population. The next level of understanding of the cellular and molecular mechanisms underlying normative function and the pathophysiology of disease lies in the ability to combine these aforementioned technologies with appropriate neuronal circuits and models to recapitulate the structure and connectivity of these complex systems in vivo and in vitro. The use of several classical histochemical stains, either alone or in combination with immunocytochemical preparations, has been determined in this report to be feasible in both post-mortem human brain tissues as well as optimally prepared mouse brain tissues, both at the regional and single cell level within the hippocampus. Moreover, these data demonstrate that histochemical labeling is a viable experimental paradigm to perform single cell analyses in fixed human post-mortem tissues.
References
Kadkol SS, Gage WR, Pasternack GR . In situ hybridization—theory and practice. Mol Diagn 1999;4:169–183.
Van Deerlin VMD, Ginsberg SD, Lee VM-Y, et al. The use of fixed human post mortem brain tissue to study mRNA expression in neurodegenerative diseases: applications of microdissection and mRNA amplification. In: Geschwind DH, Gregg JP (eds). Microarrays for the Neurosciences: An Essential Guide. MIT Press: Boston, 2002, pp 201–235.
Mirnics K, Middleton FA, Marquez A, et al. Molecular characterization of schizophrenia viewed by microarray analysis of gene expression in prefrontal cortex. Neuron 2000;28:53–67.
Ginsberg SD, Che S . RNA amplification in brain tissues. Neurochem Res 2002;27:981–992.
Ginsberg SD, Hemby SE, Lee VM-Y, et al. Expression profile of transcripts in Alzheimer's disease tangle-bearing CA1 neurons. Ann Neurol 2000;48:77–87.
Ginsberg SD, Crino PB, Hemby SE, et al. Predominance of neuronal mRNAs in individual Alzheimer's disease senile plaques. Ann Neurol 1999;45:174–181.
Mufson EJ, Counts SE, Ginsberg SD . Single cell gene expression profiles of nucleus basalis cholinergic neurons in Alzheimer's disease. Neurochem Res 2002;27:1035–1048.
Eberwine J, Kacharmina JE, Andrews C, et al. mRNA expression analysis of tissue sections and single cells. J Neurosci 2001;21:8310–8314.
Fink L, Kinfe T, Seeger W, et al. Immunostaining for cell picking and real-time mRNA quantitation. Am J Pathol 2000;157:1459–1466.
Chow N, Cox C, Callahan LM, et al. Expression profiles of multiple genes in single neurons of Alzheimer's disease. Proc Natl Acad Sci USA 1998;95:9620–9625.
Hemby SE, Ginsberg SD, Brunk B, et al. Gene expression profile for schizophrenia: discrete neuron transcription patterns in the entorhinal cortex. Arch Gen Psychiatry 2002;59:631–640.
Hemby SE, Trojanowski JQ, Ginsberg SD . Neuron-specific age-related decreases in dopamine receptor subtype mRNAs. J Comp Neurol 2003;456:176–183.
Crino PB, Khodakhah K, Becker K, et al. Presence and phosphorylation of transcription factors in dendrites. Proc Natl Acad Sci USA 1998;95:2313–2318.
Ginsberg SD, Elarova I, Ruben M, et al. Single cell gene expression analysis: implications for neurodegenerative and neuropsychiatric disorders. Neurochem Res 2004;29:1054–1065.
Goldsworthy SM, Stockton PS, Trempus CS, et al. Effects of fixation on RNA extraction and amplification from laser capture microdissected tissue. Mol Carcinog 1999;25:86–91.
Luzzi V, Holtschlag V, Watson MA . Expression profiling of ductal carcinoma in situ by laser capture microdissection and high-density oligonucleotide arrays. Am J Pathol 2001;158:2005–2010.
To MD, Done SJ, Redston M, et al. Analysis of mRNA from microdissected frozen tissue sections without RNA isolation. Am J Pathol 1998;153:47–51.
Becker I, Becker K-F, Röhrl MH, et al. Single cell mutation analysis of tumors from stained histologic slides. Lab Invest 1996;75:801–807.
Tanji N, Ross MD, Cara A, et al. Effect of tissue processing on the ability to recover nucleic acid from specific renal tissue compartments by laser capture microdissection. Exp Nephrol 2001;9:229–234.
Murase T, Inagaki H, Eimoto T . Influence of histochemical and immunohistochemical stains on polymerase chain reaction. Mod Pathol 2000;13:147–151.
Ehrig T, Abdulkadir SA, Dintzis SM, et al. Quantitative amplification of genomic DNA from histological tissue sections after staining with nuclear dyes and laser capture microdissection. J Mol Diagn 2001;3:22–25.
Luo L, Salunga RC, Guo H, et al. Gene expression profiles of laser-captured adjacent neuronal subtypes. Nat Med 1999;5:117–122.
Vincent VA, DeVoss JJ, Ryan HS, et al. Analysis of neuronal gene expression with laser capture microdissection. J Neurosci Res 2002;69:578–586.
Ginsberg SD . Gene expression profiling using single cell microdissection combined with cDNA microarrays. In: Geschwind DH (ed). DNA Microarrays: The New Frontier in Gene Discovery and Gene Expression Analysis. Society for Neuroscience Press: Washington, 2001, pp 61–70.
Arakawa H, Neault JF, Tajmir-Riahi HA . Silver(I) complexes with DNA and RNA studied by fourier transform infrared spectroscopy and capillary electrophoresis. Biophys J 2001;81:1580–1587.
Somerville LL, Wang K . The ultrasensitive silver "protein" stain also detects nanograms of nucleic acids. Biochem Biophys Res Commun 1981;102:53–58.
von Bertalanffy L, Bickis I . Identification of cytoplasmic basophilia (ribonucleic acid) by fluorescence microscopy. J Histochem Cytochem 1956;4:481–493.
Mikel UV, Becker Jr RL . A comparative study of quantitative stains for DNA in image cytometry. Analyt Quant Cytol Histol 1991;13:253–260.
Mai JK, Schmidt-Kastner R, Tefett H-B . Use of acridine orange for histologic analysis of the central nervous system. J Histochem Cytochem 1984;32:97–104.
Ginsberg SD, Galvin JE, Chiu T-S, et al. RNA sequestration to pathological lesions of neurodegenerative disorders. Acta Neuropathol 1998;96:487–494.
Ginsberg SD, Crino PB, Lee VM-Y, et al. Sequestration of RNA in Alzheimer's disease neurofibrillary tangles and senile plaques. Ann Neurol 1997;41:200–209.
Ginsberg SD, Martin LJ . Axonal transection in adult rat brain induces transsynaptic apoptosis and persistent atrophy of target neurons. J Neurotrauma 2002;19:99–109.
Ginsberg SD, Portera-Cailliau C, Martin LJ . Fimbria-fornix transection and excitotoxicity produce similar neurodegeneration in septum. Neuroscience 1999;88:1059–1071.
Ginsberg SD, Hof PR, Young WG, et al. Noradrenergic innervation of the hypothalamus of rhesus monkeys: distribution of dopamine-β-hydroxylase-immunoreactive fibers and a quantitative analysis of varicosities in the paraventricular nucleus. J Comp Neurol 1993;327:597–611.
Ginsberg SD, Hof PR, McKinney WT, et al. The noradrenergic innervation density of the monkey paraventricular nucleus is not altered by early social deprivation. Neurosci Lett 1993;158:130–134.
Ginsberg SD, Galvin JE, Lee VM-Y, et al. Accumulation of intracellular amyloid-β peptide (Aβ 1–40) in mucopolysaccharidosis brains. J Neuropathol Exp Neurol 1999;58:815–824.
Gallyas F . A principle for silver staining of tissue elements by physical development. Acta Morphol Acad Sci Hung 1971;19:57–71.
Gallyas F . Silver staining of Alzheimer's neurofibrillary changes by means of physical development. Acta Morphol Acad Sci Hung 1971;19:1–8.
Lee VM-Y, Carden MJ, Schlaepfer WW, et al. Monoclonal antibodies distinguish several differentially phosphorylated states of the two largest rat neurofilament subunits (NF-H and NF-M) and demonstrate their existence in the normal nervous system of adult rats. J Neurosci 1987;7:3474–3488.
Che S, Ginsberg SD . Amplification of transcripts using terminal continuation. Lab Invest 2004;84:131–137.
Ginsberg SD, Schmidt ML, Crino PB, et al. Molecular pathology of Alzheimer's disease and related disorders. In: Peters A, Morrison JH (eds). Cerebral Cortex, Vol. 14, Neurodegenerative and Age-related Changes in Structure and Function of Cerebral Cortex Kluwer Academic/Plenum: New York, 1999, pp 603–653.
Fasulo WH, Hemby SE . Time-dependent changes in gene expression profiles of midbrain dopamine neurons following haloperidol administration. J Neurochem 2003;87:205–219.
Tang WX, Fasulo WH, Mash DC, et al. Molecular profiling of midbrain dopamine regions in cocaine overdose victims. J Neurochem 2003;85:911–924.
Craven RA, Totty N, Harnden P, et al. Laser capture microdissection and two-dimensional polyacrylamide gel electrophoresis: evaluation of tissue preparation and sample limitations. Am J Pathol 2002;160:815–822.
Peters A, Palay SL, DeF Webster H . The Fine Structure of the Nervous System. Neurons and Their Supporting Cells, 3rd edn. Oxford University Press, New York, 1991.
Su X . A refined method for silver staining of nucleic acids fixed on nitrocellulose. Anal Biochem 1987;163:535–536.
Skopp RN, Lane LC . Agarose gel electrophoresis of denatured RNA with silver staining. Anal Biochem 1988;169:132–137.
Schümmelfeder N . Histochemical significance of the polychromatic fluorescence induced in tissue stained with acridine orange. J Histochem Cytochem 1958;6:392–393.
McMaster GK, Carmichael GG . Analysis of single- and double-stranded nucleic acids on polyacrylamide and agarose gels by using glyoxal and acridine orange. Proc Natl Acad Sci USA 1977;74:4835–4838.
Lerman LS . Structural considerations in the interaction of DNA and acridines. J Mol Biol 1961;3:18–30.
Acknowledgements
We thank Ms Joanna M Eanniello, Ms Irina Elarova, Mr Marc D Ruben, and Ms Jena Vignola for expert technical assistance and the Center for Neurodegenerative Disease Research, University of Pennsylvania School of Medicine, for the human autopsy case materials. Support for this project (SDG) comes from the NINDS (NS43939), NIA (AG10668, AG14449), NCI (CA94520), and the Alzheimer's Association. We also express our appreciation to the families of the patients studied here who made this research possible.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Ginsberg, S., Che, S. Combined histochemical staining, RNA amplification, regional, and single cell cDNA analysis within the hippocampus. Lab Invest 84, 952–962 (2004). https://doi.org/10.1038/labinvest.3700110
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/labinvest.3700110
Keywords
This article is cited by
-
Molecular analysis of H&E- and Papanicolau-stained samples—systematic review
Histochemistry and Cell Biology (2020)
-
Frontal cortex and striatal cellular and molecular pathobiology in individuals with Down syndrome with and without dementia
Acta Neuropathologica (2019)
-
Method for In-Vivo Fluorescence Imaging Contrast Enhancement through Light Modulation
Journal of Fluorescence (2017)
-
Application of NeuroTrace staining in the fresh frozen brain samples to laser microdissection combined with quantitative RT-PCR analysis
BMC Research Notes (2015)
-
Enrichment of single neurons and defined brain regions from human brain tissue samples for subsequent proteome analysis
Journal of Neural Transmission (2015)