Abstract
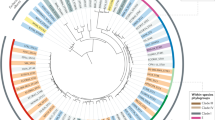
The mechanisms underlying the evolution and emergence of new bacterial pathogens are not well understood. To elucidate the evolution of pathogenic Escherichia coli strains, here we sequenced seven housekeeping genes to build a phylogenetic tree and trace the history of the acquisition of virulence genes. Compatibility analysis indicates that more than 70% of the informative sites agree with a single phylogeny, suggesting that recombination has not completely obscured the remnants of ancestral chromosomes1,2,3. On the basis of the rate of synonymous substitution for E. coli and Salmonella enterica (4.7 × 10-9 per site per year3), the radiation of clones began about 9 million years ago and the highly virulent pathogen responsible for epidemics of food poisoning, E. coli O157:H7, separated from a common ancestor of E. coli K-12 as long as 4.5 million years ago. Phylogenetic analysis reveals that old lineages of E. coli have acquired the same virulence factors in parallel, including a pathogenicity island involved in intestinal adhesion, a plasmid-borne haemolysin, and phage-encoded Shiga toxins. Such parallel evolution indicates that natural selection has favoured an ordered acquisition of genes and the progressive build-up of molecular mechanisms that increase virulence.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Guttman, D. S. & Dykhuizen, D. E. Clonal divergence in Escherichia coli as a result of recombination, not mutation. Science 266, 1380–1383 (1994).
Milkman, R. in Escherichia coli and Salmonella: Cellular and Molecular Biology 2nd edn (eds Neidhardt, F. C. et al.) 2663–2684 (American Society for Microbiology, Washington DC, 1996).
Lawrence, J. G. & Ochman, H. Molecular archaeology of the Escherichia coli genome. Proc. Natl Acad. Sci. USA 95, 9413–9417 ( 1998).
Nataro, J. P. & Kaper, J. B. Diarrheagenic Escherichia coli . Clin. Microbiol. Rev. 11, 142– 201 (1998).
Whittam, T. S. et al. Clonal relationships among Escherichia coli strains that cause hemorrhagic colitis and infantile diarrhea. Infect. Immun. 61, 1619–1629 ( 1993).
Jakobsen, I. B. & Easteal, S. A program for calculating and displaying compatibility matrices as an aid in determining reticulate evolution in molecular sequences. CABIOS 12, 291–295 (1996).
Wang, F. S., Whittam, T. S. & Selander, R. K. Evolutionary genetics of the isocitrate dehydrogenase gene (icd) in Escherichia coli and Salmonella enterica. J. Bacteriol. 179, 6551– 6559 (1997).
Bandelt, H. J. & Dress, A. W. M. Split decomposition: a new and useful approach to phylogenetic analysis of distance data. Mol. Phylogenet. Evol. 1, 242–252 (1992).
Dopazo, J., Dress, A. & von Haeseler, A. Split decomposition: a technique to analyze viral evolution. Proc. Natl Acad. Sci. USA 90, 10320– 10324 (1993).
Holmes, E. C., Urwin, R. & Maiden, M. C. The influence of recombination on the population structure and evolution of the human pathogen Neisseria meningitidis. Mol. Biol. Evol. 16, 741–749 (1999).
Takezaki, N., Rzhetsky, A. & Nei, M. Phylogenetic test of the molecular clock and linearized trees. Mol. Biol. Evol. 12, 823– 833 (1995).
Li, W.-H. Molecular Evolution (Sinauer Associates, Sunderland, Masschusetts, 1997).
Feng, P., Lampel, K. A., Karch, H. & Whittam, T. S. Genetic and phenotypic changes in the emergence of Escherichia coli O157:H7. J. Infect. Dis. 177, 1750–1753 (1998).
Karch, H. in Escherichia coli O157:H7 and Other Shiga Toxin-Producing E. coli Strains (eds Kaper, J. B. & O'Brien, A. D.) 183–194 (ASM, Washington DC, 1998).
Newland, J. W. & Neill, R. J. DNA probes for Shiga-like toxins I and II and for toxin-converting bacteriophages. J. Clin. Microbiol. 26, 1292–1297 (1988).
Schmidt, H., Beutin, L. & Karch, H. Molecular analysis of the plasmid-encoded hemolysin of Escherichia coli O157:H7 strain EDL 933. Infect. Immun. 63, 1055–1061 (1995).
Schmidt, H. & Karch, H. Enterohemolytic phenotypes and genotypes of shiga toxin-producing Escherichia coli O111 strains from patients with diarrhea and hemolytic-uremic syndrome. J. Clin. Microbiol. 34, 2364–2367 ( 1996).
Boyd, E. F. & Hartl, D. L. Chromosomal regions specific to pathogenic isolates of Escherichia coli have a phylogenetically clustered distribution. J. Bacteriol. 180, 1159– 1165 (1998).
Wieler, L. H., McDaniel, T. K., Whittam, T. S. & Kaper, J. B. Insertion site of the locus of enterocyte effacement in enteropathogenic and enterohemorrhagic Escherichia coli differs in relation to the clonal phylogeny of strains. FEMS Microbiol. Lett. 156, 49–53 (1997).
Sperandio, V. et al. Characterization of the locus of enterocyte effacement (LEE) in different enteropathogenic Escherichia coli (EPEC) and Shiga-toxin producing Escherichia coli (STEC) serotypes. FEMS Microbiol. Lett. 164, 133–139 ( 1998).
McGraw, E. A., Li, J., Selander, R. K. & Whittam, T. S. Molecular evolution and mosaic structure of α, β, and γ intimins of pathogenic Escherichia coli. Mol. Evol. Biol. 16, 12–22 (1999).
LeClerc, J. E., Li, B., Payne, W. L. & Cebula, T. A. High mutation frequencies among Escherichia coli and Salmonella pathogens. Science 274, 1208–1211 (1996).
Karaolis, D. K., Somara, S., Maneval, D. R., Jr., Johnson, J. A. & Kaper, J. B. A bacteriophage encoding a pathogenicity island, a type-IV pilus and a phage receptor in cholera bacteria. Nature 399, 375–379 (1999).
Whittam, T. S. in Escherichia coli O157:H7 and other Shiga toxin-producing E. coli strains (eds Kaper, J. B. & O'Brien, A. D.) 195– 209 (ASM, Washington DC, 1998).
Reid, S. D., Betting, D. J. & Whittam, T. S. Molecular detection and identification of intimin alleles in pathogenic Escherichia coli by multiplex PCR. J. Clin. Microbiol. 37, 2719–2722 (1999).
Thompson, J. D., Higgins, D. G. & Gibson, T. J. CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 22, 4673–4680 (1994).
Huson, D. H. SplitsTree: analyzing and visualizing evolutionary data. Bioinformatics 14, 68–73 ( 1998).
Kumar, S., Tamura, K. & Nei, M. MEGA: Molecular evolutionary genetics analysis, version 1.0. (Pennsylvania State Univ., Univ. Park, Pennsylvannia, 1993).
Felsenstein, J. PHYLIP—Phylogeny inference package (version 3.2). Cladistics 5, 164–166 ( 1989).
Acknowledgements
The authors thank S. Plock for technical assistance. This research was supported by NIH grants (to T.S.W. and R.K.S.), and the Enteric Pathogen Research Unit at the University of Maryland Medical School.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Reid, S., Herbelin, C., Bumbaugh, A. et al. Parallel evolution of virulence in pathogenic Escherichia coli. Nature 406, 64–67 (2000). https://doi.org/10.1038/35017546
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/35017546
This article is cited by
-
A bacterial autotransporter impairs innate immune responses by targeting the transcription factor TFE3
Nature Communications (2023)
-
Revisiting the Design of the Long-Term Evolution Experiment with Escherichia coli
Journal of Molecular Evolution (2023)
-
Colonization of gut microbiota by plasmid-carrying bacteria is facilitated by evolutionary adaptation to antibiotic treatment
The ISME Journal (2022)
-
Phylogroup stability contrasts with high within sequence type complex dynamics of Escherichia coli bloodstream infection isolates over a 12-year period
Genome Medicine (2021)
-
Population structure and genetic diversity of non-O157 Shiga toxin-producing Escherichia coli (STEC) clinical isolates from Michigan
Scientific Reports (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.