Abstract
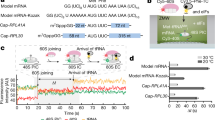
Initiation of eukaryotic protein synthesis begins with the ribosome separated into its 40S and 60S subunits1. The 40S subunit first binds eukaryotic initiation factor (eIF) 3 and an eIF2–GTP–initiator transfer RNA ternary complex. The resulting complex requires eIF1, eIF1A, eIF4A, eIF4B and eIF4F to bind to a messenger RNA and to scan to the initiation codon2. eIF5 stimulates hydrolysis of eIF2-bound GTP and eIF2 is released from the 48S complex formed at the initiation codon before it is joined by a 60S subunit to form an active 80S ribosome3,4,5,6,7,8. Here we show that hydrolysis of eIF2-bound GTP induced by eIF5 in 48S complexes is necessary but not sufficient for the subunits to join. A second factor termed eIF5B (relative molecular mass 175,000) is essential for this process. It is a homologue of the prokaryotic initiation factor IF2 (refs 6, 7) and, like it8,9,10,11,12, mediates joining of subunits and has a ribosome-dependent GTPase activity that is essential for its function.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Merrick, W. C. Mechanism and regulation of eukaryotic protein synthesis. Microbiol. Rev. 56, 291–315 ( 1992).
Pestova, T. V., Borukhov, S. I. & Hellen, C. U. T. Eukaryotic ribosomes require initiation factors 1 and 1A to locate initiation codons. Nature 394, 854–859 (1998).
Chakrabarti, A. & Maitra, U. Functions of eukaryotic initiation factor 5 in the formation of an 80S ribosomal polypeptide chain initiation complex. J. Biol. Chem. 266, 14039–14045 (1991).
Das, K., Chesevich, J. & Maitra, U. Molecular cloning and expression of cDNA for mammalian translation initiation factor 5. Proc. Natl Acad. Sci. USA 90, 3058–3062 (1993).
Huang, H.-K., Yoon, H., Hannig, E. M. & Donahue, T. F. GTP hydrolysis controls stringent selection of the AUG start codon during translation initiation in Saccharomyces cerevisiae. Genes Dev. 11, 2396–2413 (1997).
Choi, S. K., Lee, J. H., Zoll, W. L., Merrick, W. C. & Dever, T. E. Promotion of Met-tRNAMet binding to ribosomes by yIF2, a bacterial IF2 homolog in yeast. Science 280, 1757–1760 (1998).
Lee, J. H., Choi, S. K., Roll-Mecak, A., Burley, S. K. & Dever, T. E. Universal conservation in translation initiation revealed by human and archaeal homologs of bacterial translation factor IF2. Proc. Natl Acad. Sci. USA 96, 4342–4347 (1999).
Sacerdot, C., Dessen, P., Hershey, J. W. B., Plumbridge, J. A. & Grunberg-Manago, M. Sequence of the initiation factor IF2 gene; unusual protein features and homologies with elongation factors. Proc. Natl Acad. Sci. USA 81, 7787– 7791 (1984).
Kolakofsky, D., Dewey, K. F., Hershey, J. W. B. & Thach, R. E. Guanosine 5′-triphosphatase activity of initiation factor f2. Proc. Natl Acad. Sci. USA 61, 1066– 1070 (1968).
Godefroy-Colburn, T. et al. Light-scattering studies showing the effect of initiation factors on the reversible dissociation of Escherichia coli ribosomes. J. Mol. Biol. 94, 461– 478 (1975).
Luchin, S. et al. In vitro study of two dominant inhibitory GTPase mutants of Escherichia coli translation initiation factor IF2. Direct evidence that GTP hydrolysis is necessary for factor recycling. J. Biol. Chem. 274, 6074–6079 ( 1999).
Lockwood, A. H., Sarkar, P. & Maitra, U. Release of polypeptide chain initiation factor IF-2 during initiation complex formation. Proc. Natl Acad. Sci. USA 69, 3602–3605 ( 1972).
Merrick, W. C., Kemper, W. M. & Anderson, W. F. Purification and characterization of homogenous initiation factor M2A from rabbit reticulocytes. J. Biol. Chem. 250, 5556–5562 (1975).
Trachsel, H., Emi, B., Schreier, M. H. & Staehelin, T. Initiation of mammalian protein synthesis. II. The assembly of the initiation complex with purified initiation factors. J. Mol. Biol. 116, 755–767 (1977).
Benne, R., Brown-Luedi, M. L. & Hershey, J. W. B. Purification and characterization of protein synthesis initiation factors eIF-1, eIF-4C, eIF-4D, and eIF-5 from rabbit reticulocytes. J. Biol. Chem. 253, 3070– 3077 (1978).
Peterson, D. T., Safer, B. & Merrick, W. C. Role of eukaryotic initiation factor 5 in the formation of 80S initiation complexes. J. Biol. Chem. 254, 7730–7735 (1979).
Pestova, T. V., Shatsky, I. N., Fletcher, S. P., Jackson, R. J. & Hellen, C. U. T. A prokaryotic-like mode of binding of cytoplasmic eukaryotic ribosomes to the initiation codon during internal initiation of translation of Hepatitis C and Classical Swine fever virus RNAs. Genes Dev. 12, 67– 83 (1998).
Acknowledgements
We thank W. Merrick for discussions, D. Etchison and R. Schneider for antibodies, and L. Siconolfi-Baez for sequencing eIF5B. These studies were supported by grants from the NIH to C.U.T.H. and T.V.P.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Pestova, T., Lomakin, I., Lee, J. et al. The joining of ribosomal subunits in eukaryotes requires eIF5B. Nature 403, 332–335 (2000). https://doi.org/10.1038/35002118
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/35002118
This article is cited by
-
The molecular basis of translation initiation and its regulation in eukaryotes
Nature Reviews Molecular Cell Biology (2024)
-
The germline factor DDX4 contributes to the chemoresistance of small cell lung cancer cells
Communications Biology (2023)
-
Start codon-associated ribosomal frameshifting mediates nutrient stress adaptation
Nature Structural & Molecular Biology (2023)
-
Nonsense-mediated RNA decay: an emerging modulator of malignancy
Nature Reviews Cancer (2022)
-
Structural basis for the transition from translation initiation to elongation by an 80S-eIF5B complex
Nature Communications (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.