Abstract
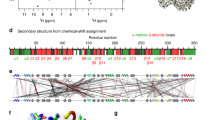
The geometric information used to solve three-dimensional (3D) structures of proteins by NMR spectroscopy resides in short (<5 Å) interproton-distance data1,2. To obtain these distances, the 1H-NMR spectrum must first be assigned using correlation and nuclear Overhauser effect (NOE) experiments to demonstrate through-bond (scalar) and through-space connectivities, respectively. Because the NOE is proportional to r–6, distance information can then be derived. The increased resolution afforded by extending NMR experiments into a second dimension3 enables one to detect and interpret effects that would not be possible in one dimension owing to extensive spectral overlap and much reduced information. A number of small protein structures have previously been solved in this way1,2. Extending this methodology to larger proteins, however, requires yet an additional improvement in resolution as overlap of cross-peaks in the two-dimensional (2D) NMR spectra present a major barrier to their unambiguous identification. One way of increasing the resolution is to extend the 2D-NMR experiments into a third dimension. We report here the applicability of three-dimensional NMR to macromolecules using the 46-residue protein al-purothionin as an example.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Wüthrich, K. NMR of Proteins (Wiley, New York, 1986).
Clore, G. M. & Gronenborn, A. M. Protein Engineering 1, 275–288 (1987).
Ernst, R. R., Bodenhausen, G. & Wokaun, A. Principles of Nuclear Magnetic Resonance in One and Two Dimensions (Clarendon, Oxford, 1987).
Griesinger, C., Sørensen, O. W. & Ernst, R. R. J. magn. Reson. 73, 574–579 (1987).
Griesinger, C., Sørensen, O. W. & Ernst, R. R. J. Am. chem. Soc. 109, 7227–7228 (1987).
Brauschweiler, R., Madsen, J. C., Griesinger, C., Sørensen, O. W. & Ernst, R. R. J. magn. Reson. 73, 380–385 (1987).
Jeener, J., Meier, B. H., Bachmann, P. & Ernst, R. R. J. chem. Phys. 71, 4546–4553 (1979).
Anil Kumar, Ernst, R. R. & Wüthrich, K. Biochem. biophys. Res. Commun. 95, 1–6 (1980).
Aue, W. P., Bartholdi, E. & Ernst, R. R. J. chem. Phys. 64, 2229–2246 (1976).
Braunschweiler, L. & Ernst, R. R. J. magn. Reson. 53, 521–528 (1983).
Davis, D. G. & Bax, A. J. Am. chem. Soc. 107, 2821–2822 (1985).
Bauer, C., Freeman, R., Frenkiel, T., Keeler, J. & Shaka, A. J. J. magn. Reson. 58, 442–457 (1984).
Clore, G. M. et al. J. molec. Biol. 193, 571–578 (1987).
Jones, T. A. J. appl. Crystallogr. 11, 268–272 (1978).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Oschkinat, H., Griesinger, C., Kraulis, P. et al. Three-dimensional NMR spectroscopy of a protein in solution. Nature 332, 374–376 (1988). https://doi.org/10.1038/332374a0
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/332374a0
This article is cited by
-
CONNJUR Workflow Builder: a software integration environment for spectral reconstruction
Journal of Biomolecular NMR (2015)
-
Structural insights on the pamoic acid and the 8 kDa domain of DNA polymerase beta complex: Towards the design of higher-affinity inhibitors
BMC Structural Biology (2008)
-
A nomenclature and data model to describe NMR experiments
Journal of Biomolecular NMR (2006)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.