Abstract
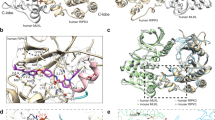
When activated, membrane-bound receptors for Fas and tumour-necrosis factor initiate programmed cell death by recruiting the death domain of the adaptor protein FADD1 (Mort1; ref. 2) to the membrane. FADD then activates caspase 8 (ref. 3) (also known as FLICE4 or MACH5) through an interaction between the death-effector domains of FADD and caspase 8. This ultimately leads to the apoptotic response. Death-effector domains and homologous protein modules known as caspase-recruitment domains6 have been found in several proteins1,2,3,4,5,6,7,8,9,10,11,12,13,14 and are important regulators of caspase (FLICE) activity and of apoptosis. Here we describe the solution structure of a soluble, biologically active mutant of the FADD death-effector domain. The structure consists of six antiparallel, amphipathic α-helices and resembles the overall fold of the death domains of Fas15 and p75 (ref. 16). Despite this structural similarity, mutations that inhibit protein–protein interactions involving the Fas death domain have no effect when introduced into the FADD death-effector domain. Instead, a hydrophobic region of the FADD death-effector domain that is not present in the death domains is vital for binding to FLICE and for apoptotic activity.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Chinnaiyan, A. M., O'Rourke, K., Tewari, M. & Dixit, V. M. FADD, a novel death domain-containing protein, interacts with the death domain of Fas and initiates apoptosis. Cell 81, 505–512 (1995).
Boldin, M. P. et al. Anovel protein that interacts with the death domain of Fas/APO1 contains a sequence motif related to the death domain. J. Biol. Chem. 270, 7795–7798 (1995).
Alnemri, E. S. et al. Human ICE/CED-3 protease nomenclature. Cell 87, 171 (1996).
Muzio, M. et al. FLICE, a novel FADD-homologous ICE/CED-3-like protease, is recruited to the CD95 (Fas/APO-1) death-inducing signaling complex. Cell 85, 817–827 (1996).
Boldin, M. P., Goncharov, T. M., Goltsev, Y. W. & Wallach, D. Involvement of MACH, a novel MORT1/FADD-interacting protease, in Fas/APO-1- and TNF receptor-induced cell death. Cell 85, 803–815 (1996).
Hofmann, K., Bucher, P. & Tschopp, J. The CARD domain: a new apoptotic signalling motif. Trends Biochem. Sci. 22, 155–156 (1997).
Duan, H. & Dixit, V. M. RAIDD is a new ‘death’ adaptor molecule. Nature 385, 86–89 (1997).
Bertin, J. et al. Death effector domain-containing herpesvirus and poxvirus proteins inhibit both Fas- and TNFR1-induced apoptosis. Proc. Natl Acad. Sci. USA 94, 1172–1176 (1997).
Thome, M. et al. Viral FLICE-inhibitory proteins (FLIPs) prevent apoptosis induced by death receptors. Nature 386, 517–521 (1997).
Hu, S., Vinchenz, C., Buller, M. & Dixit, V. M. Anovel family of viral death effector domain-containing molecules that inhibit both CD-95- and tumor necrosis factor receptor-1-induced apoptosis. J. Biol. Chem. 272, 9621–9624 (1997).
Irmler, M. et al. Inhibition of death receptor signals by cellular FLIP. Nature 388, 190–195 (1997).
Shu, H. B., Halpin, D. R. & Goeddel, D. V. Casper is a FADD- and caspase-related inducer of apoptosis. Immunity 6, 751–763 (1997).
Srinivasula, S. M. et al. FLAME-1, a novel FADD-like anti-apoptotic molecule that regulates Fas/TNFR1-induced apoptosis. J. Biol. Chem. 272, 18542–18545 (1997).
Goltsev, Y. V. et al. CASH, a novel caspase homologue with death effector domains. J. Biol. Chem. 272, 19641–19644 (1997).
Huang, B., Eberstadt, M., Olejniczak, E. T., Meadows, R. P. & Fesik, S. W. NMR structure and mutagenesis of the Fas (APO-1/CD95) death domain. Nature 384, 638–641 (1996).
Liepinsh, E., Ilag, L. L., Otting, G. & Ibanez, C. F. NMR structure of the death domain of the p75 neurotrophin receptor. EMBO J. 16, 4999–5005 (1997).
Eberstadt, M., Huang, B., Olejnizak, E. T. & Fesik, S. W. The lymphoproliferation mutation in Fas locally unfolds the Fas death domain. Nature Struct. Biol. 4, 983–986 (1997).
Watanabe-Fukunaga, R., Brannan, C. I., Copeland, N. G., Jenkins, N. A. & Nagata, S. Lymphoproliferation disorder in mice explained by defects in fas antigen that mediates apoptosis. Nature 356, 314–317 (1992).
Tartaglia, L. A., Ayres, T. M., Wong, G. H. & Goeddel, D. V. Anovel domain within the 55 kd TNF receptor signals cell death. Cell 74, 845–853 (1993).
Keane, M. M., Ettenberg, S. A., Lowrey, G. A., Russell, E. K. & Lipkowitz, S. Fas expression and function in normal and malignant breast cell lines. Cancer Res. 56, 4791–4798 (1996).
Yamazaki, T., Lee, W., Arrowsmith, C. H., Muhandiram, D. R. & Kay, L. E. Asuite of triple resonance NMR experiments for the backbone assignement of 15N, 13C, 2H labeled proteins with high sensitivity. J. Am. Chem. Soc. 116, 11655–11666 (1994).
Clore, G. M. & Gronenborn, A. M. Multidimensional heteronuclear nuclear magnetic resonance of proteins. Meth. Enzymol. 239, 349–363 (1994).
Logan, T. M., Olejniczak, E. T., Xu, R. X. & Fesik, S. W. Ageneral method for assigning NMR spectra of denatured proteins using 3D HC(CO)NH-TOCSY triple resonance experiments. J. Biomol. NMR 3, 225–231 (1993).
Neri, D., Szyperski, T., Otting, G., Senn, H. & Wüthrich, K. Stereospecific nuclear magnetic resonance assignments of the methyl groups of valien and leucine in the DNA-binding domain of the 434 repressor by biosynthetically directed fractional 13C labeling. Biochemistry 28, 7510–7516 (1989).
Vuister, G. W. & Bax, A. Quantitative J correlation: a new approach for measuring homonuclear three-bond J(HNHα) coupling constants in 15N-enriched proteins. J. Am. Chem. Soc. 115, 7772–7777 (1993).
Kuszewski, J., Nilges, M. & Brünger, A. T. Sampling and efficiency of metric matrix distance geometry: a novel partial metrization algorithm. J. Biomol. NMR 2, 33–56 (1992).
Brünger, A. T. X-PLOR 3.2 Manual (Yale Univ. Press, New Haven, 1992).
Carson, M. J. Ribbon models of macromolecules. J. Mol. Graph. 5, 103–106 (1987).
Acknowledgements
The 750-MHz NMR data were acquired at the National Magnetic Resonance Facility at Madison, which is supported by NIH, NSF, and the University of Wisconsin.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Eberstadt, M., Huang, B., Chen, Z. et al. NMR structure and mutagenesis of the FADD (Mort1) death-effector domain. Nature 392, 941–945 (1998). https://doi.org/10.1038/31972
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/31972
This article is cited by
-
FADD as a key molecular player in cancer progression
Molecular Medicine (2022)
-
Comprehensive review of ASC structure and function in immune homeostasis and disease
Molecular Biology Reports (2020)
-
Network Connectivity, Centrality and Fragmentation in the Greek-Key Protein Topology
The Protein Journal (2019)
-
A motif within the armadillo repeat of Parkinson’s-linked LRRK2 interacts with FADD to hijack the extrinsic death pathway
Scientific Reports (2018)
-
Structural Characterizations of the Fas Receptor and the Fas-Associated Protein with Death Domain Interactions
The Protein Journal (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.