Abstract
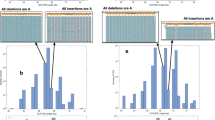
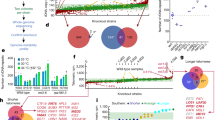
Halobacterium halobium is an obligately halophilic archaebacterium1 of interest to molecular biologists for many reasons2–5, one of which is the unexplained high frequency (10−4–10−2 mutants per cell plated) at which it yields readily identifiable and unstable mutants4. We showed previously5 that the genome of H. halobium contains many (>50) families of repeated sequences whose members are dispersed on both chromosome and plasmid. Here we report that most if not all of the members of most of these repeat sequence families effect or are affected by spontaneous genomic rearrangements. Quantitative analyses show that such repeat sequence-associated rearrangements (which may be of several kinds) occur at high frequencies (>4 × 10−3 events per family per cell generation), while unique-sequence DNAs are physically stable. The presence of so many families of elements of such great instability in the halobacterial genome gives it an unusual degree of structural and perhaps functional plasticity.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Woese, C. R. Scient. Am. 244(6), 94–106 (1981).
Bayley, S. T. & Morton, R. A. CRC crit. Rev. Microbiol. 6, 151–205 (1978).
Weidinger, G., Klotz, G. & Goebel, W. Plasmid 2, 377–386 (1979).
Pfeifer, F., Weidinger, G. & Goebel, W. J. Bact. 145, 375–381 (1981).
Sapienza, C. & Doolittle, W. F. Nature 295, 384–389 (1982).
Southern, E. M. J. molec. Biol. 98, 503–517 (1975).
Feller, W. An Introduction to Probability Theory and its Applications 3rd edn (Wiley, New York, 1968).
Schnabel, H. et al. EMBO J. (in the press).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Sapienza, C., Rose, M. & Doolittle, W. High-frequency genomic rearrangements involving archaebacterial repeat sequence elements. Nature 299, 182–185 (1982). https://doi.org/10.1038/299182a0
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/299182a0
This article is cited by
-
Comparative chloroplast genome analysis of seven extant Citrullus species insight into genetic variation, phylogenetic relationships, and selective pressure
Scientific Reports (2023)
-
Haloferax volcanii for biotechnology applications: challenges, current state and perspectives
Applied Microbiology and Biotechnology (2020)
-
Archaeal genetics — the third way
Nature Reviews Genetics (2005)
-
Distribution and characterization of plasmid-related sequences in the chromosomal DNA of different thermophilic Methanobacterium strains
Molecular and General Genetics MGG (1993)
-
Analysis of secondary, integration sites for IS117 in Streptomyces lividans and their role in the generation of chromosomal deletions
Molecular and General Genetics MGG (1993)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.