Abstract
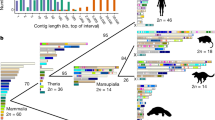
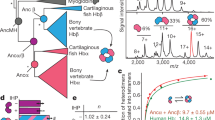
Recent years have seen rapid growth in amino acid sequence data on globins and nucleotide sequence data on haemoglobin genes and pseudogenes, and cladistic analysis1 of these data continues to reveal new facets of globin evolution. Our present findings demonstrate: (1) avian and mammalian embryonic α genes (π and ξ, respectively) had a monophyletic origin involving an α locus duplication about 400 Myr ago soon after the duplication which separated α and β genes; (2) much later in phylogeny, independent β-gene duplications produced the embryonic ρ locus of birds and embryonic ɛ and fetal γ loci of mammals. This parallels the earlier finding2 that myoglobins evolved more than once from generalized globin ancestors. Here we support the view2 that such globin evolution resulted from natural selection acting on mutations in duplicated genes. Thus, our evidence contradicts the neutralist view3,4 in which almost all amino acid substitutions in descent to extant globins evaded positive selection.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Goodman, M. Prog. Biophys. molec. Biol. 38, 105–164 (1981).
Goodman, M. J. molec. Evol. 17, 114–120 (1981).
Kimura, M. Nature 217, 624–626 (1968).
Kimura, M. J. molec. Evol. 17, 110–113 (1981).
Goodman, M. in Calcium-Binding Proteins: Structure and Function (eds Siegel, F. L., Carafoli, E., Kretsinger, R. H., MacLennan, D. H. & Wasserman, R. H.) 347–354 (Elsevier, New York, 1980).
Baba, M. L., Darga, L. L., Goodman, M. & Czelusniak, J. J. molec. Evol. 17, 197–213 (1981).
De Jong, W. W., Zweers, A. & Goodman, M. Nature 292, 538–540 (1981).
Tashian, R. E., Hewett-Emmett, D. & Goodman, M. in Protides of the Biological Fluids Colloq. 28 (ed. Peeters, H.) 153–156 (Pergamon, Oxford, 1980).
Beintema, J. et al. J. molec. Evol. 10, 49–71 (1977).
Ladner, R. C., Heidner, E. J. & Perutz, M. F. J. mlec. Biol. 114, 385–414 (1977).
Fermi, G. & Perutz, M. F. J. molec. Biol. 114, 421–431 (1977).
Li, W-H., Gojóbori, T. & Nei, M. Nature 292, 237–239 (1981).
Cleary, M. L., Schon, E. A. & Lingrel, J. B. Cell 26, 181–190 (1981).
Efstratiadis, A. et al. Cell 21, 653–668 (1980).
Hewett-Emmett, D. et al. Fedn Proc. 40, 1591 (1981).
Goodman, M., Romero-Herrera, A. E., Czelusniak, J., Dene, H. & Tashian, R. E. in Macromolecular Sequences in Systematic and Evolutionary Biology (ed. Goodman, M) (Plenum, New York, in the press).
Dodgson, J. B., McCune, K. C., Rusling, D. J., Krust, A. & Engel, J. D. Proc. natn. Acad. Sci. U.S.A. 78, 5998–6002 (1981).
Aschauer, J., Sanguansermsri, T. & Braunitzer, G. Hoppe-Seyler's Z. physiol. Chem. 362, 1159–1162 (1981).
Roninson, I. B. & Ingram, V. M. Proc. natn. Acad. Sci. U.S.A. 78, 4782 (1981).
Hardison, R. C. J. biol. Chem. 2566, 11780–11786 (1981).
Hansen, J. N., Konkel, D. A. & Leder, P. J. biol. Chem. 257, 1048–1052 (1982).
Holmquist, R. J. molec. Biol. 135, 929–938 (1979).
Wilson, J. T. et al. J. biol. Chem. 255, 2807–2815 (1980).
Heindell, H. C. et al. Cell 15, 43–54 (1978).
Proudfoot, N. J. & Maniatis, T. Cell 21, 537–544 (1980).
Nishioka, Y. & Leder, P. Cell 18, 875–882 (1979).
Vanin, E. F., Goldberg, G. I., Tucker, P. W. & Smithies, O. Nature 286, 222–226 (1980).
Nishioka, Y., Leder, A. & Leder, P. Proc. natn. Acad. Sci. U.S.A. 77, 2806–2809 (1980).
Paddock, G. V. & Gaubatz, J. Eur. J. Biochem. 117, 269–273 (1981).
Richardson, L., Cappello, J., Cochran, M. D., Armentrout, R. W. & Brown, R. D. Devl Biol. 78, 161–172 (1980).
Williams, J. G., Kay, R. M. & Patient, R. K. Nucleic Acids Res. 8, 4247–4258 (1980).
Richards, R. I. et al. Nucleic Acids Res. 7, 1137–1146 (1980).
Niessing, J. Biochem. Int. 2, 113–120 (1981).
Haupe, A., Therwath, A., Soriano, P. & Galibert, F. Gene 14, 11–21 (1980).
Barelle, F. E., Shoulders, C. C. & Proudfoot, N. J. Cell 21, 621–626 (1980).
Slightom, J. L., Blechl, A. E. & Smithies, O. Cell 21, 627–638 (1980).
Konkel, D. A., Maizel, J. V. Jr & Leder, P. Cell 18, 865–873 (1981).
Cleary, M. L., Haynes, J. R., Schon, E. A. & Lingrel, J. B. Nucleic Acids Res. 8, 4791–4802 (1980).
Lacy, E. & Maniatis, T. Cell 21, 545–553 (1980).
Van Ooyen, A., van den Berg, J., Mantai, N. & Weissman, C. Science 206, 337–344 (1979).
Hardison, R. C. et al. Cell 18, 1285–1297 (1979).
Spritz, R. A., De Riel, J. K., Forget, G. B. & Weissman, S. Cell 21, 639–646 (1980).
Lawn, R. M., Efstratiadis, A., O'Connell, C. & Maniatis, T. Cell 21, 647–651 (1980).
Partington, G. A. & Baralle, F. E. J. molec. Biol. 145, 463–470 (1981).
Haynes, J. R. et al. J. biol. Chem. 255, 6355–6367 (1980).
Haynes, J. R., Rosteck, P. Jr & Lingrel, J. B. Proc. natn. Acad. Sci. U.S.A. 77, 7127–1731 (1980).
Kretschmer, P. J. et al. J. biol. Chem. 256, 1975–1982 (1981).
Barrie, P. A., Jeffreys, A. J. & Scott, A. F. J. molec. Biol. 149, 319–336 (1981).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Czelusniak, J., Goodman, M., Hewett-Emmett, D. et al. Phylogenetic origins and adaptive evolution of avian and mammalian haemoglobin genes. Nature 298, 297–300 (1982). https://doi.org/10.1038/298297a0
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/298297a0
This article is cited by
-
Recent genome duplications facilitate the phenotypic diversity of Hb repertoire in the Cyprinidae
Science China Life Sciences (2021)
-
The effects of linkage on comparative estimators of selection
BMC Evolutionary Biology (2013)
-
Conversion events in gene clusters
BMC Evolutionary Biology (2011)
-
Genomic organization and evolution of the Atlantic salmon hemoglobin repertoire
BMC Genomics (2010)
-
GIGA: a simple, efficient algorithm for gene tree inference in the genomic age
BMC Bioinformatics (2010)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.