Abstract
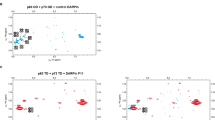
Tumor suppressor p53 forms a homo-tetramer through its COOH-terminal oligomerization domain and acts as a sequence-specific transcription factor. We have analysed the interrelation among the transcriptional activities, the structure and the cancer-related mutations in the oligomerization domain by using a comprehensive missense mutation library. Here, we examined the ability of 184 mutant p53s in the domain to form an oligomer by expressing these mutant p53s in yeast, and compared the data with the previous information. We showed that specific residues in the α-helix and the β-strand of the oligomerization domain were critical for both oligomer formation and sequence-specific transactivation, and the activities were closely related. In particular, the α-helix was more sensitive to amino-acid substitutions than the β-strand. We found identity in the interrelation between the two activities, that is, monomer mutants were transcriptionally inactive whereas dimer and tetramer mutants retained their transcriptional activities. In TP53 mutation databases, a small number of mutations have been reported in this domain. Surprisingly, most do not encode p53s defective in functional properties. These results indicate that, although oligomer formation is essential for p53 transactivation function, the inactivation of oligomer formation and therefore the inactivation of transactivation may not be essential for tumor suppression by p53 because they do not lead to oncogenic proteins.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Ashcroft M, Kubbutat MH and Vousden KH . (1999). Mol. Cell. Biol., 19, 1751–1758.
Barlev NA, Liu L, Chehab NH, Mansfield K, Harris KG, Halazonetis TD and Berger SL . (2001). Mol. Cell, 8, 1243–1254.
Brachmann RK, Vidal M and Boeke JD . (1996). Proc. Natl. Acad. Sci. USA, 93, 4091–4095.
Chene P and Bechter E . (1999). J. Mol. Biol., 288, 891–897.
Chene P, Mittl P and Grutter M . (1997). J. Mol. Biol., 273, 873–881.
Clore GM, Omichinski JG, Sakaguchi K, Zambrano N, Sakamoto H, Appella E and Gronenborn AM . (1995). Science, 267, 1515–1516.
Davison TS, Yin P, Nie E, Kay C and Arrowsmith CH . (1998). Oncogene, 17, 651–656.
DiGiammarino EL, Lee AS, Cadwell C, Zhang W, Bothner B, Ribeiro RC, Zambetti G and Kriwacki RW . (2002). Nat. Struct. Biol., 9, 12–16.
el-Deiry WS, Tokino T, Velculescu VE, Levy DB, Parsons R, Trent JM, Lin D, Mercer WE, Kinzler KW and Vogelstein B . (1993). Cell, 75, 817–825.
Halazonetis TD and Kandil AN . (1993). EMBO J., 12, 5057–5064.
Ishioka C, Englert C, Winge P, Yan YX, Engelstein M and Friend SH . (1995). Oncogene, 10, 1485–1492.
Ishioka C, Shimodaira H, Englert C, Shimada A, Osada M, Jia LQ, Suzuki T, Gamo M and Kanamaru R . (1997). Biochem. Biophys. Res. Commun., 232, 54–60.
Jeffrey PD, Gorina S and Pavletich NP . (1995). Science, 267, 1498–1502.
Kakudo Y, Shibata H, Otsuka K, Kato S and Ishioka C . (2005). Cancer Res., 65, 2108–2114.
Kato S, Han SY, Liu W, Otsuka K, Shibata H, Kanamaru R and Ishioka C . (2003). Proc. Natl. Acad. Sci. USA, 100, 8424–8429.
Lee W, Harvey TS, Yin Y, Yau P, Litchfield D and Arrowsmith CH . (1994). Nat. Struct. Biol., 1, 877–890.
Lomax ME, Barnes DM, Hupp TR, Picksley SM and Camplejohn RS . (1998). Oncogene, 17, 643–649.
Marchenko ND, Zaika A and Moll UM . (2000). J. Biol. Chem., 275, 16202–16212.
Mateu MG and Fersht AR . (1998). EMBO J., 17, 2748–2758.
Mihara M, Erster S, Zaika A, Petrenko O, Chittenden T, Pancoska P and Moll UM . (2003). Mol. Cell, 11, 577–590.
Miller M, Lubkowski J, Rao JK, Danishefsky AT, Omichinski JG, Sakaguchi K, Sakamoto H, Appella E, Gronenborn AM and Clore GM . (1996). FEBS Lett., 399, 166–170.
Mittl PR, Chene P and Grutter MG . (1998). Acta Crystallogr D Biol. Crystallogr., 54, 86–89.
Miyashita T and Reed JC . (1995). Cell, 80, 293–299.
Oda E, Ohki R, Murasawa H, Nemoto J, Shibue T, Yamashita T, Tokino T, Taniguchi T and Tanaka N . (2000). Science, 288, 1053–1058.
Owen-Schaub LB, Zhang W, Cusack JC, Angelo LS, Santee SM, Fujiwara T, Roth JA, Deisseroth AB, Zhang WW, Kruzel E and Radinsky R . (1995). Mol. Cell. Biol., 15, 3032–3040.
Polyak K, Xia Y, Zweier JL, Kinzler KW and Vogelstein B . (1997). Nature, 389, 300–305.
Roth J, Dobbelstein M, Freedman DA, Shenk T and Levine AJ . (1998). EMBO J., 17, 554–564.
Sakaguchi K, Herrera JE, Saito S, Miki T, Bustin M, Vassilev A, Anderson CW and Appella E . (1998). Genes Dev., 12, 2831–2841.
Shaulian E, Zauberman A, Ginsberg D and Oren M . (1992). Mol. Cell. Biol., 12, 5581–5592.
Shaulian E, Zauberman A, Milner J, Davies EA and Oren M . (1993). EMBO J., 12, 2789–2797.
Slingerland JM and Benchimol S . (1991). J. Cell. Physiol., 148, 391–395.
Soussi T, Kato S, Levy PP and Ishioka C . (2005). Hum. Mutat., 25, 6–17.
Srivastava S, Wang S, Tong YA, Pirollo K and Chang EH . (1993). Oncogene, 8, 2449–2456.
Stommel JM, Marchenko ND, Jimenez GS, Moll UM, Hope TJ and Wahl GM . (1999). EMBO J., 18, 1660–1672.
Tanaka H, Arakawa H, Yamaguchi T, Shiraishi K, Fukuda S, Matsui K, Takei Y and Nakamura Y . (2000). Nature, 404, 42–49.
Tao W and Levine AJ . (1999). Proc. Natl. Acad. Sci. USA, 96, 6937–6941.
Unger T, Mietz JA, Scheffner M, Yee CL and Howley PM . (1993). Mol. Cell. Biol., 13, 5186–5194.
Unger T, Nau MM, Segal S and Minna JD . (1992). EMBO J., 11, 1383–1390.
Varley JM, McGown G, Thorncroft M, Cochrane S, Morrison P, Woll P, Kelsey AM, Mitchell EL, Boyle J, Birch JM and Evans DG . (1996). Oncogene, 12, 2437–2442.
Zacchi P, Gostissa M, Uchida T, Salvagno C, Avolio F, Volinia S, Ronai Z, Blandino G, Schneider C and Del Sal G . (2002). Nature, 419, 853–857.
Zheng H, You H, Zhou XZ, Murray SA, Uchida T, Wulf G, Gu L, Tang X, Lu KP and Xiao ZX . (2002). Nature, 419, 849–853.
Acknowledgements
We thank Shuang-Yin Han and Wen Liu for their contribution to the construction of the p53 mutation library. We also thank Yuka Fujimaki for technical assistance. This study was supported in part by grants-in-aid from the Ministry of Education and Science, Sports and Culture (to CI and SK) and the Gonryo Medical Foundation (CI).
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplemenatry Information accompanies the paper on Oncogene website (http://www.nature.com/onc)
Rights and permissions
About this article
Cite this article
Kawaguchi, T., Kato, S., Otsuka, K. et al. The relationship among p53 oligomer formation, structure and transcriptional activity using a comprehensive missense mutation library. Oncogene 24, 6976–6981 (2005). https://doi.org/10.1038/sj.onc.1208839
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1208839
Keywords
This article is cited by
-
Structure-dependent recruitment and diffusion of guest proteins in liquid droplets of FUS
Scientific Reports (2022)
-
TP53 mutations in functional corticotroph tumors are linked to invasion and worse clinical outcome
Acta Neuropathologica Communications (2022)
-
Molecular principles of recruitment and dynamics of guest proteins in liquid droplets
Scientific Reports (2021)
-
NEK10 tyrosine phosphorylates p53 and controls its transcriptional activity
Oncogene (2020)
-
The multiple mechanisms that regulate p53 activity and cell fate
Nature Reviews Molecular Cell Biology (2019)